Ligand Preparation
Introduction
Accurately determining the structure and conformation of small molecule drugs is fundamental to subsequent calculations and analyses in drug design.
The Ligand Preparation module on the Hermite platform offers functionalities for processing small molecule structures. These include filtering molecular properties, generating isomers and conformations, and determining the protonation state of ionizable groups.
This tutorial guides you through using the Hermite platform's Ligand Preparation module to prepare the following ligand:
1. Ligand Preparation
1.1 Import and Check Ligand Files
Navigate to "File" → "Import Structure".
In the "Import Structure" window, click "Select Files", choose the local "9ligands.sdf" file, and click "Import". The file will then be available for viewing in the 3D Workspace.
1.2 Ligand Preparation Entry
Go to "Function" → "General" → "Ligand Preparation".
1.3 Select Ligands
Click the "Select File" button to upload the local file "9ligands.sdf".
Note:For operating instructions on more ligand selection methods, please refer to "Protein Selection, Ligand Selection and Pocket Setting".
1.4 Generation of Isomers and Conformations
Enable "Generate Isomers" and enter 4 in the "Generate isomers (per ligand)" field. (Keep other options at default)
Check "Possible Protonation States" and keep the default values.
Under "Conformation", check "Generate Conformations" and enter 8 to create 8 conformations for each isomer of each ligand.
Click "Next".
Note:Descriptions of all parameters are shown below:
| Parameter | Meaning | Note | |
| Generate Isomers | Chiral Isomers | Generating chiral isomers | Set a limit for the number of chiral centers to a maximum of 5. Enable the option "Retain specified chiralities (vary other chiral centers)" to maintain specific chiralities while allowing changes in other chiral centers. |
| Cis-/Trans- Isomers | Generating cis-trans isomers | ||
| Generate isomers (per ligand) | Determine how many isomers each ligand generates | (Number of Isomers Generated) * (Number of Configurations Generated) shall not exceed 100. | |
| Possible Protonation States | Select the pH range and determine the protonation environment | ||
| Conformation | Retain Original Conformation | Retain the original conformation | Ensure that the input ligand contains structural information (SMILES has no structural information, so it is recommended to check "Generate Conformation". |
| Generate Conformations | Generate new conformations | (Number of Isomers Generated) * (Number of Configurations Generated) shall not exceed 100. | |
| Sample ring conformations | Ring sampling | When dealing with macrocyclic systems (typically those with more than nine ring systems) or polypeptide systems (generally consisting of more than 10 amino acids), it is recommended to enable this option. |
1.5 Ligand Filtering and Task Submission
Filter Setting: Filter ligands based on a range of chemical properties. (This step will be skipped here)
Click "Submit" to initiate the task.
2. Ligand Preparation Results View
2.1 Job Entry
Navigate to "Job" and search for the task using "Job ID" 167861. Results usually become available within 1 min after submission for this task.
Click "Show" (first icon under "Operation") to view the task results.
2.2 Result List
Enlarge the result list to examine the generated ligands. The list includes the following information:
| Name | Meaning |
| Original Ligand | Displays the 2D structural diagram and name of the original ligand |
| Generated Ligand | Shows the 2D structural diagram and name of the generated ligand |
| Isomer | Type of isomer |
| MW | Molecular weight |
| LogP | Lipid-water partition coefficient |
| HBA | Number of hydrogen bond receptors |
| HBD | Number of hydrogen bond donors |
| ROTB | Number of rotatable keys |
Advanced Search: Filter and select ligands based on five chemical properties for display in the results list.
Operations in the 'Operation' Column:
3D: Click to visualize the ligand's conformation in the 3D Workspace.
Download: Click to download the ligand's .sdf file.
Use the "Ligand" and "Isomer" filters in the upper left corner to sort and display corresponding ligands or isomers in the result list.
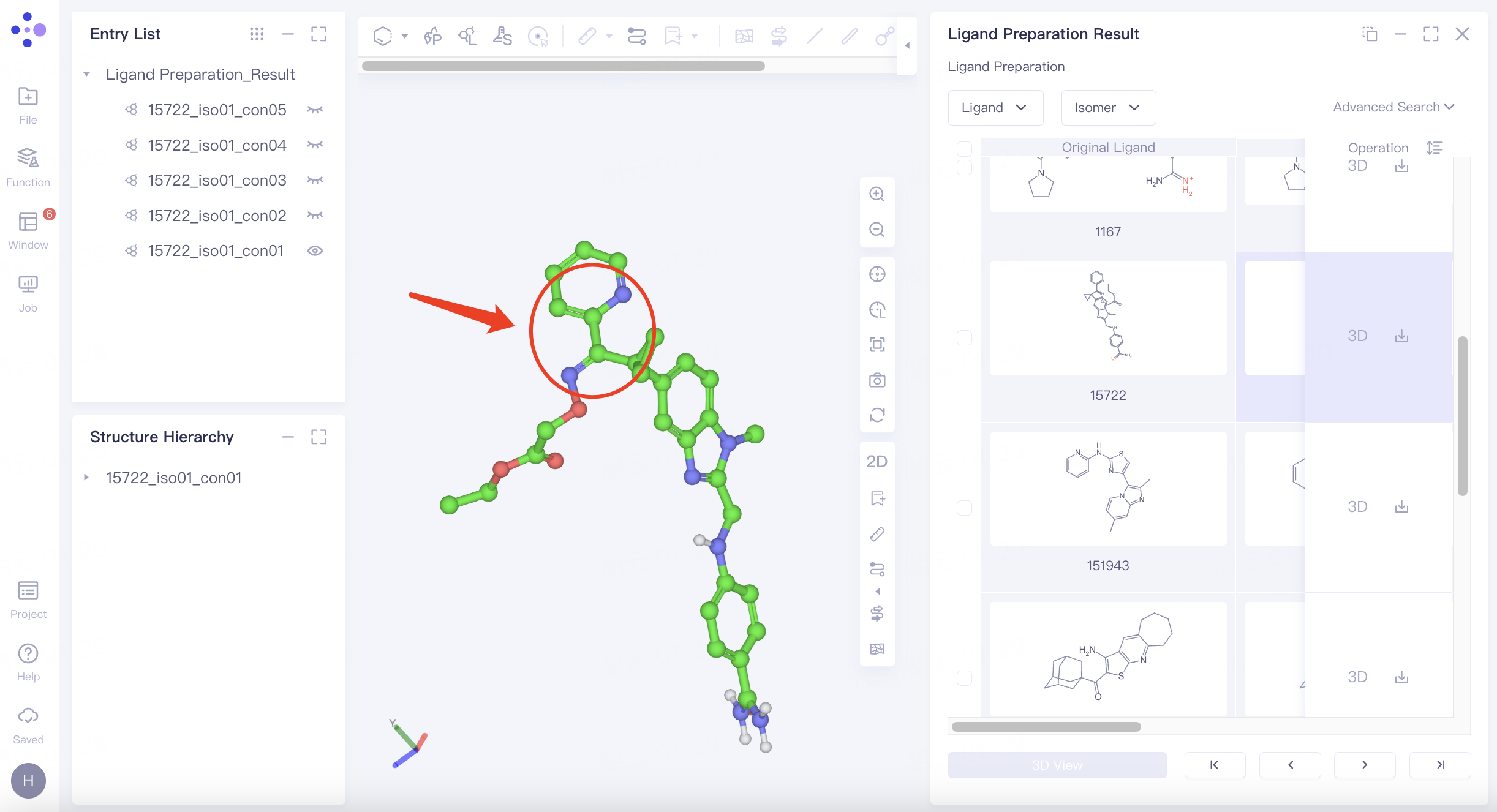
2.3 Ligand structure viewing
Ligand 15722 generates a trans isomer. To view the ligand, click on the "3D" icon to the right of the ligand named "15722" to display the ligand in the 3D Workspace window.
The pyridine and imine groups connected on both sides of the carbon-carbon single bond form a conjugate, which makes the carbon-carbon single bond non-rotatable, thus generating cis-trans isomers.