Similar Molecules Search
Introduction
In the process of optimizing compounds, or when adopting fragment-based drug design (Fragment Based Drug Design, FBDD) strategies, it is often necessary to search for similar compounds in the compound library. Searching for similar compounds already in the compound library can help us Predict whether the compound under development has other targets and ADME properties for subsequent research.
The Similar Molecules Search module of the Hermite platform provides a molecular search function based on the SMARTS representation of compound fragments. With the massive virtual compound library built into the Hermite platform, you can quickly search for molecules containing the same molecular fragments in a relatively large chemical space.
This tutorial is based on the Substructure Search module of the Hermite platform to search the Alinda database for molecules with compound structures.
Molecules used in this tutorial: [H]N([H])c1nnc(c(n1)-c1c([H])c([H])c([H])c1[H])-c1c([H])c(nc(c1[H])C([H])([H])[H])C([H])([H])[H], abbreviated as 4g.
1. Usage
1.1 Entrance
The left general menu bar Function → Molecular Recommendation → Substructure Search .
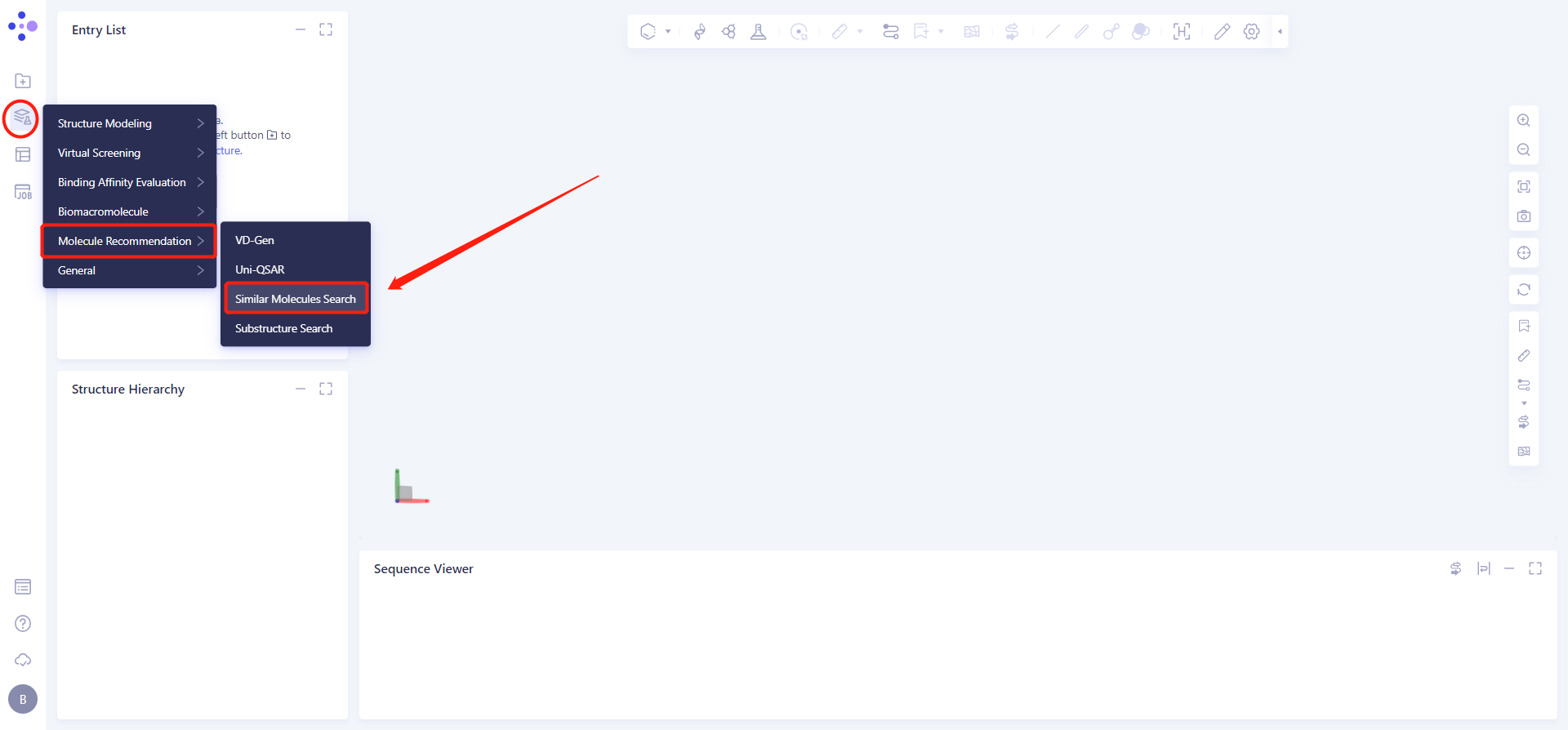
The operation box of Substructure Search (shown in the red box) appears on the right. The overall interface is as follows:
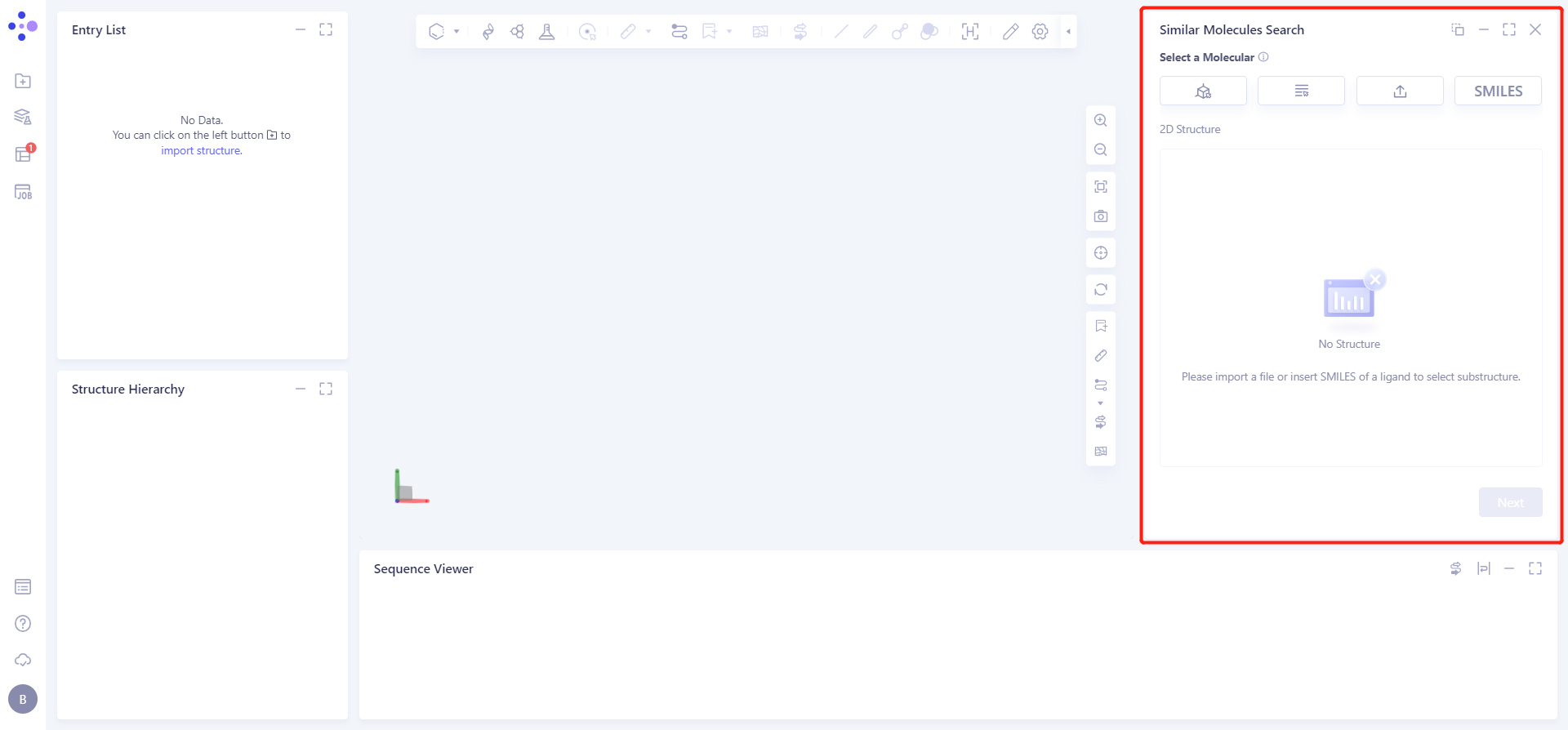
1.2 Operation
1.2.1 upload ligand structure
Click Select from SMILES checkbox → Import SMILES interface pops up → Enter molecule → Click OK.
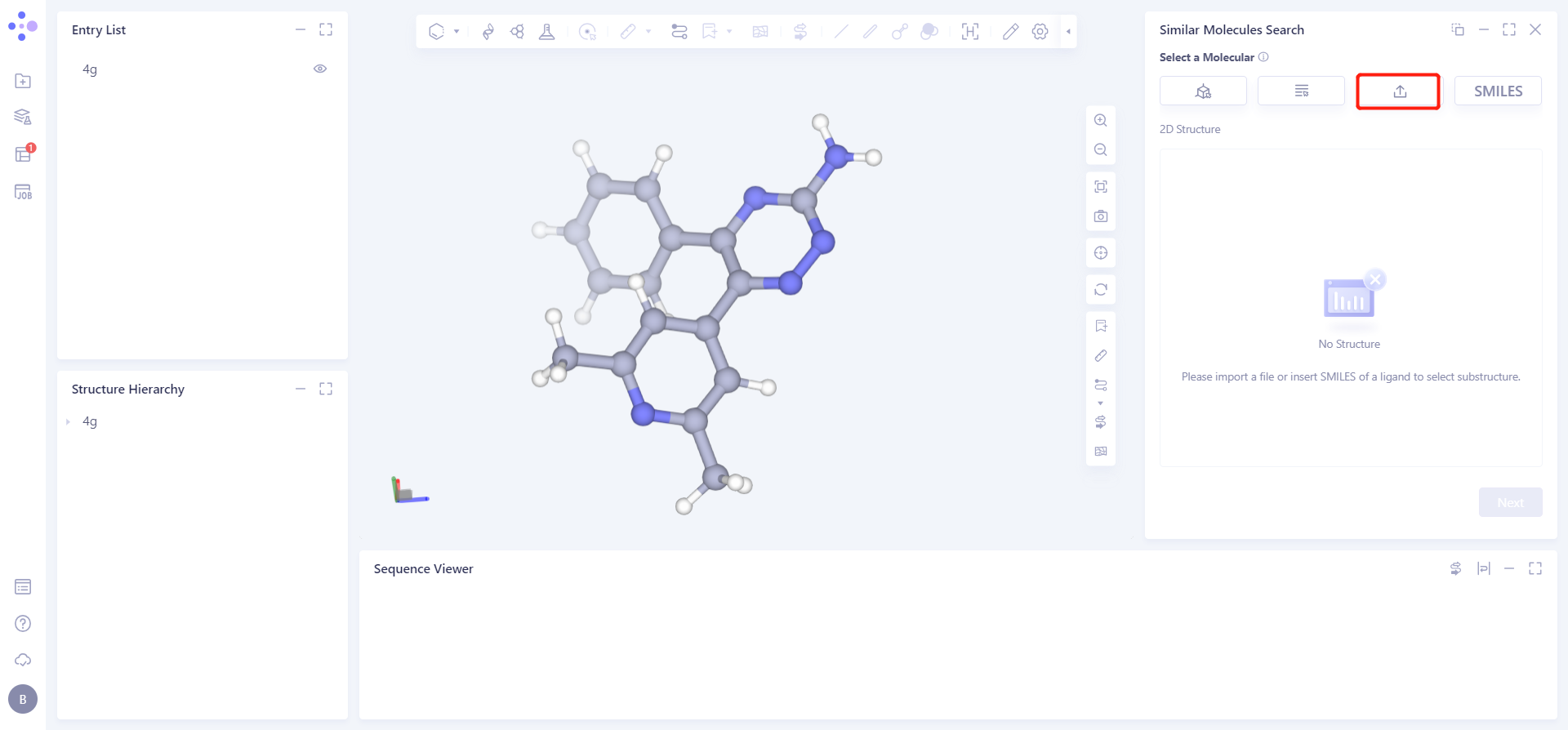
1.2.2 select dataset
Click the From Database checkbox → pop up the Select Ligands interface → select the desired molecular database (optional) → click OK.
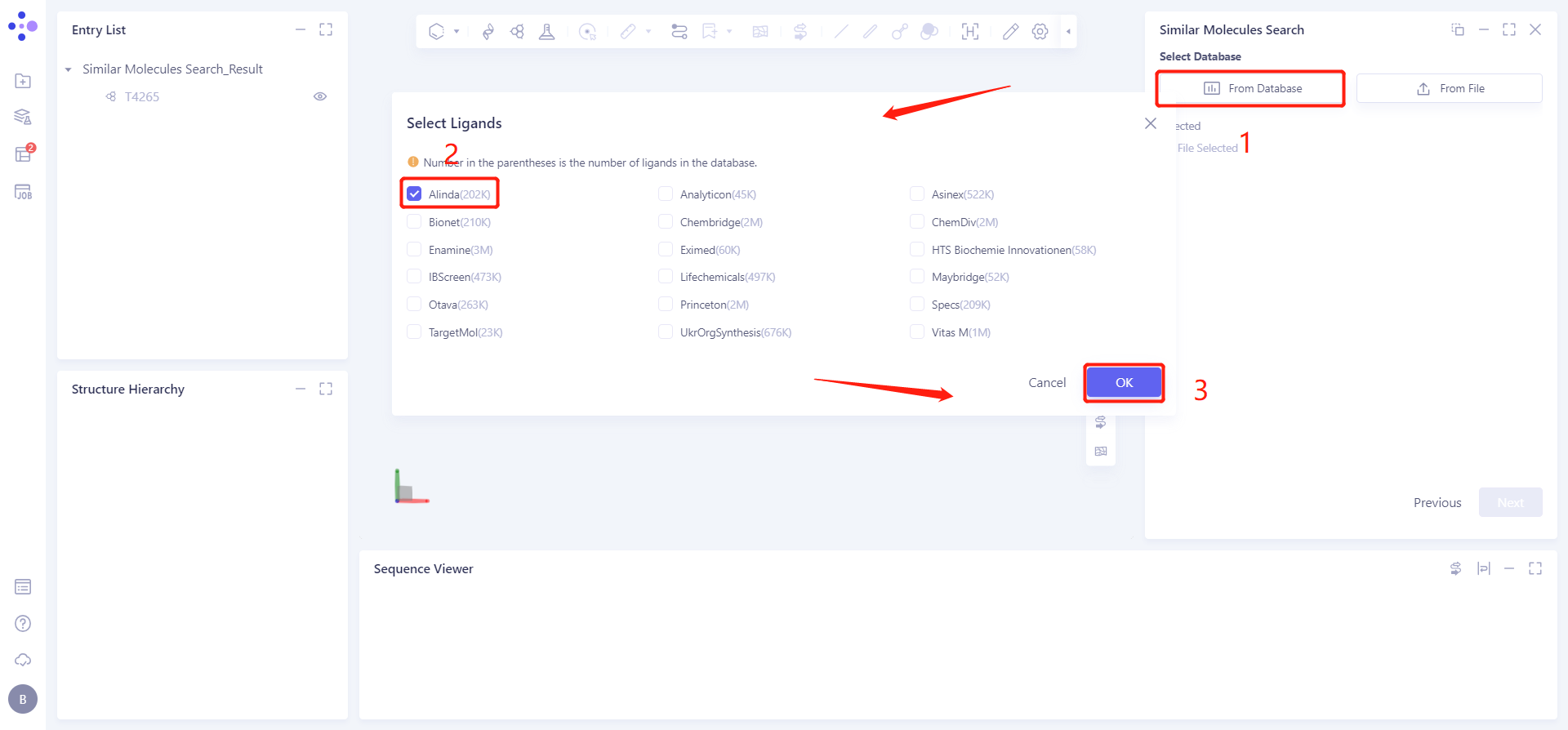
1.2.3 settings
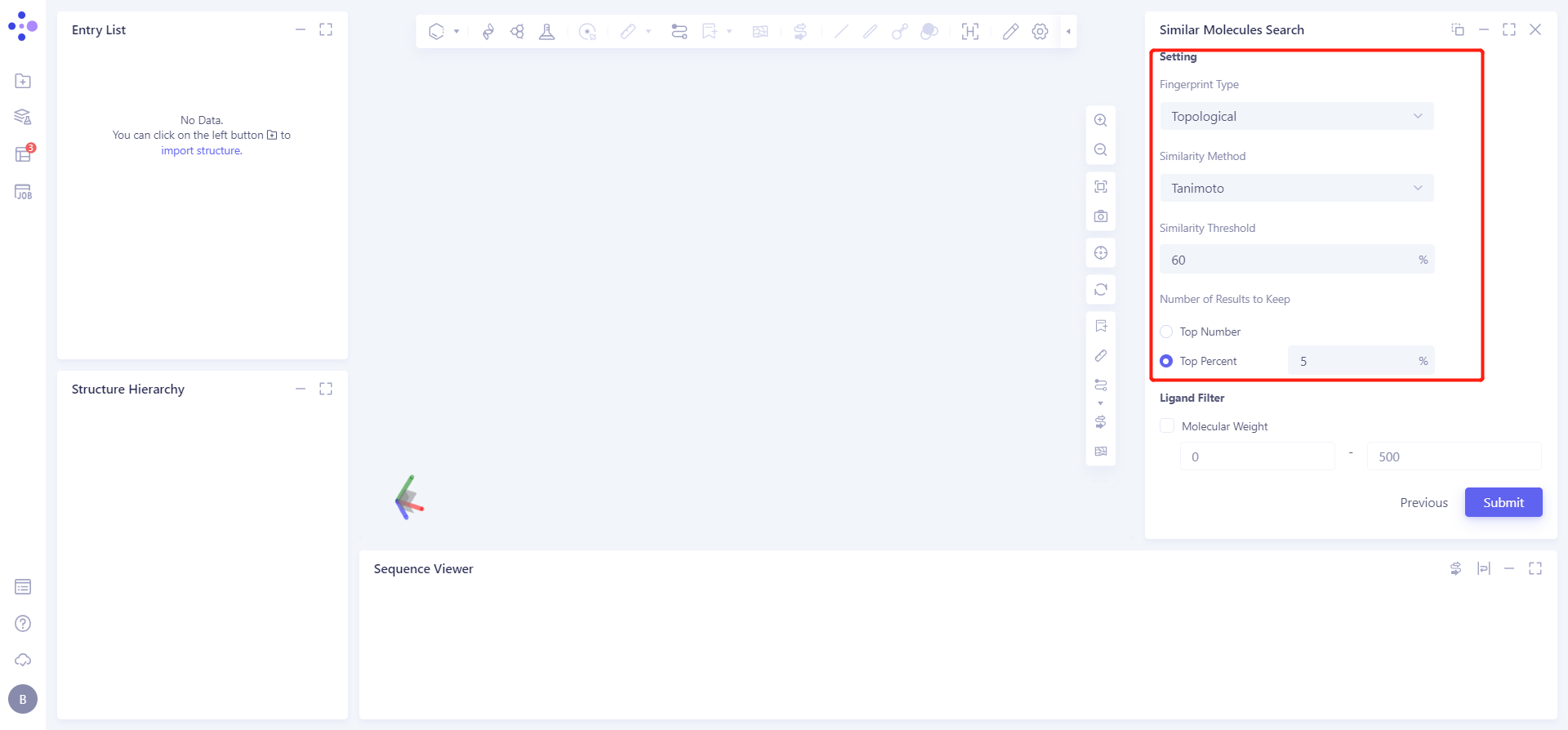
In Settings, Similarty Threshold is set to 60, and the other options remain the default settings;
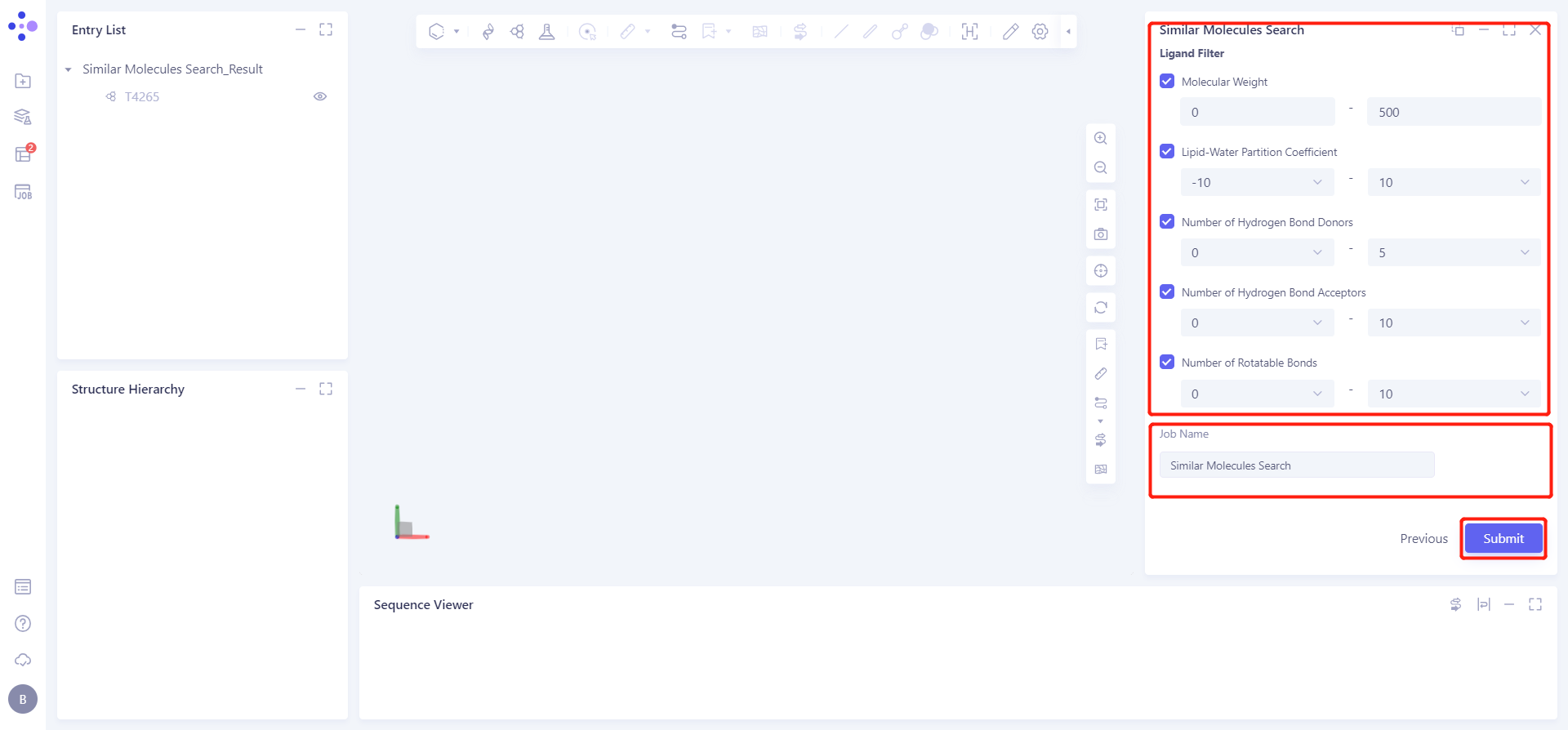
Check the filter conditions under Ligand Filter, the parameters are the default parameters;
The Job Name is Similar Molecules Search.
Click Submit to submit the task.
2. Results analysis
2.1 Entrance
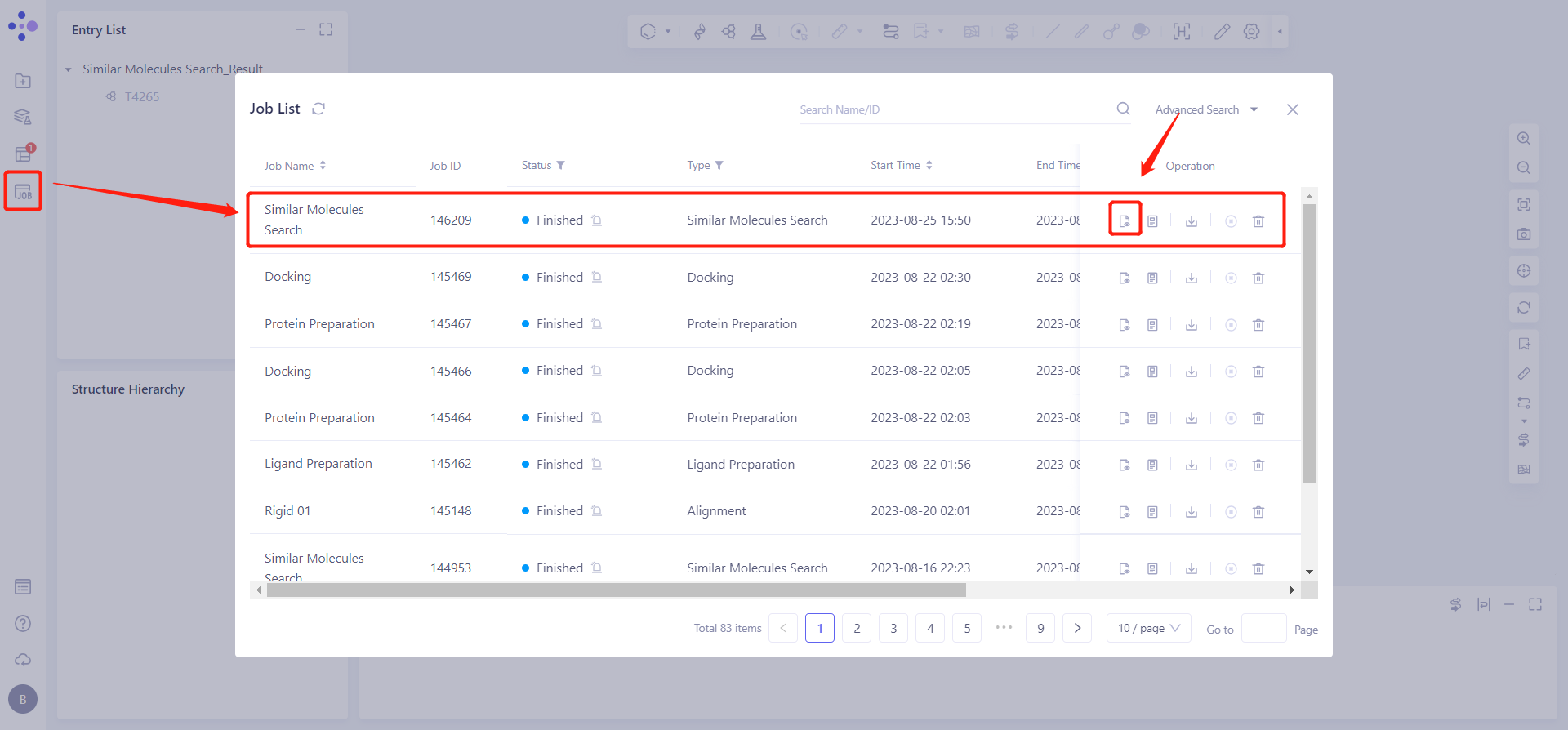
The left general menu bar Menu Job → Find the task Similar Molecules Search → Click Show in the Operation column to view the result of the task.
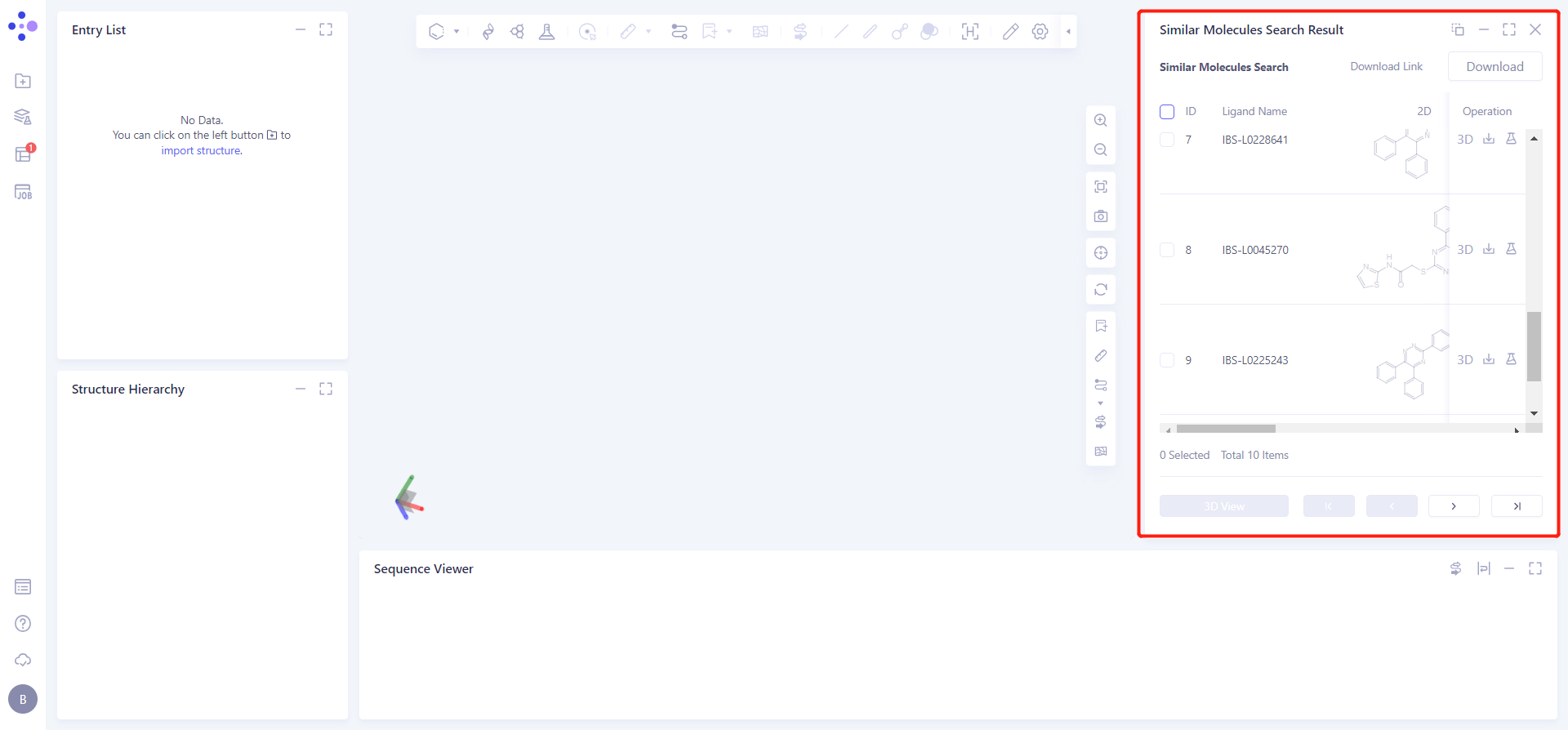
The display interface is shown on the right.
 |  |
2.2 Results display and download
Ten molecules were screened that were similar to the small molecule 4g and matched the parameters we set.
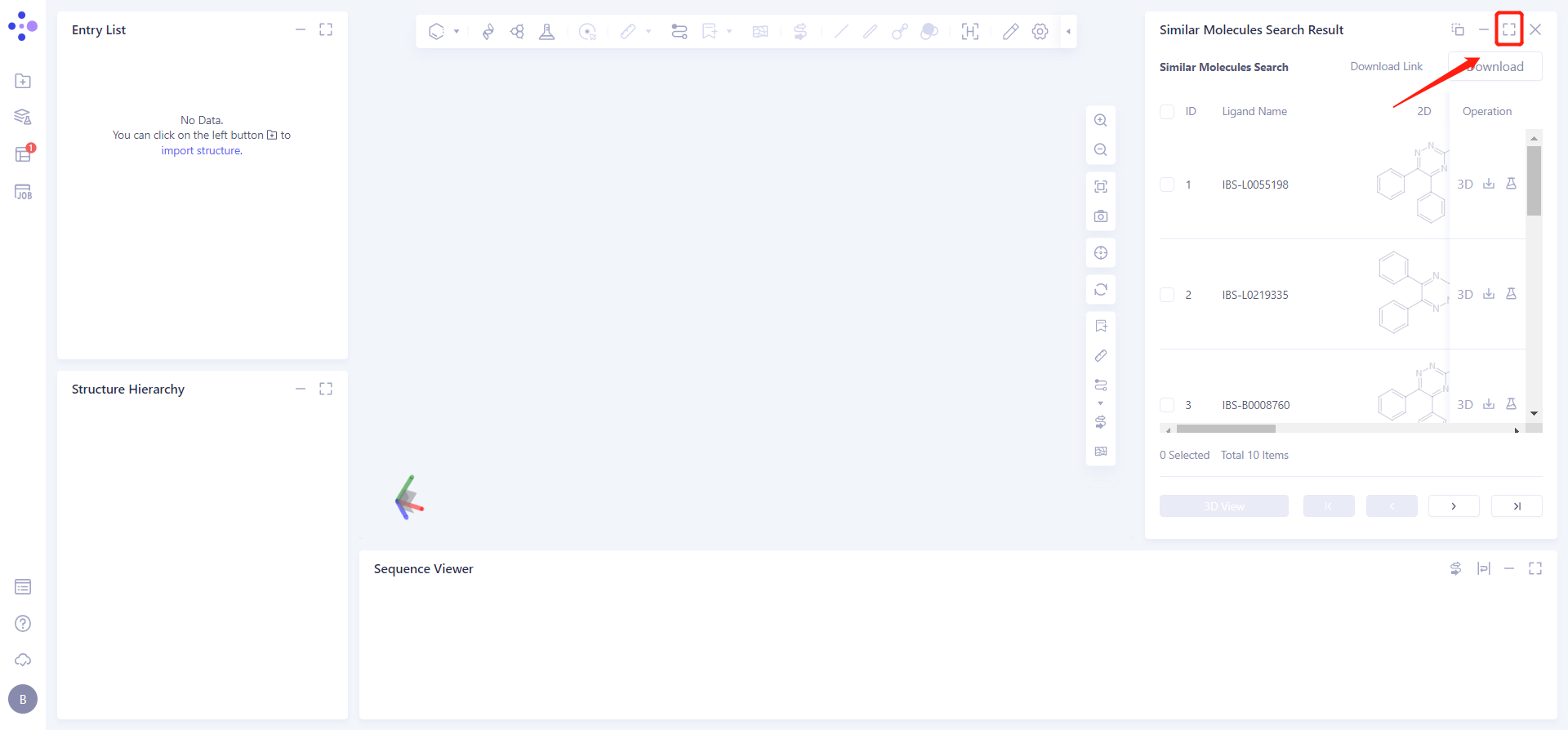
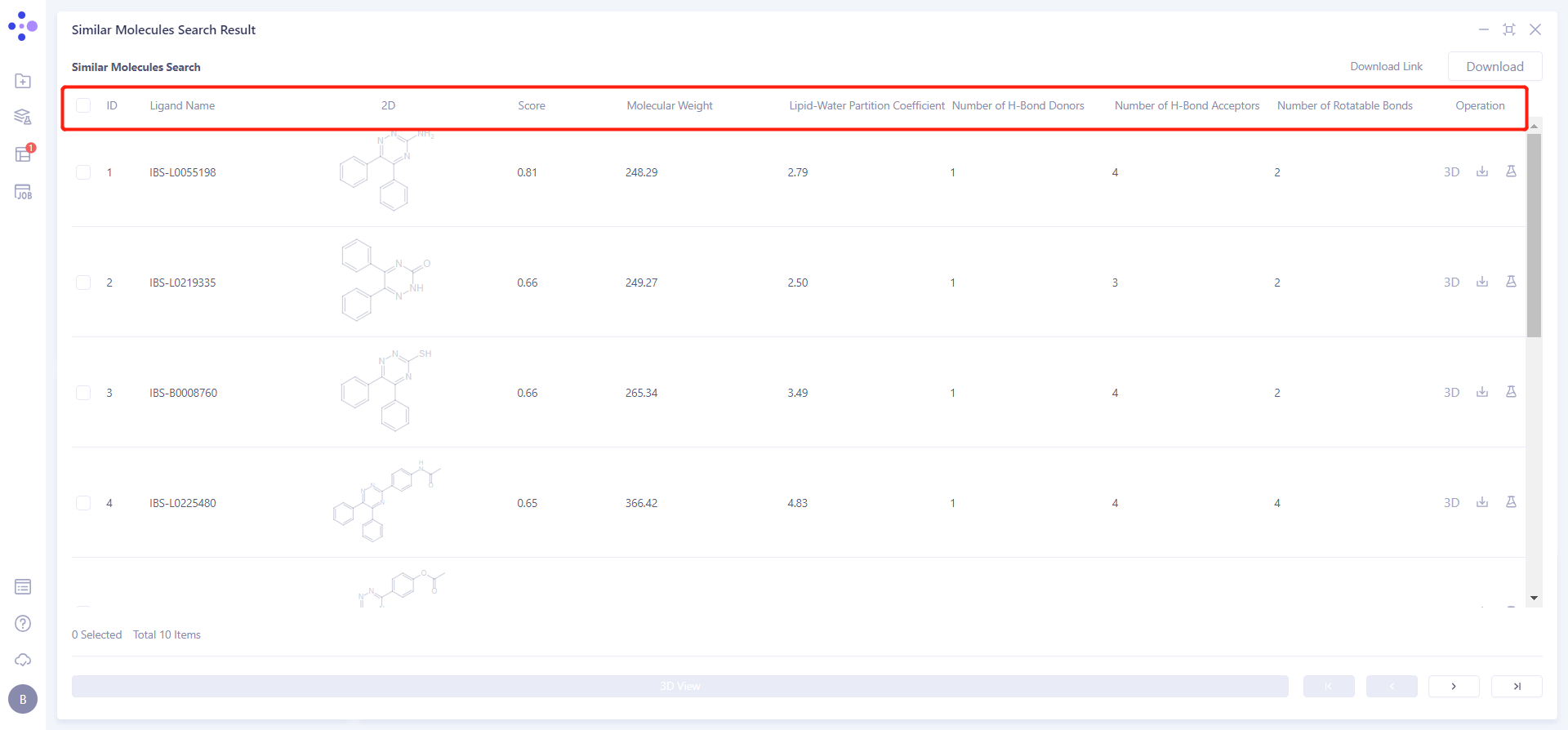
Click the symbol in the upper right corner to expand the result list and check the result information:
Ligand Name: Molecular name;
2D: Two-dimensional image of molecules;
Molecular Weight: Molecular weight;
Lipid-Water Partition Coefficient: Lipid-Water Partition Coefficient;
Number of H-Bond Donors: Determine the number of hydrogen bond donors;
Number of H-Bond Acceptors: Determine the number of H-Bond Acceptors;
Number of Rotatable Bonds: Determines the number of rotatable Bonds.
 |  |
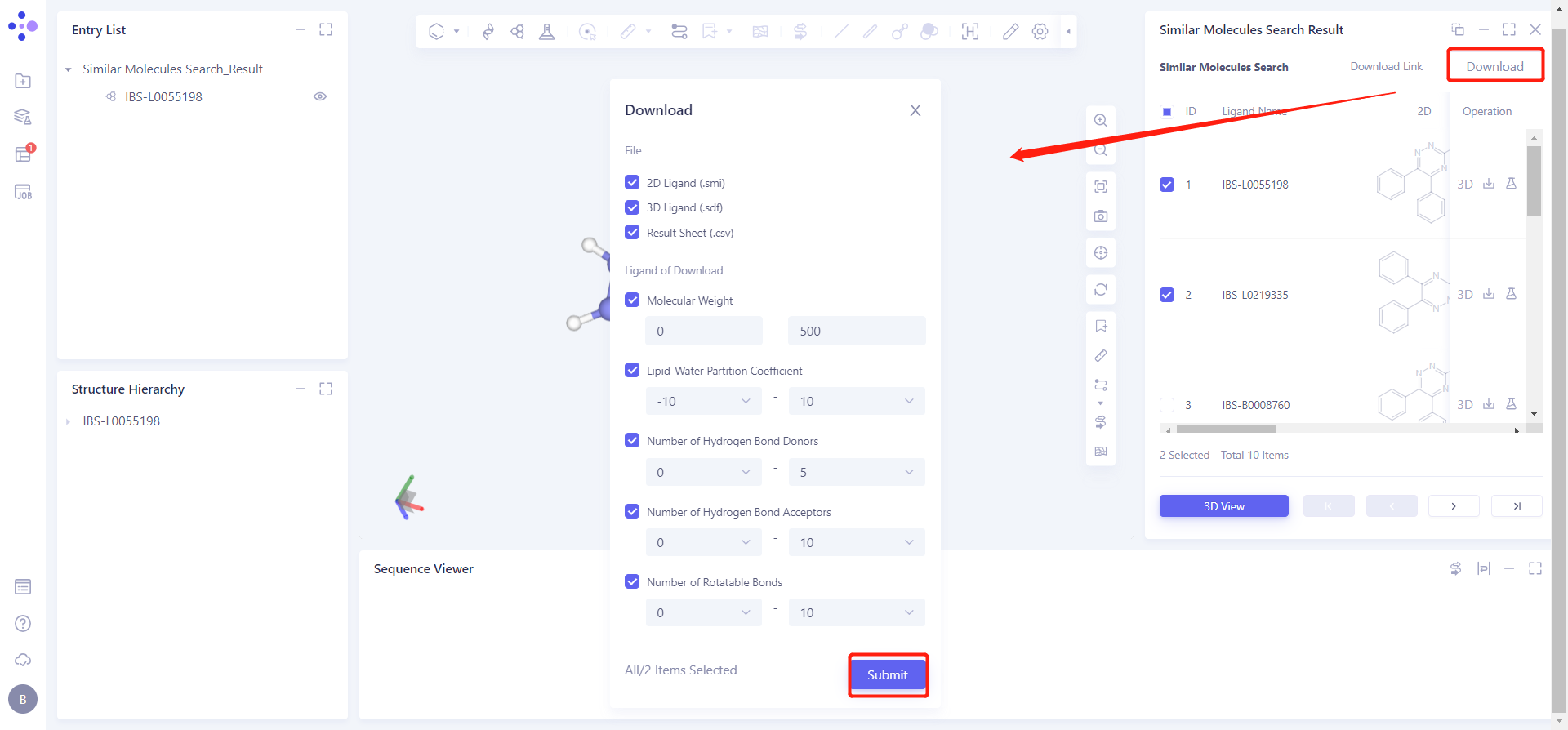
Display and download the required molecules based on the searched molecular structure and property information:
Check the IBS-L0055198 checkbox, click 3D View below, and the molecule will be displayed in the 3D Workspace window.
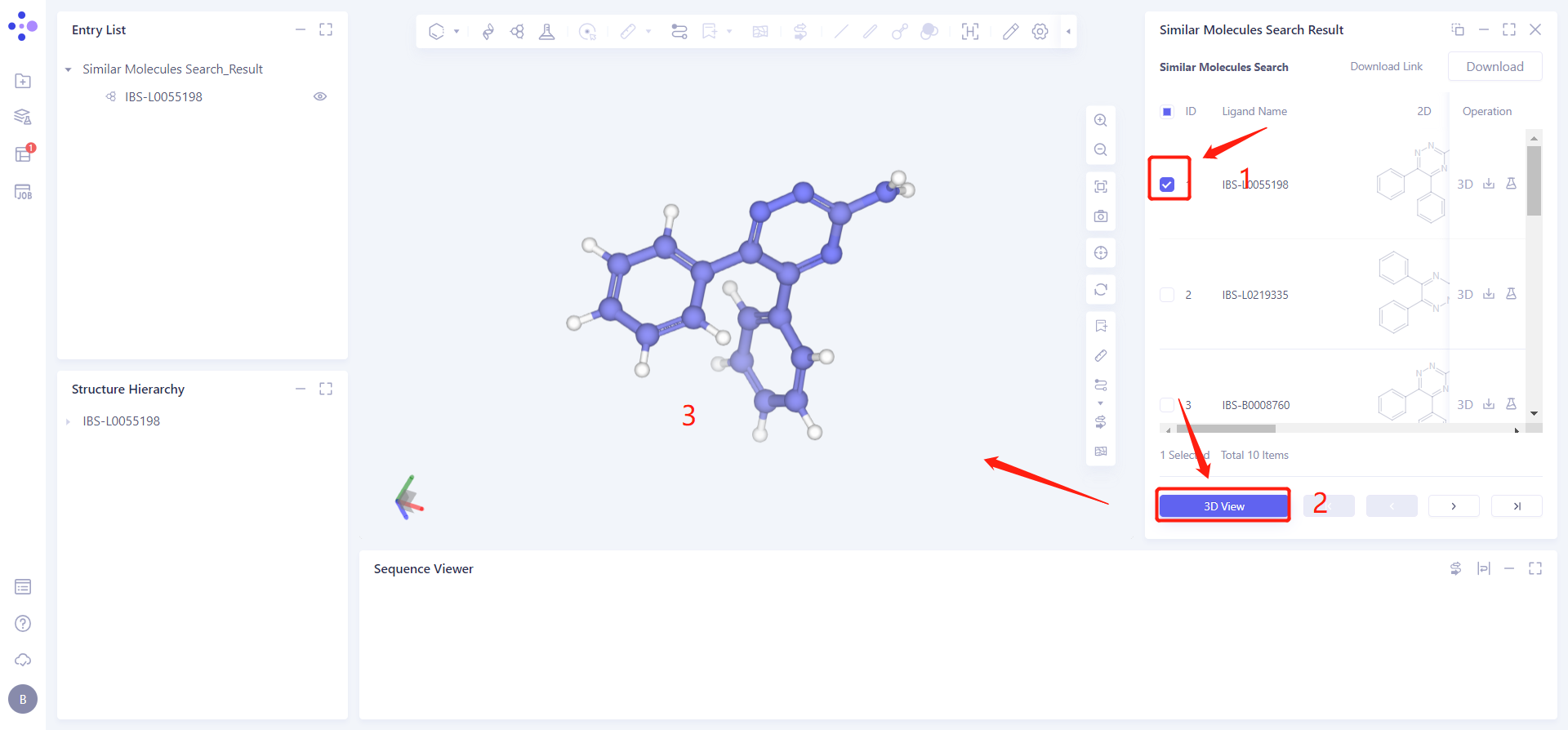
Click Download → pop-up Download interface in the upper right corner, select Filter information → click Submit.
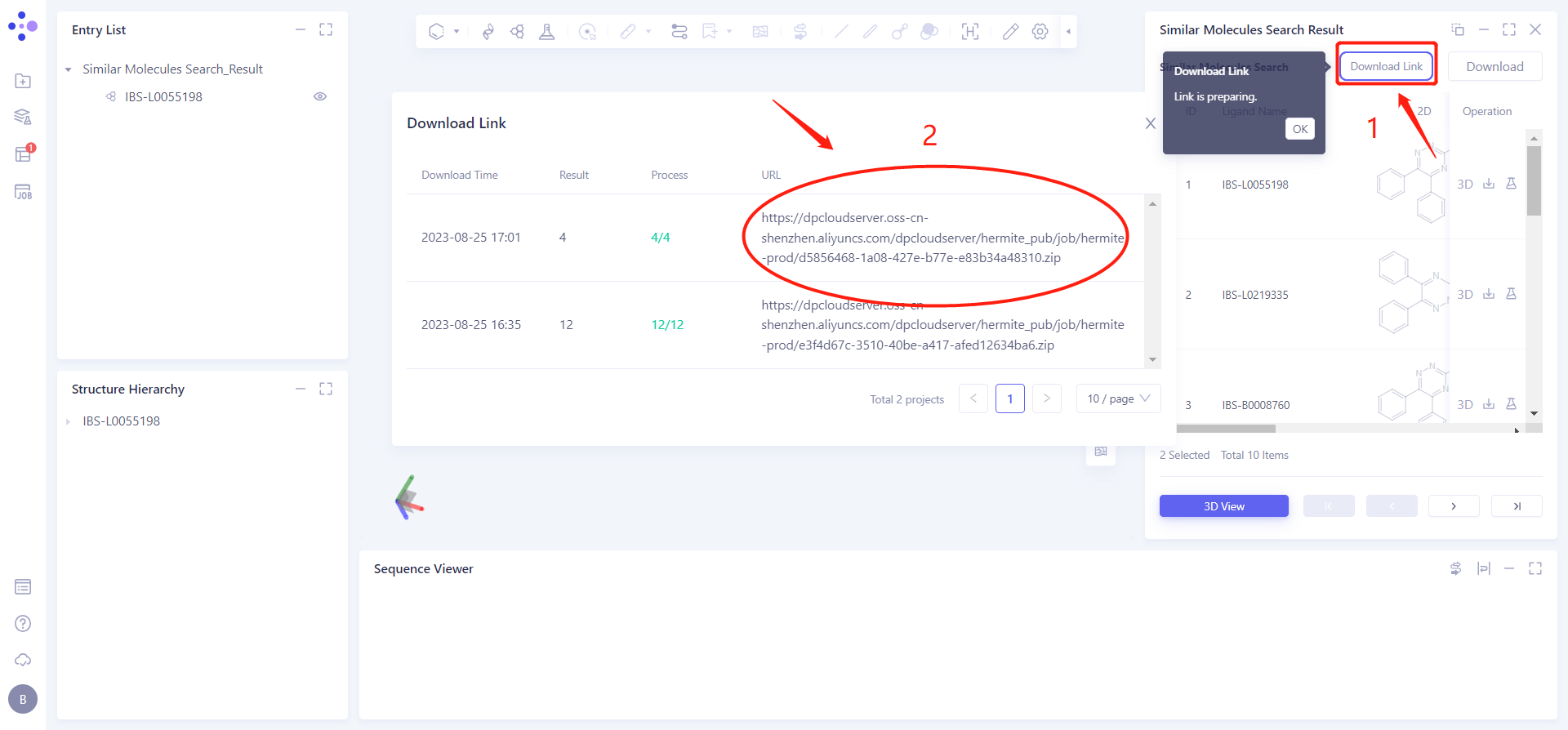
The Download Link window flashes, and the gray part on the left prompts "Link is preparing", click OK.
Click Download Link → pop-up Download Link interface → click the link under the URL to download the result data.