Uni-FEP (case of targeted exploration)
Introduction
Free Energy Perturbation (FEP) [1] is currently recognized as the most accurate and best mobility affinity calculation method, which can accurately evaluate the receptor-ligand binding free energy. For example, calculating the relative binding free energy of ligands A and B , while and are expensive to calculate. Since free energy is a state function, the thermodynamic cycle can be designed to calculate instead. Therefore, the relative binding free energy [2].
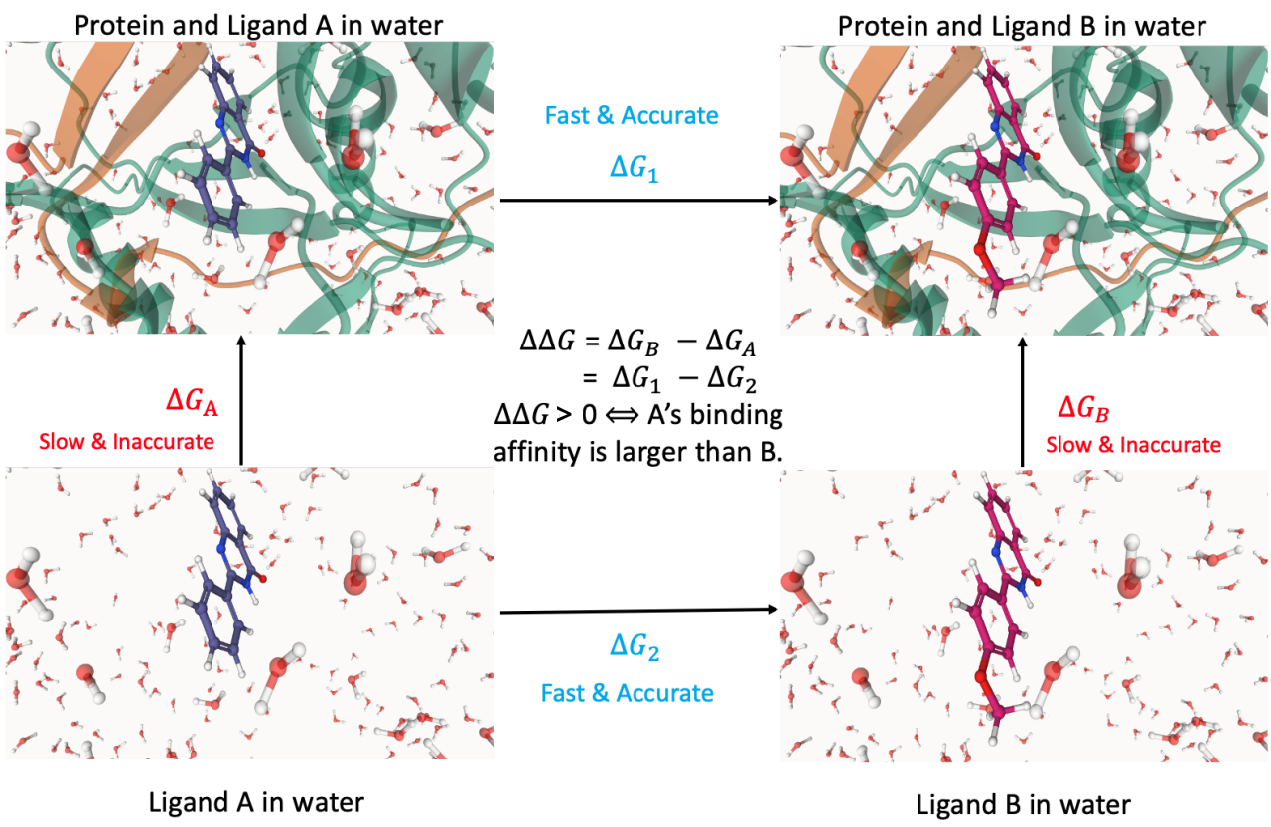
Uni-FEP is a complete FEP solution developed by Deeppotential Technology. It combines free energy perturbation theory, molecular dynamics, enhanced sampling algorithm and high-performance computing to efficiently evaluate the binding of receptors and ligands with chemical precision. Affinity, suitable for lead compound optimization stage. Uni-FEP has always been committed to improving the accuracy and speed of calculation. In terms of accuracy, Uni-FEP uses automatic construction of atomic mapping algorithms, REST2 enhanced sampling algorithms, statistical methods to optimize the design and regression analysis of perturbation graphs, and has many simulation parameters tuned for soft key interactions; In terms of speed, through systematic GPU high-performance optimization, a single FEP calculation under recommended parameters can be completed within 10-15 hours, while the Hermite ® platform based on Cloud Service can perform hundreds of FEP calculations a day.
HNE
Human neutrophil elastase (HNE) is an important class of proteolytic enzymes present in neutrophil azure granules. It belongs to the chymotrypsin family and is similar to other common proteolytic enzymes, such as proteinase 3 (Proteinase 3, P3), cathepsin G (Cathepsin, G) have a greater degree of homology. The main physiological functions of HNE in the human body are to resist and remove foreign pathogens that invade the human body. In addition, HNE also plays an important role in many inflammatory response diseases. Studies have shown that abnormal secretion of HNE is closely related to the occurrence of cardiopulmonary diseases such as chronic obstructive pulmonary disease, acute lung injury, cystic fibrosis, bronchiectasis, and pulmonary hypertension. There are corresponding negative regulatory proteins of HNE in the human body, such as serine protease inhibitors, α2 globulin and secretory leukocyte protease inhibitors. Such negative regulatory proteins are crucial to maintaining the normal function of HNE, and the balance between negative regulatory proteins and proteolytic enzymes such as HNE is broken, and various diseases as described above occur. Therefore, the development of effective HNE inhibitors to inhibit over-secreted HNE is expected to become a new strategy for the treatment of the above diseases. At present, there is only one drug on the market around this target, and several drugs are in different stages of clinical research. In this case, the paper [3] of Bayer Company disclosing the development and optimization process of BAY 85-8501 lead compounds was selected, and the more representative inhibitor molecules 18, 20, 22, 25, 27, 29 (BAY 85-8501) were compared. FEP simulation was carried out. The selected crystal structure PDB is 5A0A.
1. Create a project
1.1 Login system
Login address: https://hermite.dp.tech

1.2 Create a project
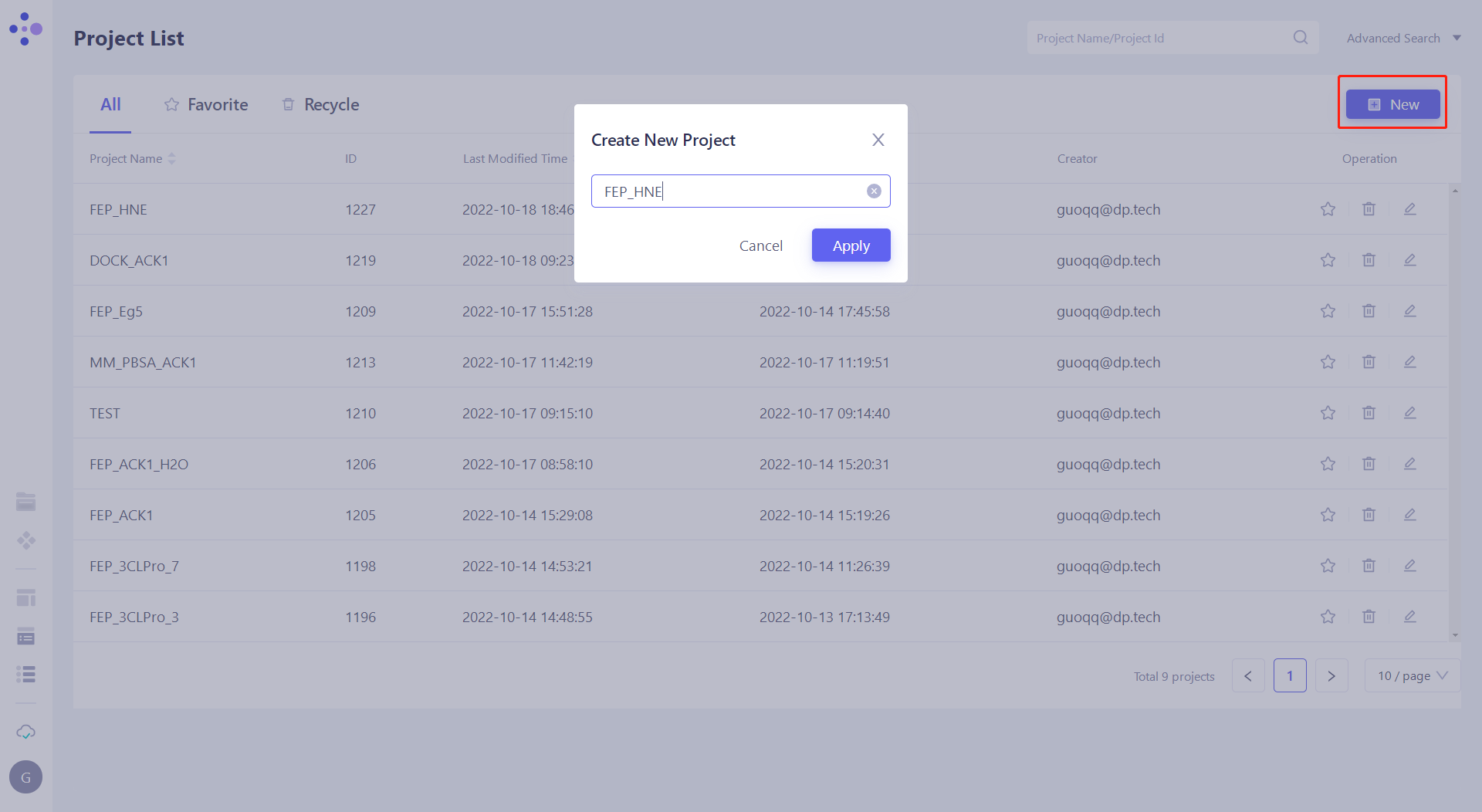
2. Import and prepare the system
2.1 Import structure
| Operation | Display |
|---|---|
| Click File → Get PDB, enter the PDB ID 5A0A | 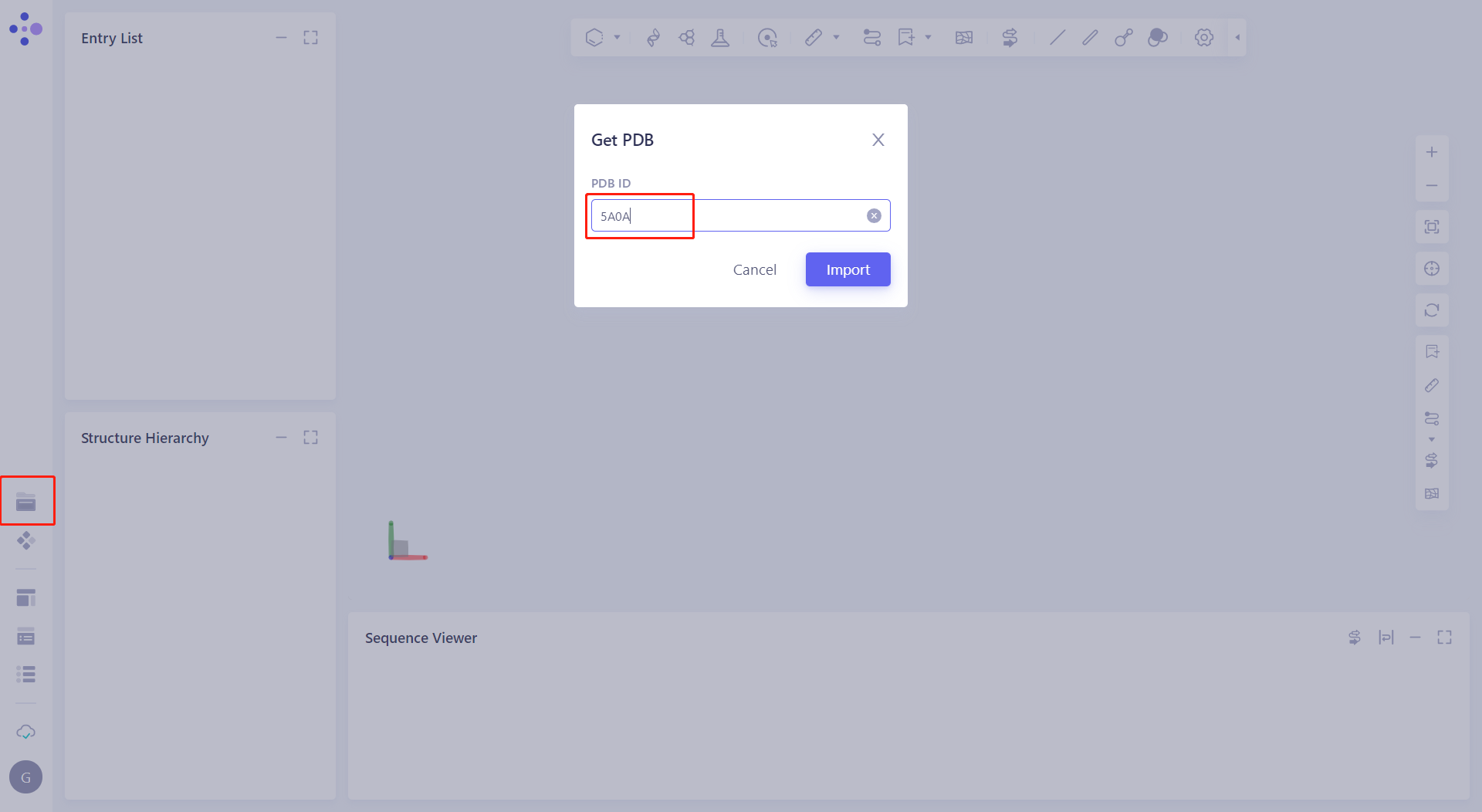 |
2.2 Manual preparation work
| Operation | Display |
|---|---|
| 2.2.1 Manually handle the protein In the Structure Hierarchy window, select the Ligand hierarchy, E NAG 1247 and E EPE 1250, right-click the mouse and select "Delete"; Select all water (except 2044) under the Solvents hierarchy, right-click the mouse and select "Delete"; Select the Others hierarchy, right-click the mouse and select "Delete". | 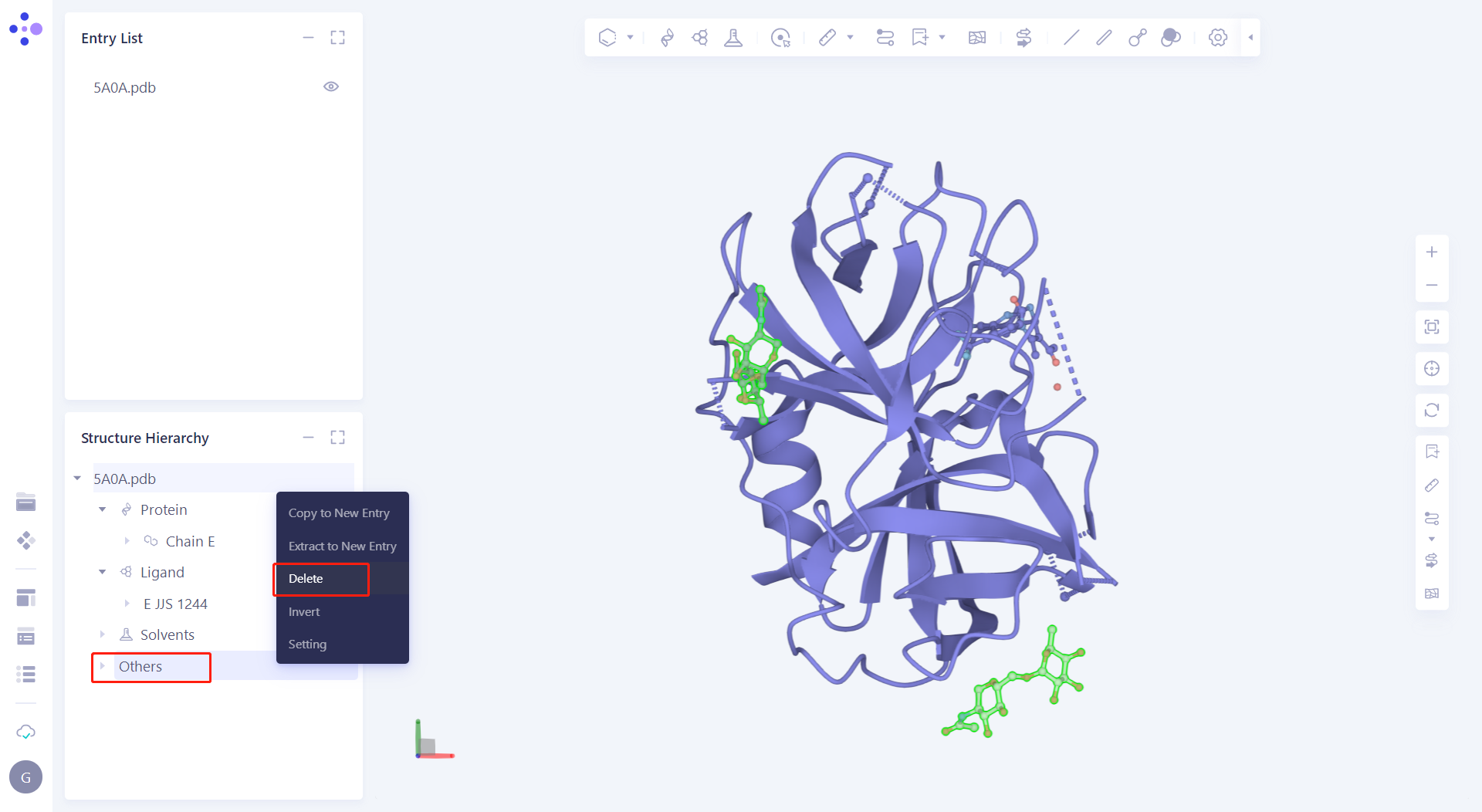 |
| 2.2.2 Manual extraction of eutectic ligand Put eutectic ligand E JJS 1244 "Extract to New Entry" and rename it to "Lig_JJS". | 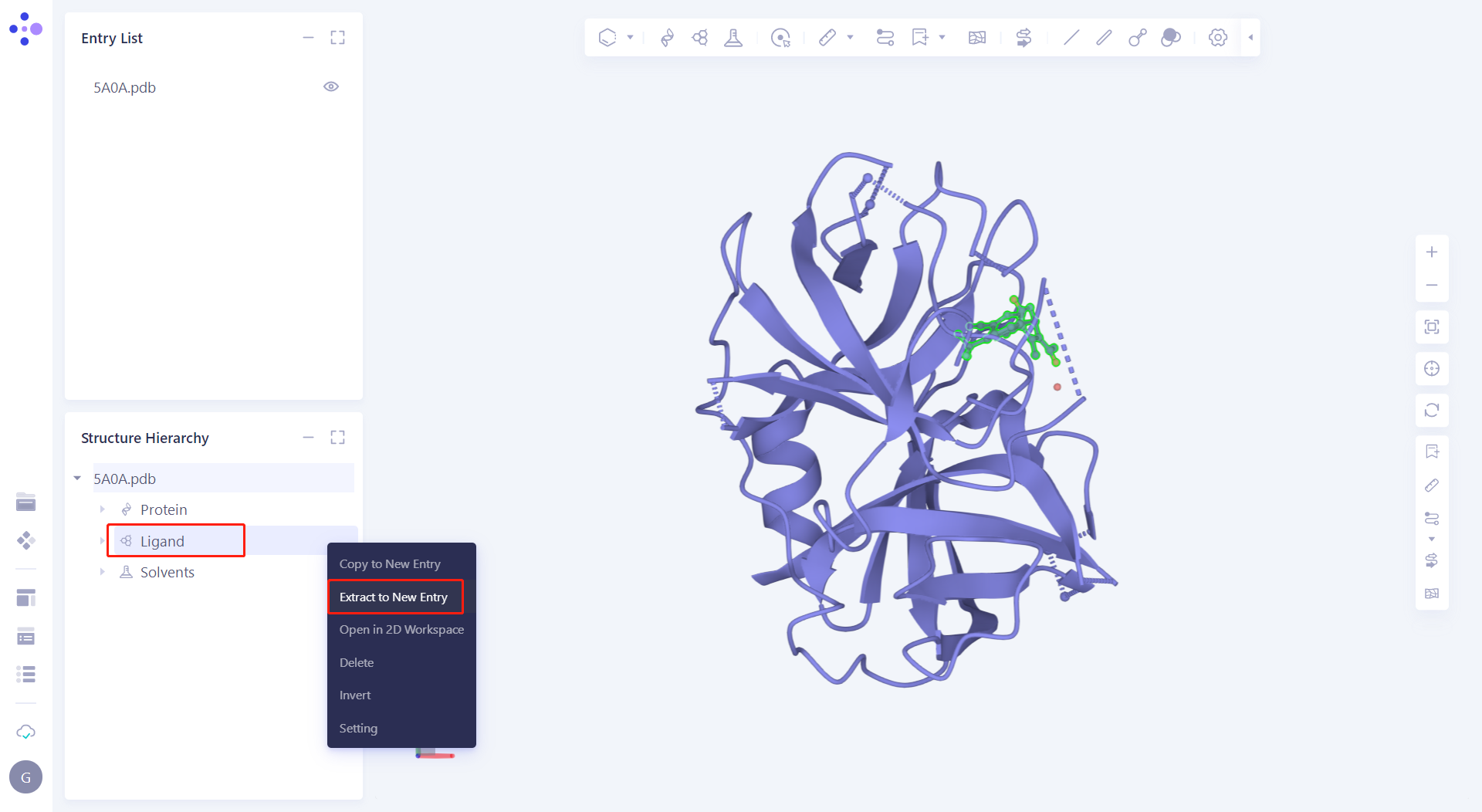 |
2.3 Open the FEP function interface
| Operation | Display |
|---|---|
| Click Function → Binding Affinity Evaluation → FEP Calculation | 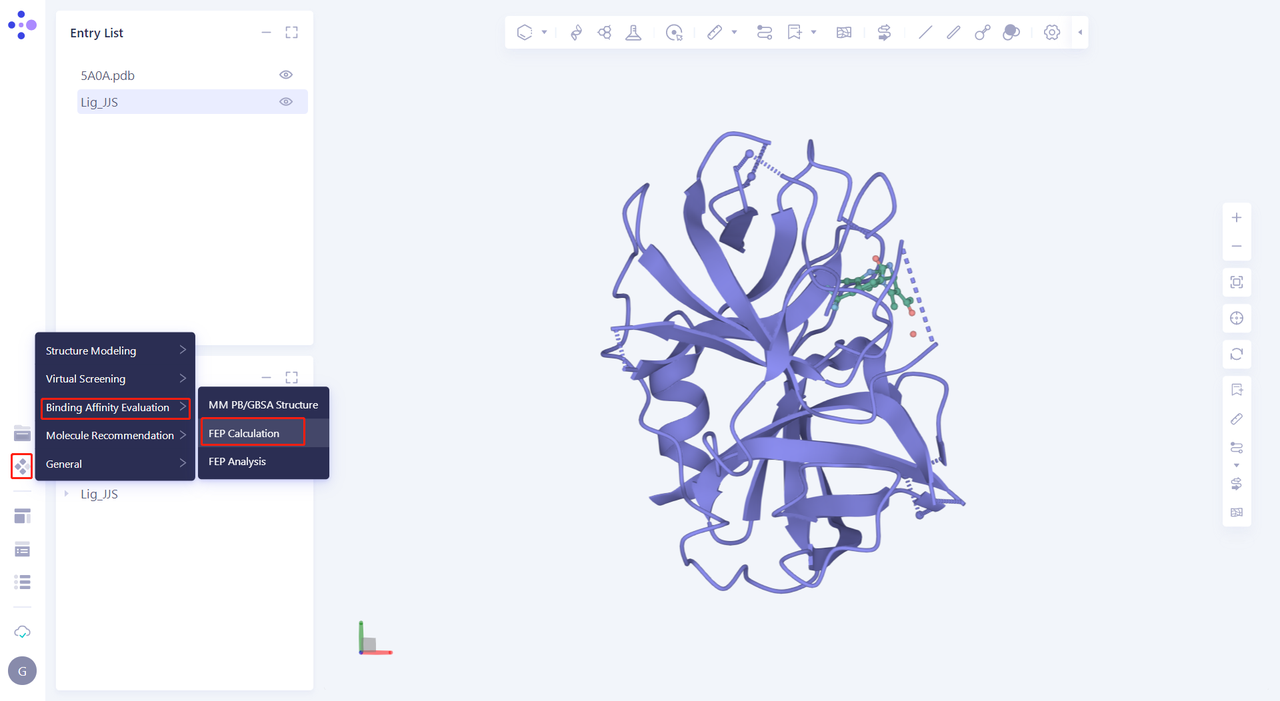 |
2.4 Processing protein
| Operation | Display |
|---|---|
| 2.4.1 Import the protein calculated by the user's FEP Click "From 3D", select "5A0A.pdb", and then click "OK" to upload the protein to the FEP function. | 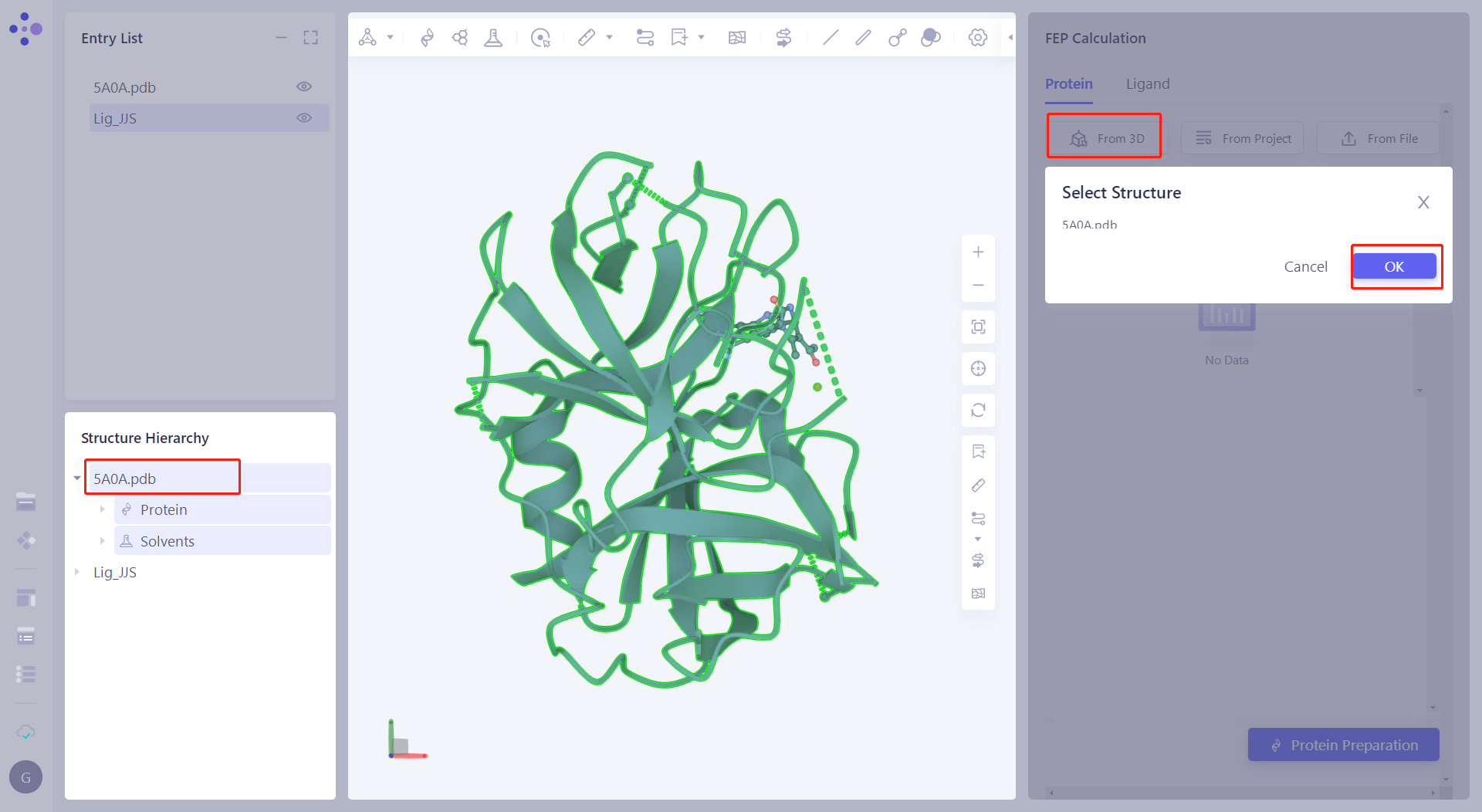 |
| 2.4.2 Automatic processing of proteins Click "Protein Preparation" to treat proteins, keep the "E-chain" in "Select Polymer to Keep" and "Select Other Groups to Keep", and most click "Next" to submit the task. | 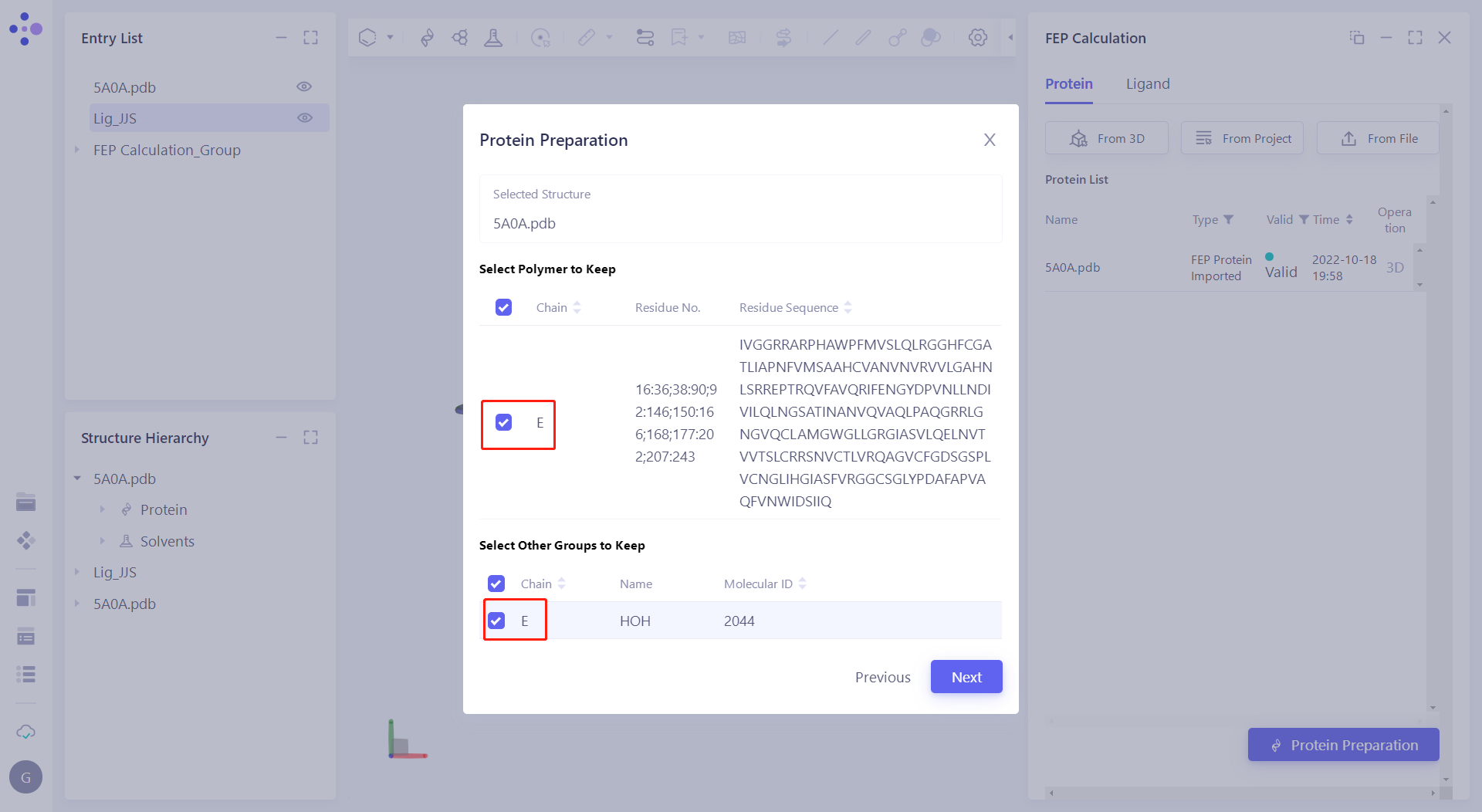 |
| In the "Protein Preparation" parameter settings, submit tasks according to the default parameters. | 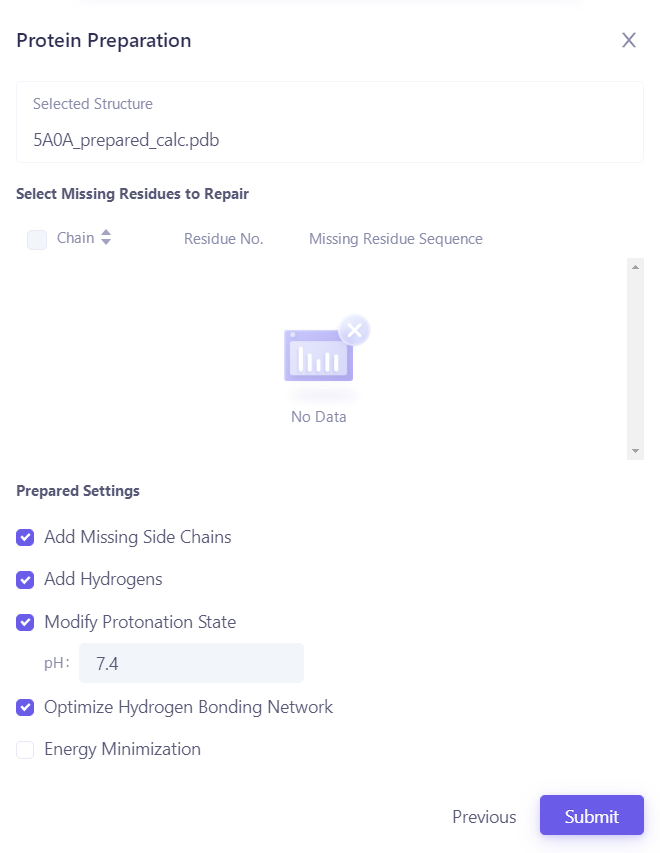 |
2.5 Molecular superposition
| Operation | Display |
|---|---|
| 2.5.1 Uploading Small Molecules In the Ligand module, click "From File" to upload the small molecule structure. | 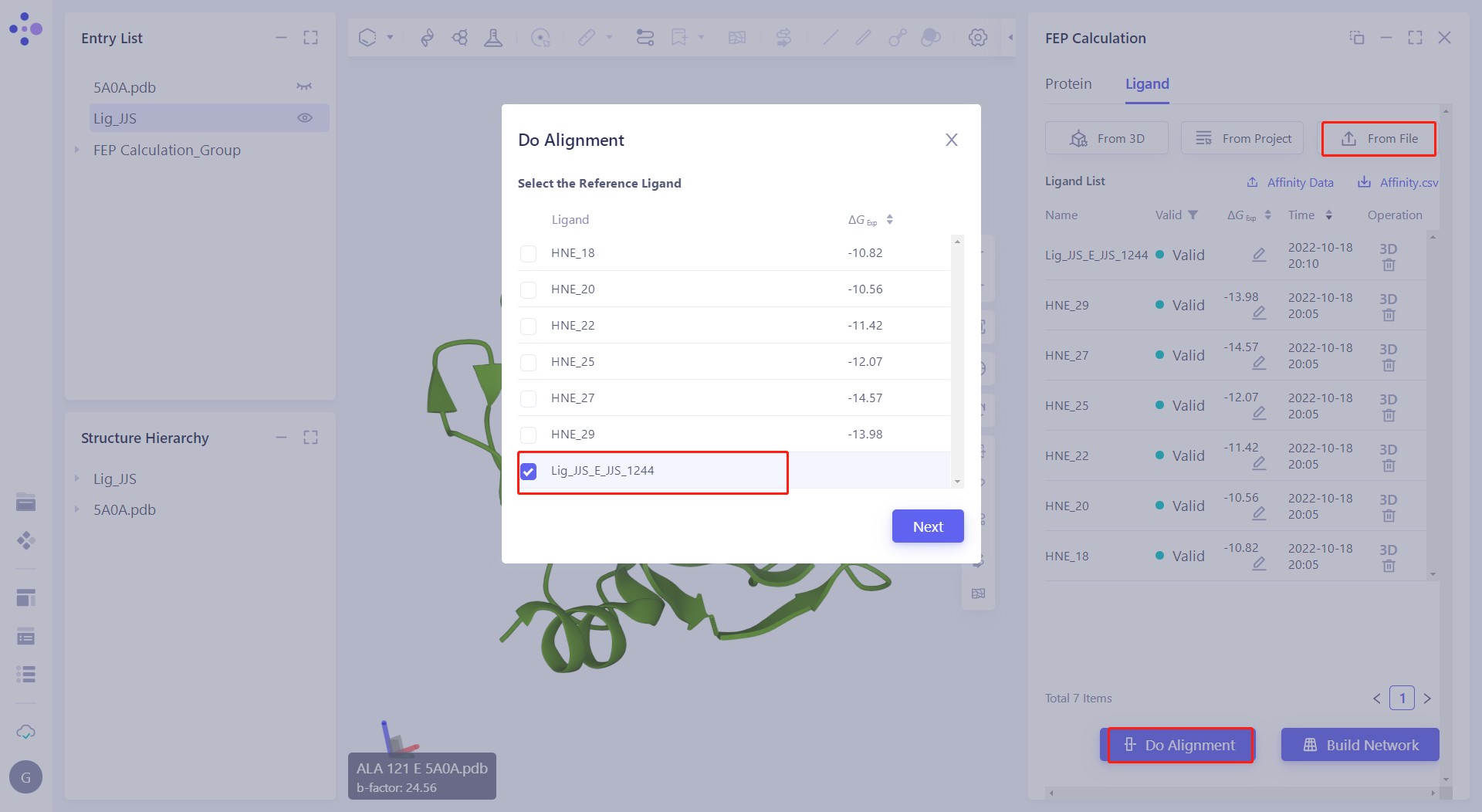 |
| 2.5.2 Select and Align Reference Molecules Click "Do Alignment" to do molecular overlap on the molecules. Check "Lig_JJS_EJJS_1244" as reference. Then check all molecules in Select Alignment Ligands, and select "Rigid Alignment" in the alignment method to align all molecules with reference. < br/> Note: After the molecules are aligned, delete the Lig_JJS_EJJS_1244. | 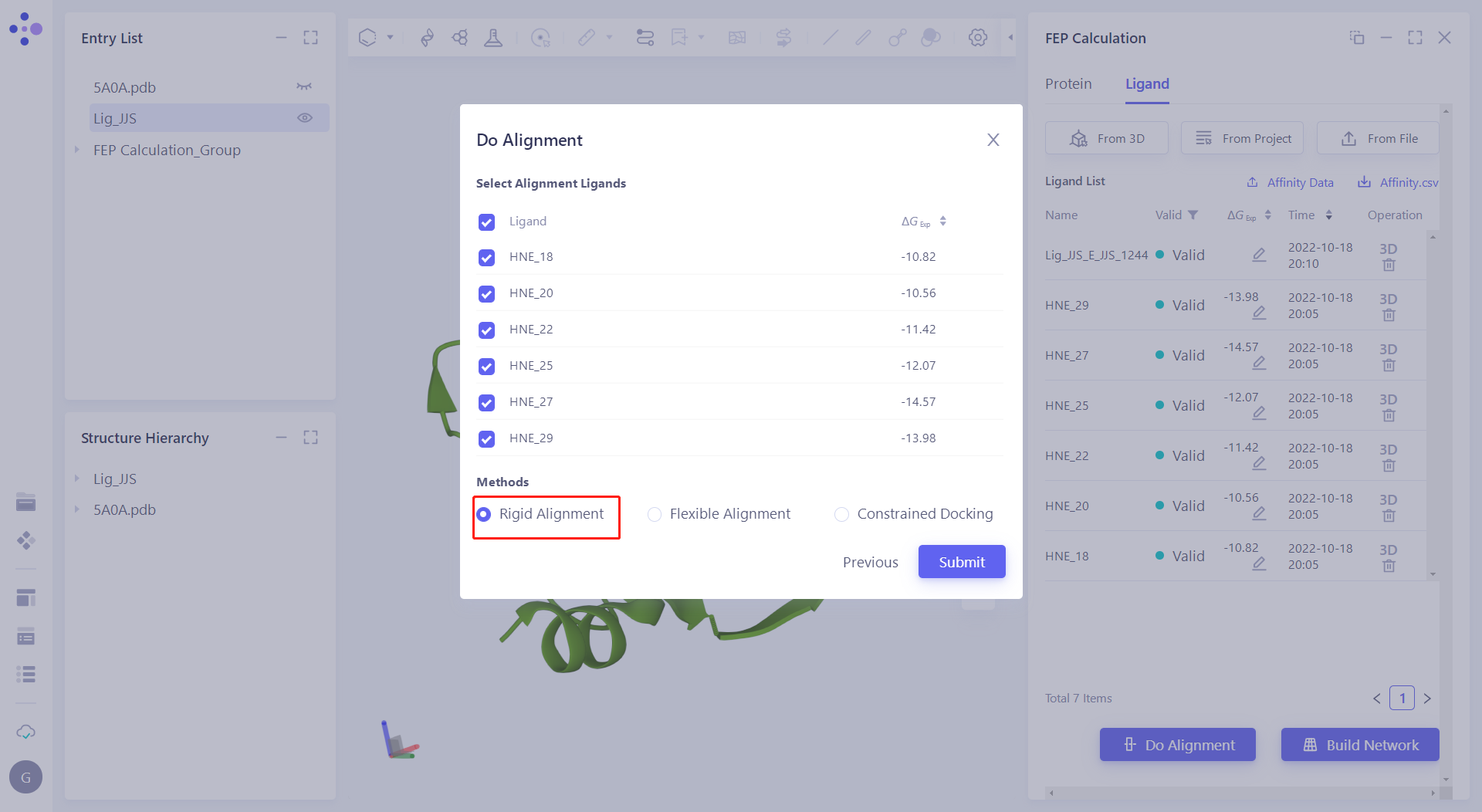 |
2.6 Perturbation Graph Construction and Mapping Checking
| Operation | Display |
|---|---|
| 2.6.1 Construction of the perturbation graph Click "Build Network" and submit the task according to the default parameters. | 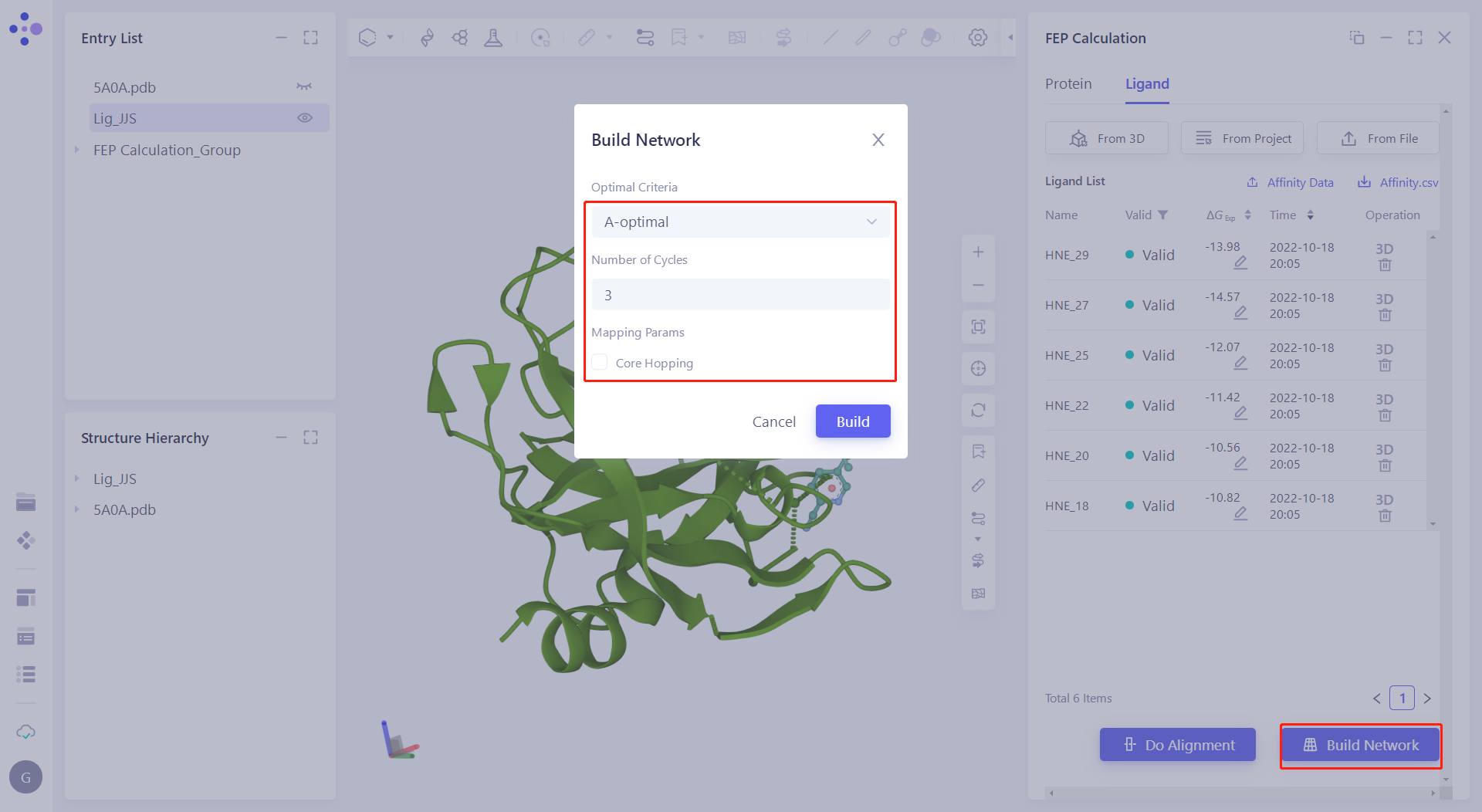 |
| 2.6.2 Perturbation graph modification and Mapping inspection After the task is completed, click "show" in "Perturbation" in the Job List to display the results of perturbation and mapping. | 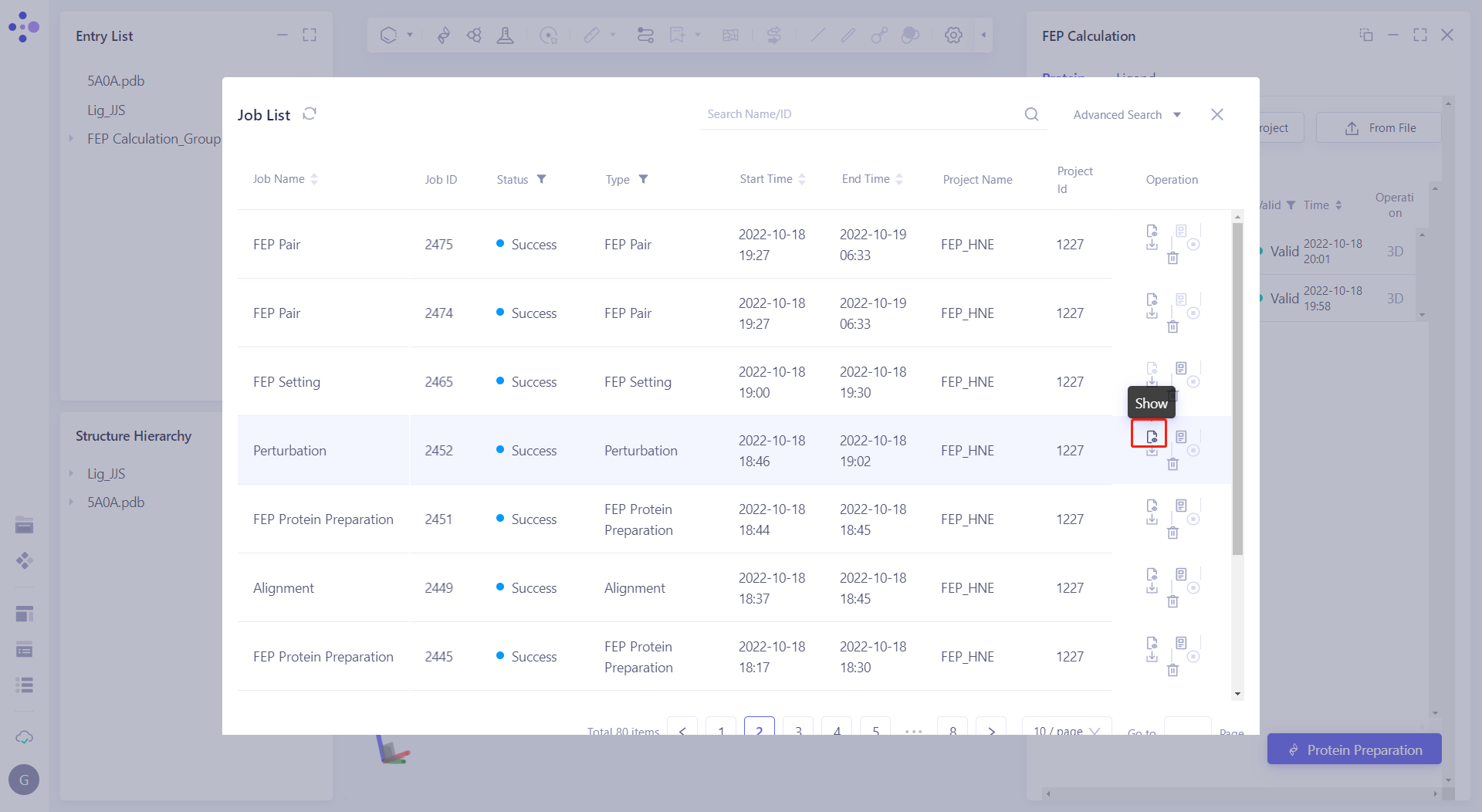 |
| Method 1 to modify the perturbation graph If you delete "HNE27 -> HNE18", first select the Edge between the two molecules, and click "-" on the right to delete the pairs. If you add "HNE27 -> HNE18" pairs, first select HNE27, then select HNE18, and then click "+" on the right to add new pairs. | 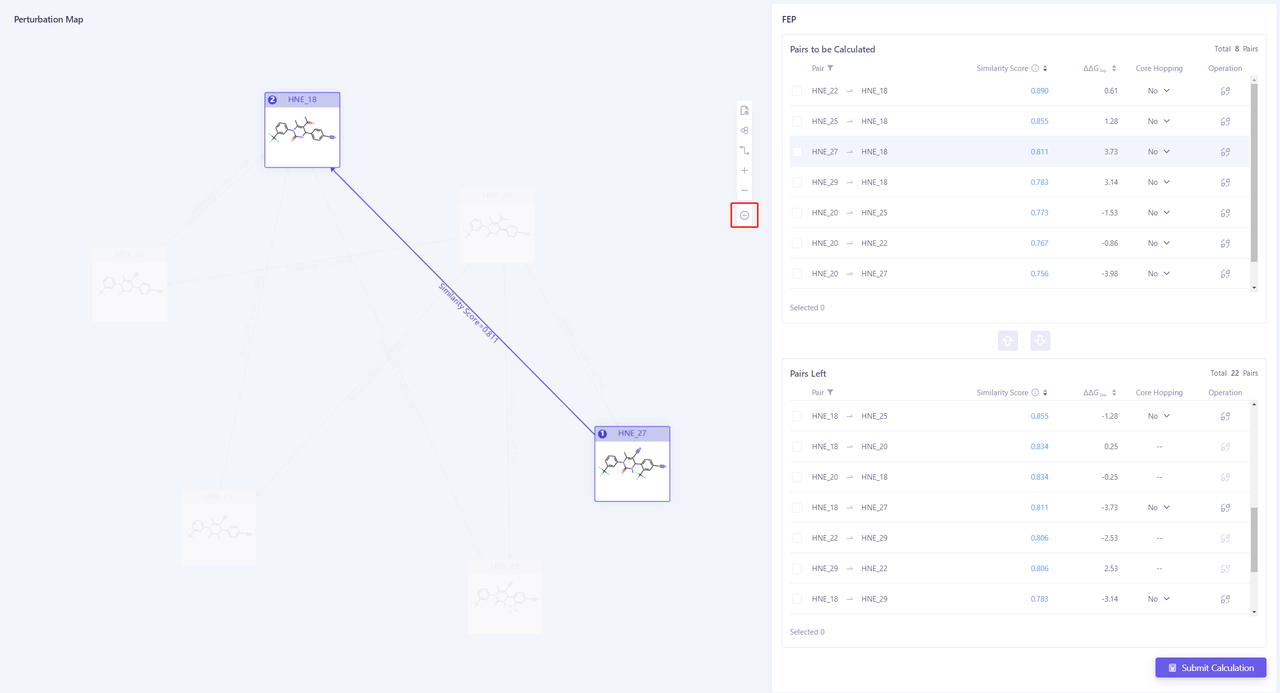 |
| Method 2 to modify the perturbation graph If you want to "add" or "delete" pairs, you can directly select the pairs in the table on the right, and then click the "up and down arrows" in the middle of the two tables to "add" or "delete". |  |
| Click "Mapping" in the upper right corner of the interface to check the mapping of each pair. | 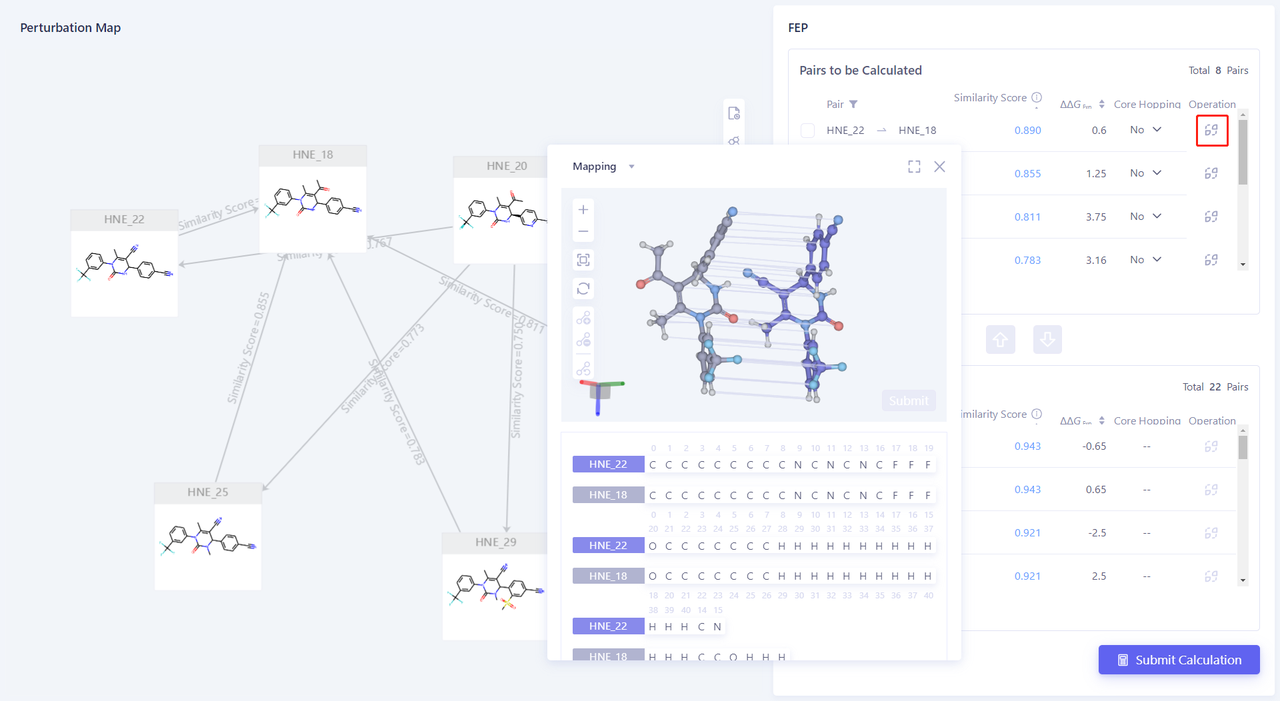 |
| To delete the mapping between two atoms, select the two atomic lines first, and click the "-" on the left to delete the mapping. If you add new mappings, you can select the two atoms separately, and then click the "+" on the left to add a new mapping. | 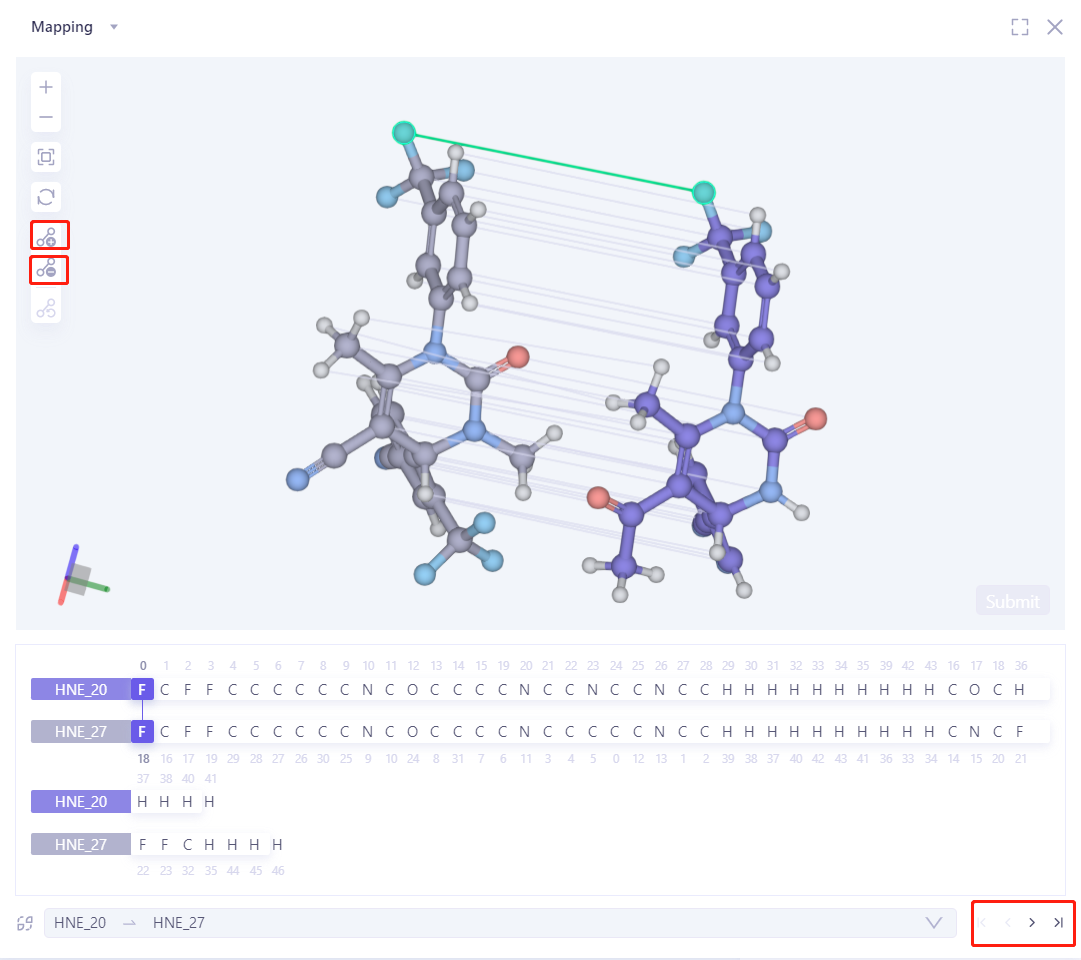 |
3. Task Submission
3.1 Determine the system used for calculation
| Operation | Display |
|---|---|
| 3.1.1 Determine the protein to be used for the FEP calculation Select the treated protein "5A0A_prepared" for the FEP calculation. |  |
| 3.1.2 Determine the Pairs to be calculated After confirming that the Pairs to be submitted are correct, click "Next". Tips: The operator has modified the perturbation graph before clicking Submit Task, so the pairs submitted for this task are different from the Pairs of the default perturbation graph. | 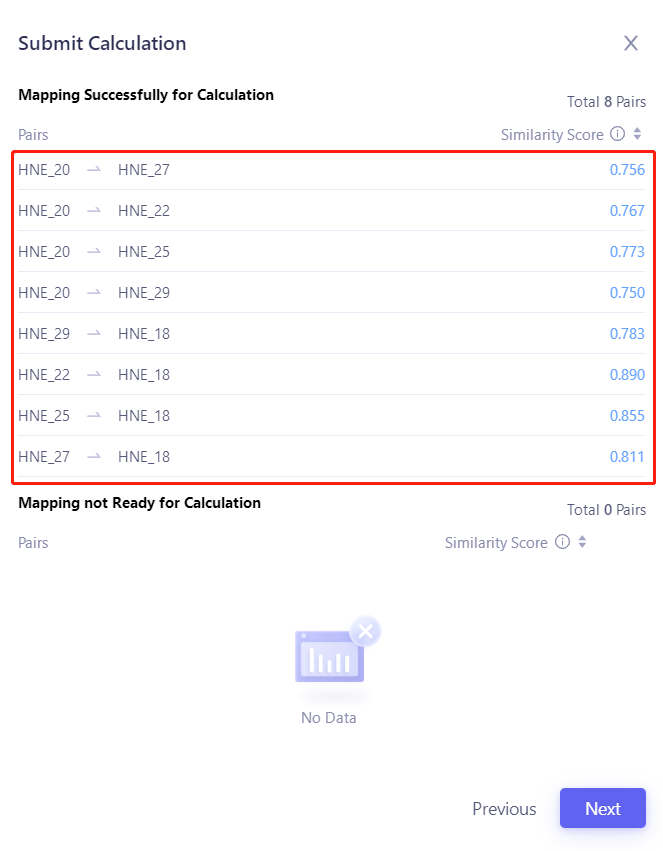 |
3.2 FEP parameter setting
| Operation | Display |
|---|---|
| After clicking "Next", the "Submit Calculation" parameter will pop up, and the simulation time will be modified in "Simulation Time". | 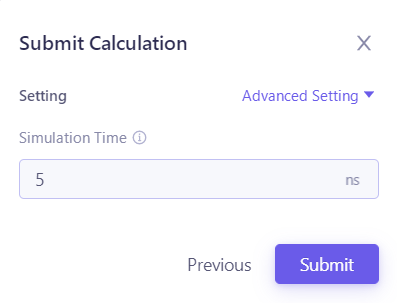 |
| Click "Advanced Settings" to view the calculation parameters of FEP , and finally click "Submit" to submit the task. In this case, the task is submitted according to the default parameters. | 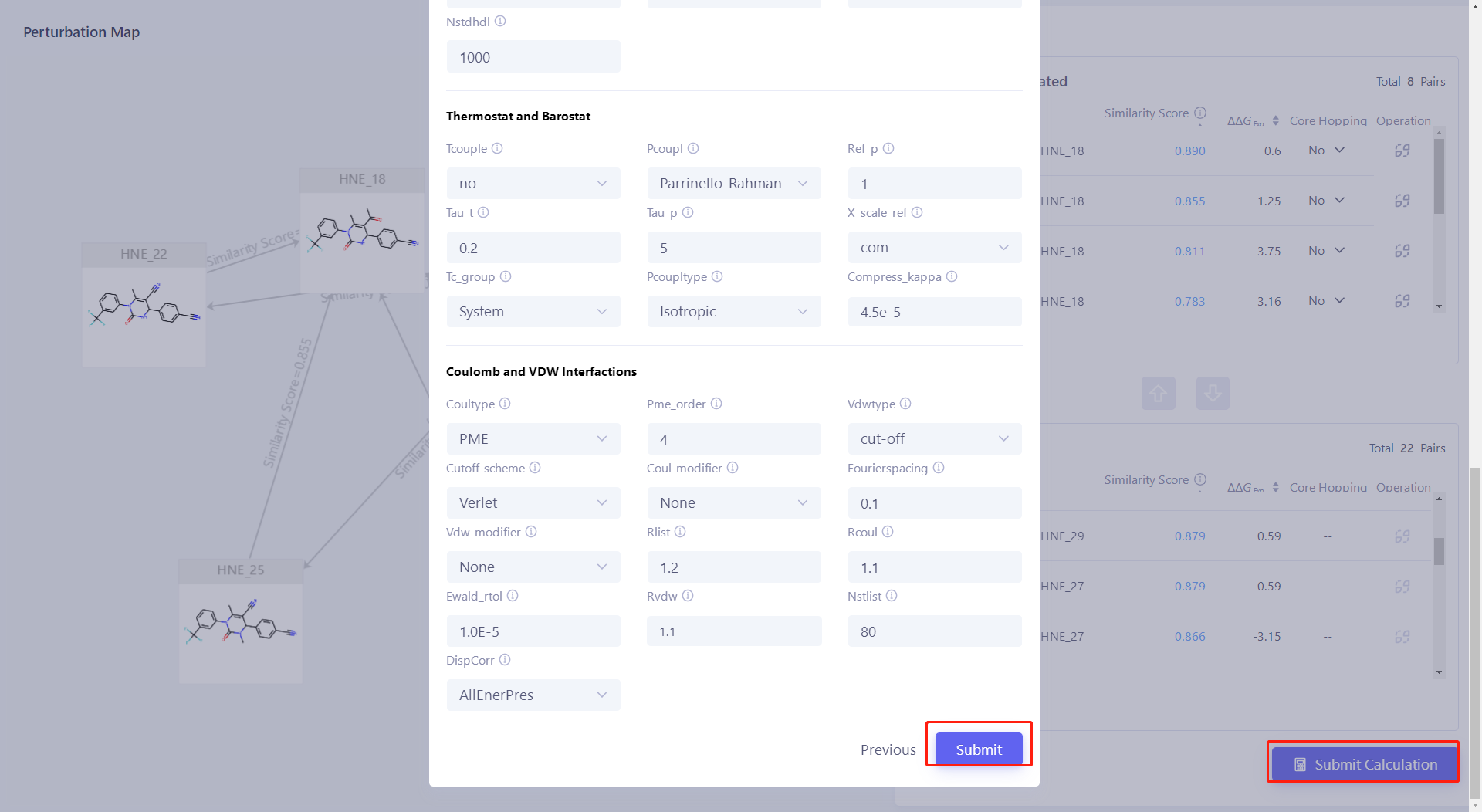 |
4. View FEP results
| Operation | Display |
|---|---|
| After the FEP task calculation is completed, click Function → Binding Affinity Evaluation → FEP Analysis | 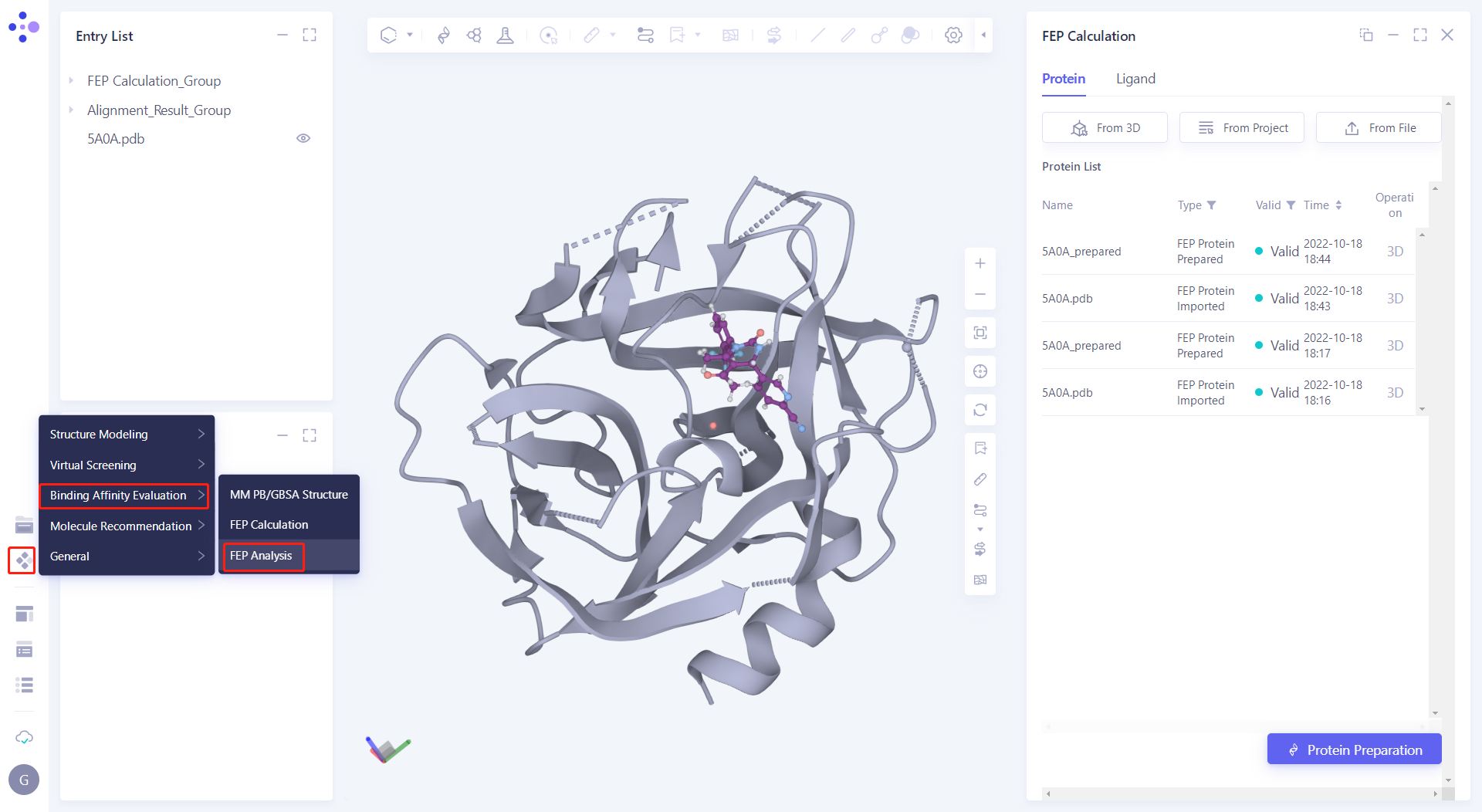 |
| Click "Analysis" in the lower right corner of the interface to submit the task. | 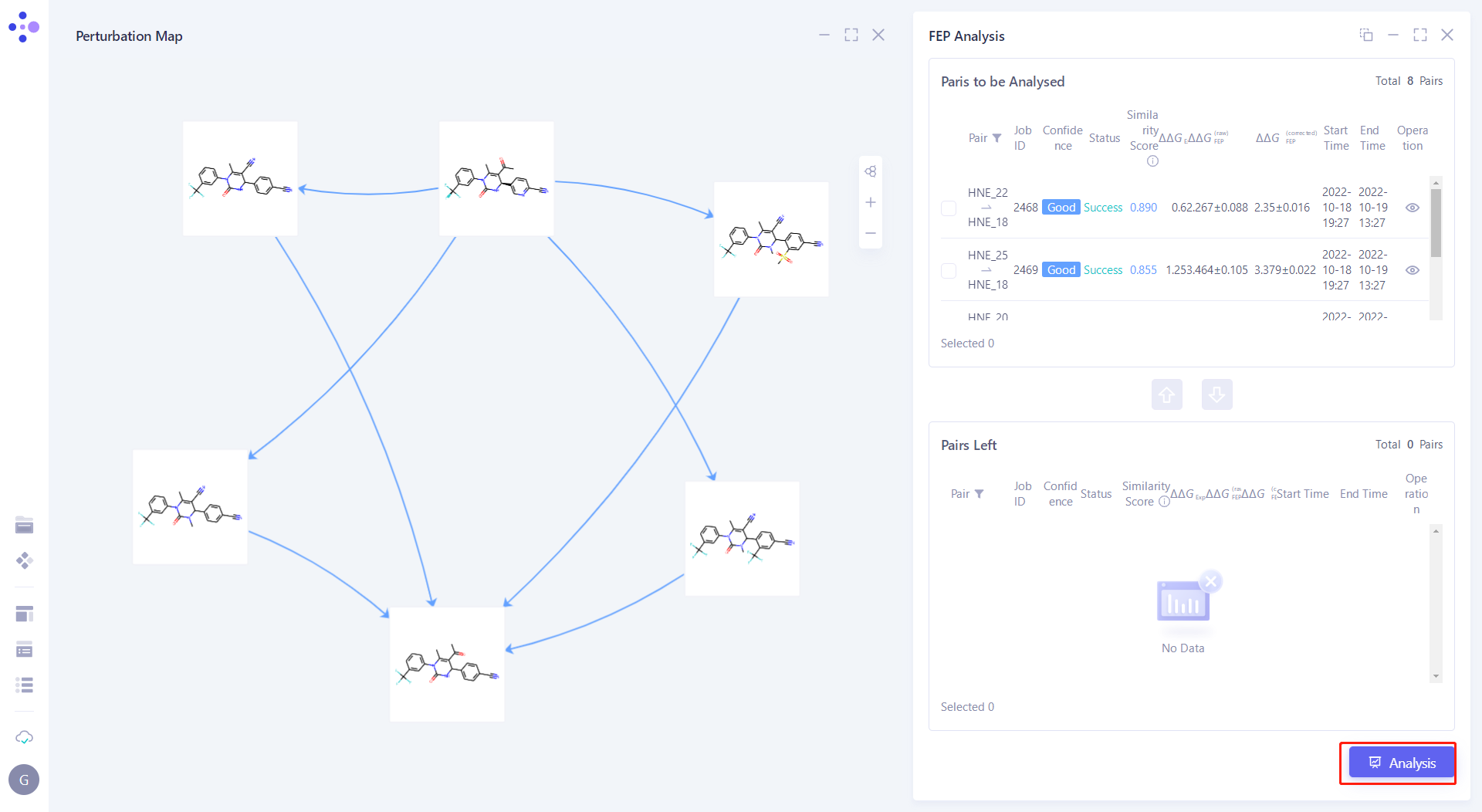 |
| After the FEP Analysis task calculation is completed, click "Show" in the "FEP Analysis" action sheet in the "Job List". | 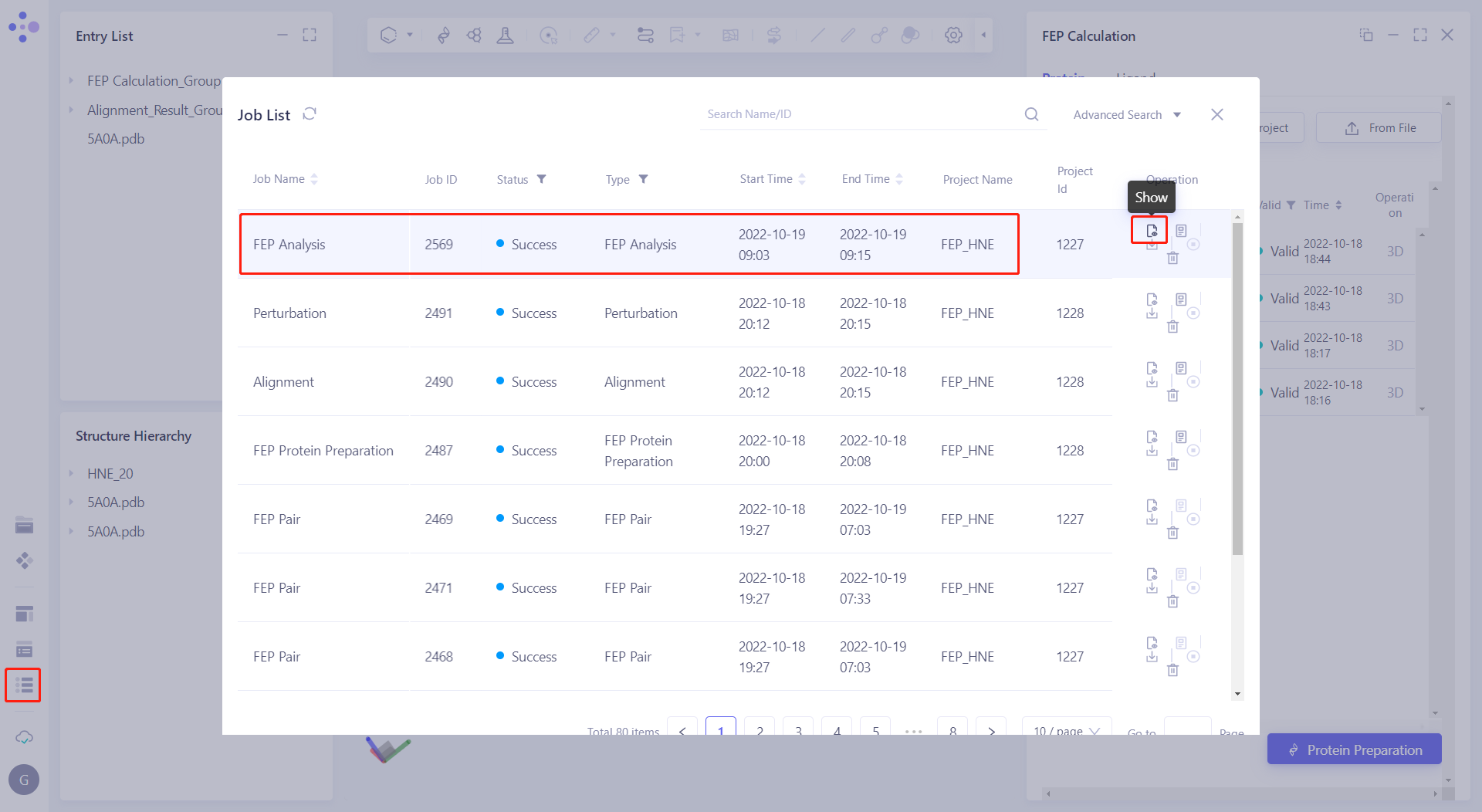 |
| Click "Node" to check " Ligand Name, ΔG EXP, ΔG FEP and σ (ΔG FEP) " to display the experimental affinity and calculated affinity. | 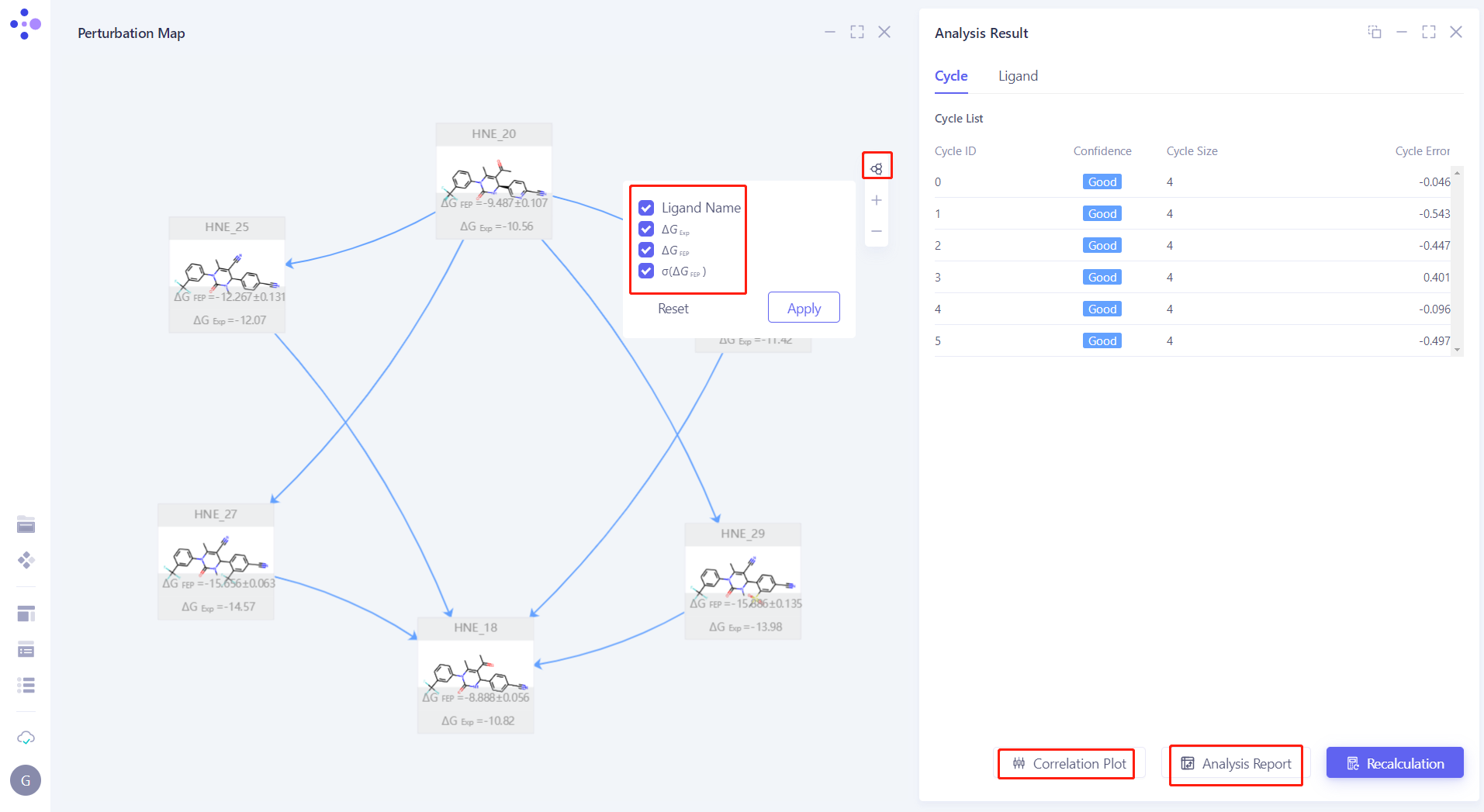 |
| Click "Correlation Plot" to analyze the experimental ΔG and calculate the correlation of ΔG, RMSE and R ² are 1.28kcal/mol and 0.96, respectively. | 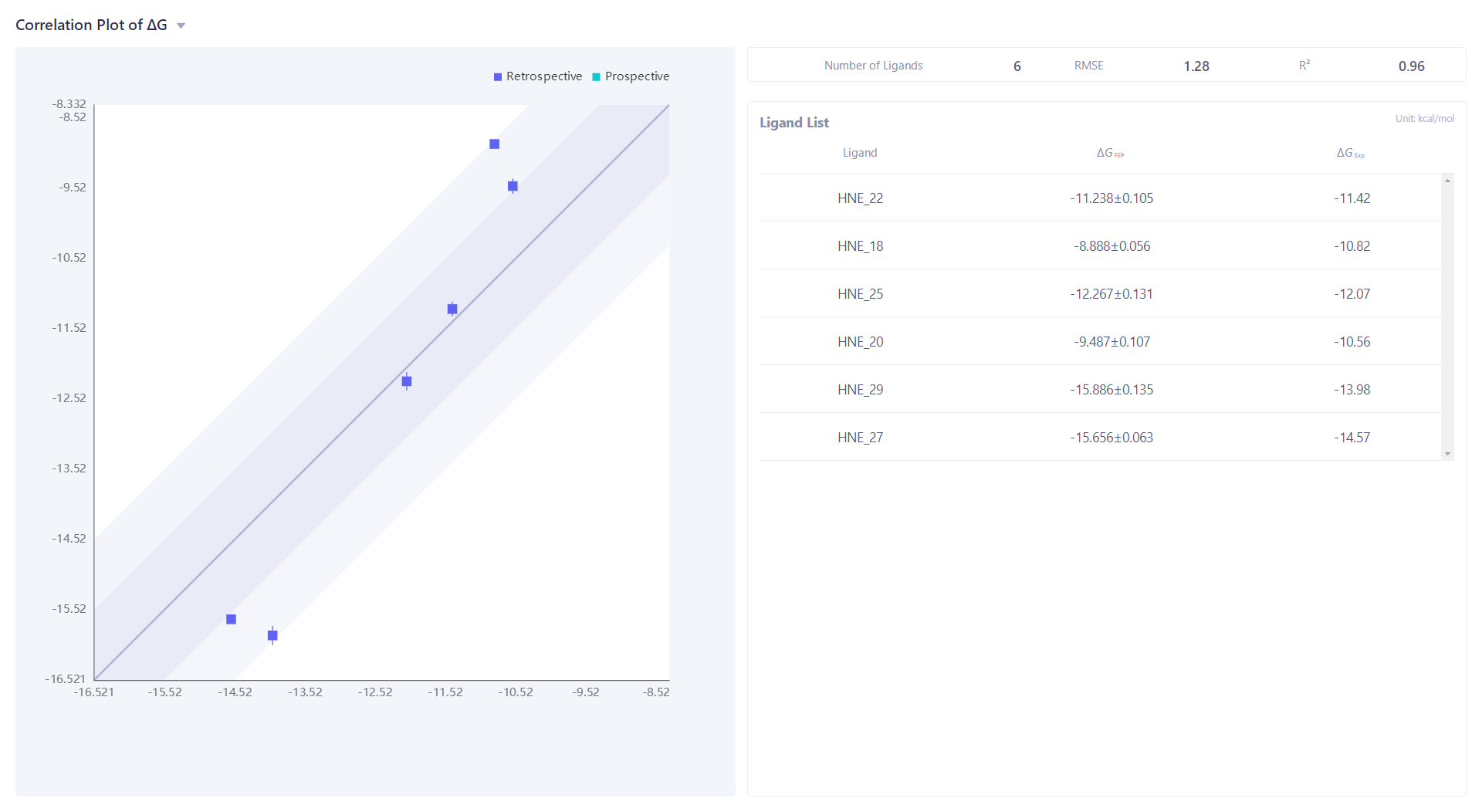 |
| Click "Analysis Report" to analyze the reliability of the calculation results. | 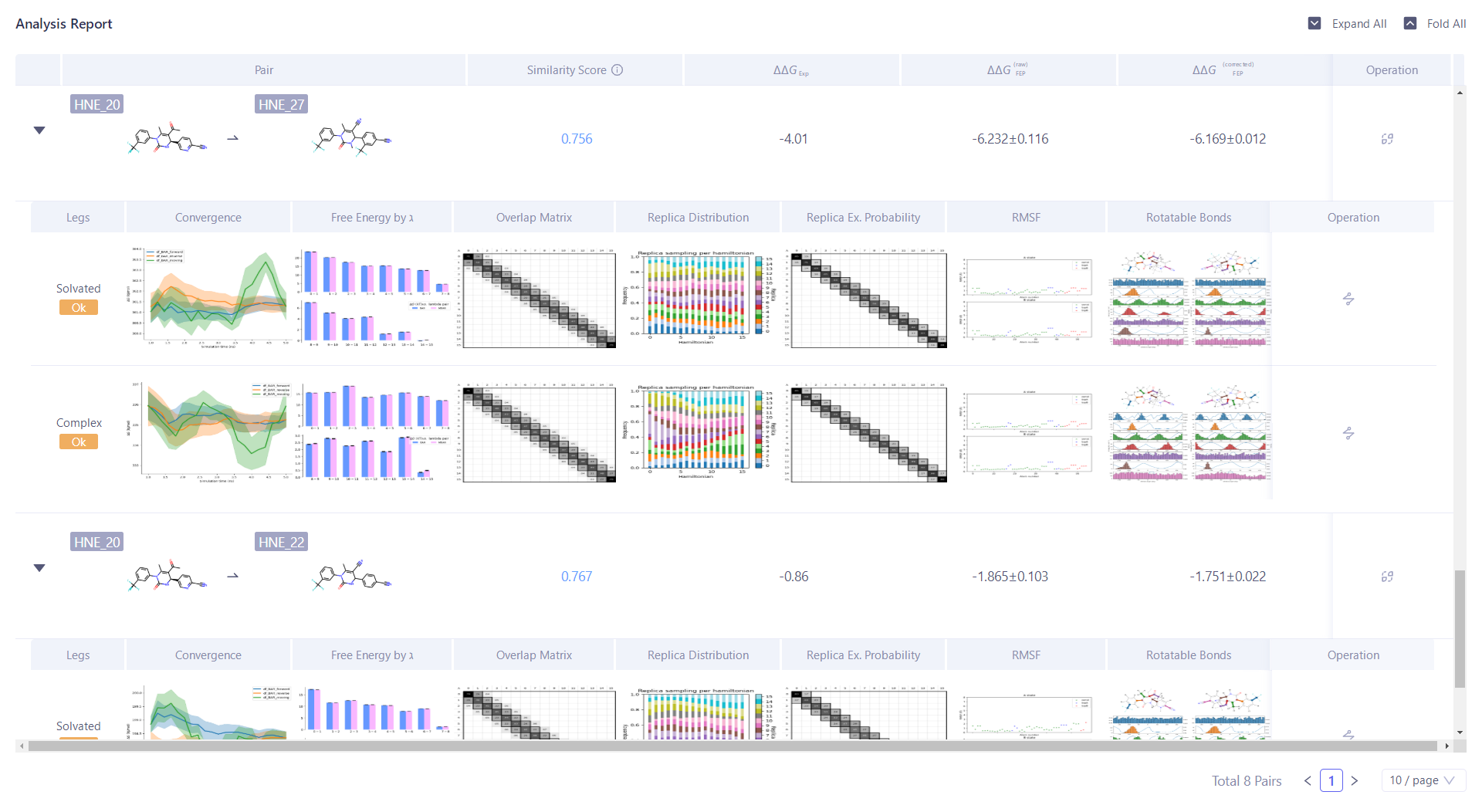 |
5. Summary
In this case, taking human neutrophil elastase (HNE) as an example, some key transition molecules in the optimization process of the clinical candidate molecule BAY 85-8501 and its lead compound developed by Bayer were carried out between small molecules and proteins. The binding energy is calculated. The results show that for a given target molecule, Uni-FEP can accurately evaluate the relative binding free energy between the compound and the target , thereby guiding the optimization of the lead compound.
6. Reference
[1] Chipot, Christophe, and Andrew Pohorille. "Free energy calculations." Springer series in chemical physics 86 (2007): 159-184.
[2] Cournia, Zoe, Bryce Allen, and Woody Sherman. "Relative binding free energy calculations in drug discovery: recent advances and practical considerations." Journal of chemical information and modeling 57.12 (2017): 2911-2937.
[3] Von Nussbaum, F.; Li, V. M. J.; Allerheiligen, S.; Anlauf, S.; Bärfacker, L.; Bechem, M.; Delbeck, M.; Fitzgerald, M. F.; Gerisch, M.; Gielen-Haertwig, H.; Haning, H.; Karthaus, D.; Lang, D.; Lustig, K.; Meibom, D.; Mittendorf, J. ; Rosentreter, U.; Schäfer, M.; Schäfer, S.; Schamberger, J.; Telan, L. A.; Tersteegen, A. Cover Pictures: Freezing the Bioactive Conformation to Boost Potency: The Identification of BAY 85-8501, a Selective and Potent Inhibitor of Human Neutrophil Elastase for Pulmonary Diseases. ChemMedChem 2015, 10 (7), 1117-1117.