VD Gen: a Novel Molecular Design Based on Protein Pocket
Introduction
The discovery and design of novel compounds with high affinity to target proteins is one of the core tasks of structure-based drug design (SBDD). Different from the traditional methods based on Virtual Screening, molecular de novo design (de novo Molecule Design) does not require the preparation of a huge virtual compound library in advance, but directly generates the molecular structures to be evaluated according to the requirements. The VD Gen module in the Hermite ® platform provides protein pocket-based molecular de novo design capabilities. Based on the Virtual Dynamics artificial intelligence algorithm independently developed by Deep Potential Technology, VD Gen can directly generate a new molecular 3D structure in the protein active pocket designated by the user, and evaluate the affinity between the molecule and the protein.
PDE10A is an important regulator of striatal signaling that, when inhibited, can normalize dysfunctional activity. This tutorial is based on the VD Gen module in the Hermite ® platform for early optimization of molecular fragments of PDE10A inhibitors.
1. Entrance
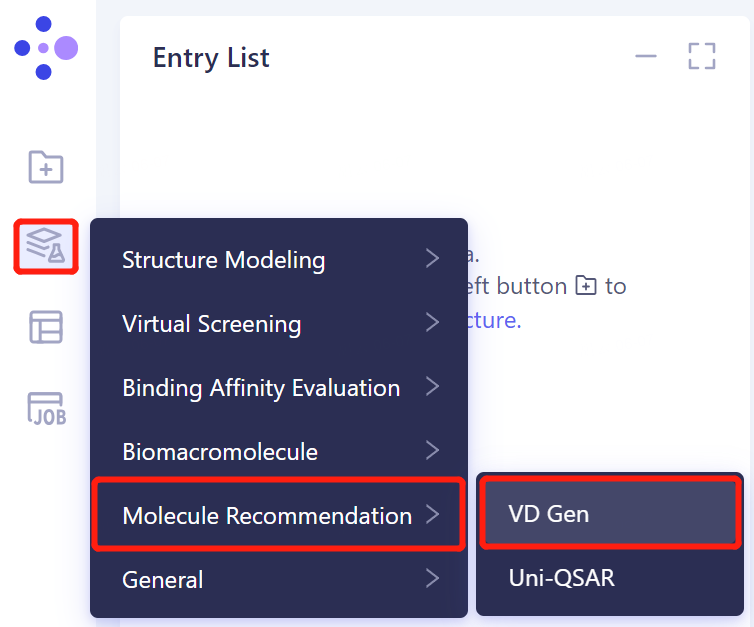
- The left general menu bar Menu → Function → Molecule Recommendation → VD Gen.
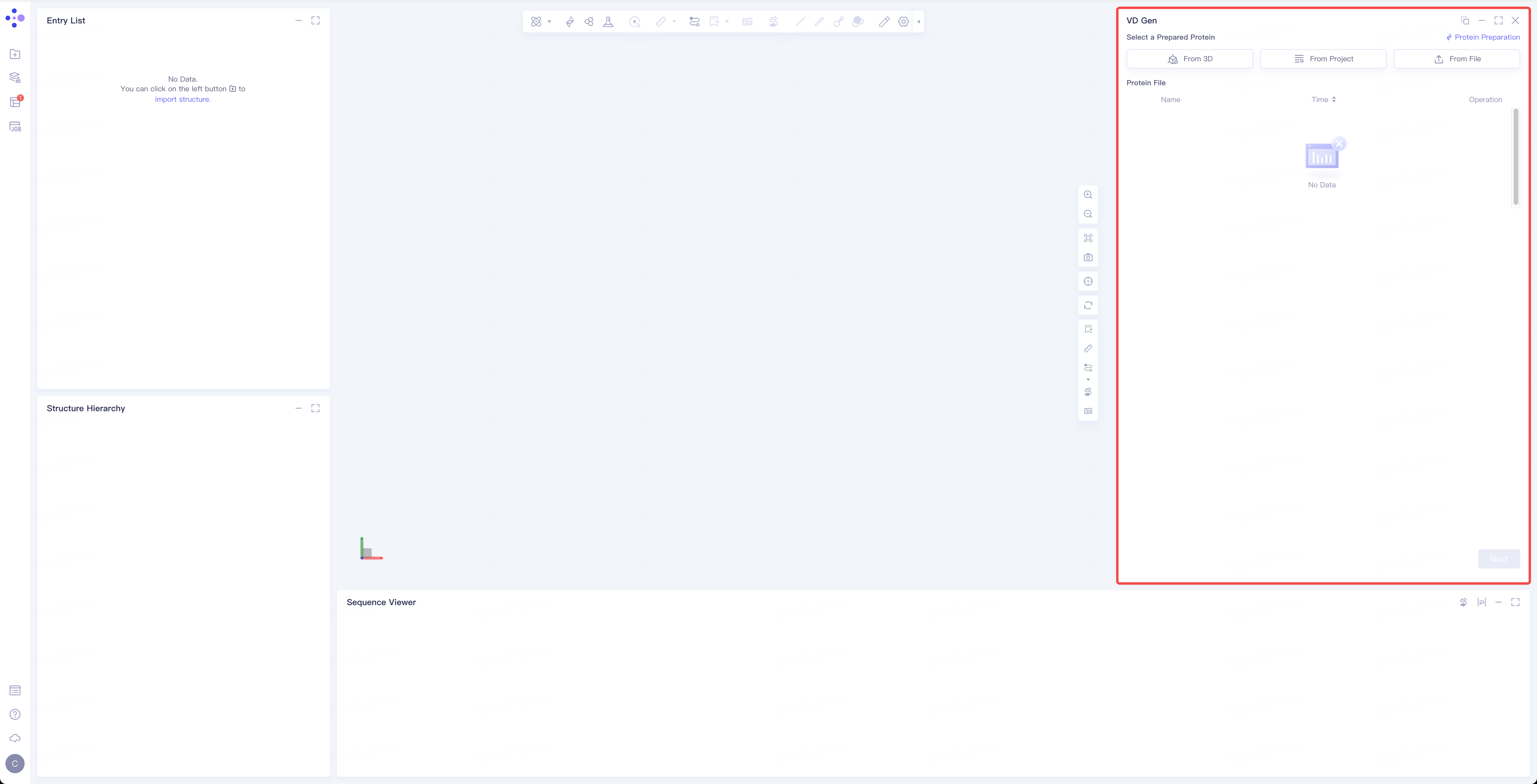
- The operation interface of VD Gen (shown in the red box) appears on the right side, and the overall interface is as follows:
2. Operation
2.1 Upload Protein Structure
-
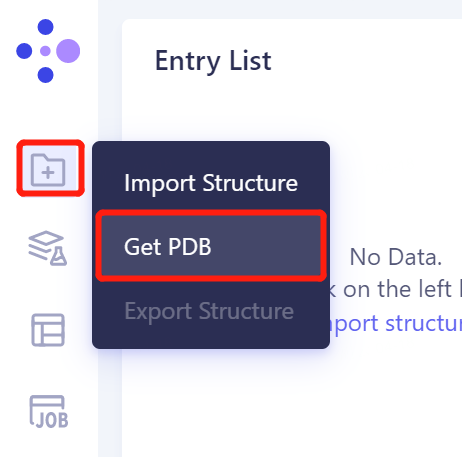
In the left general menu bar File → Get PDB → the pop-up interface, input the PDB ID: 8DI4 → Import.
- The two amino acid chains of 8DI4 are homodimers.
 |  |
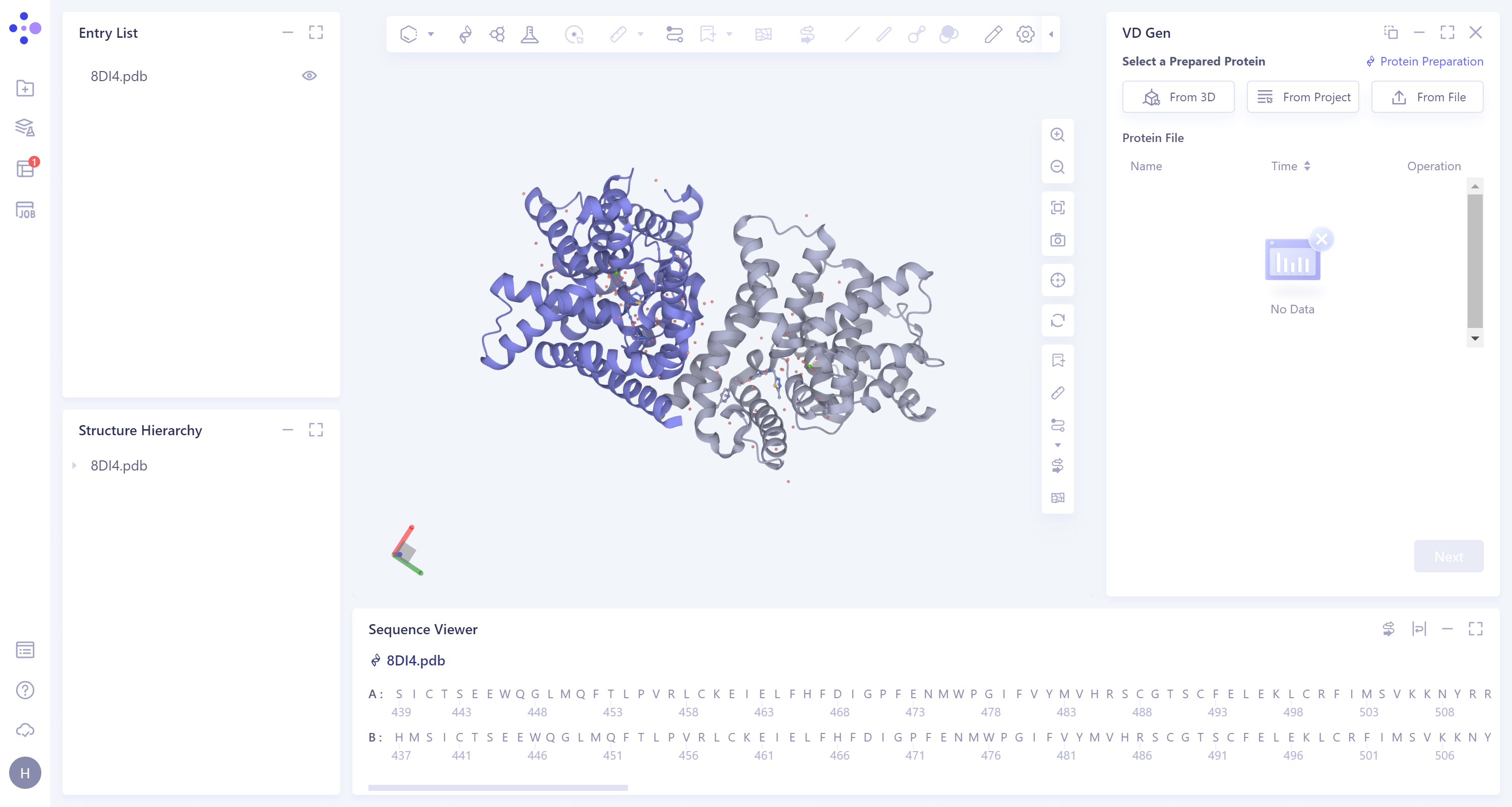
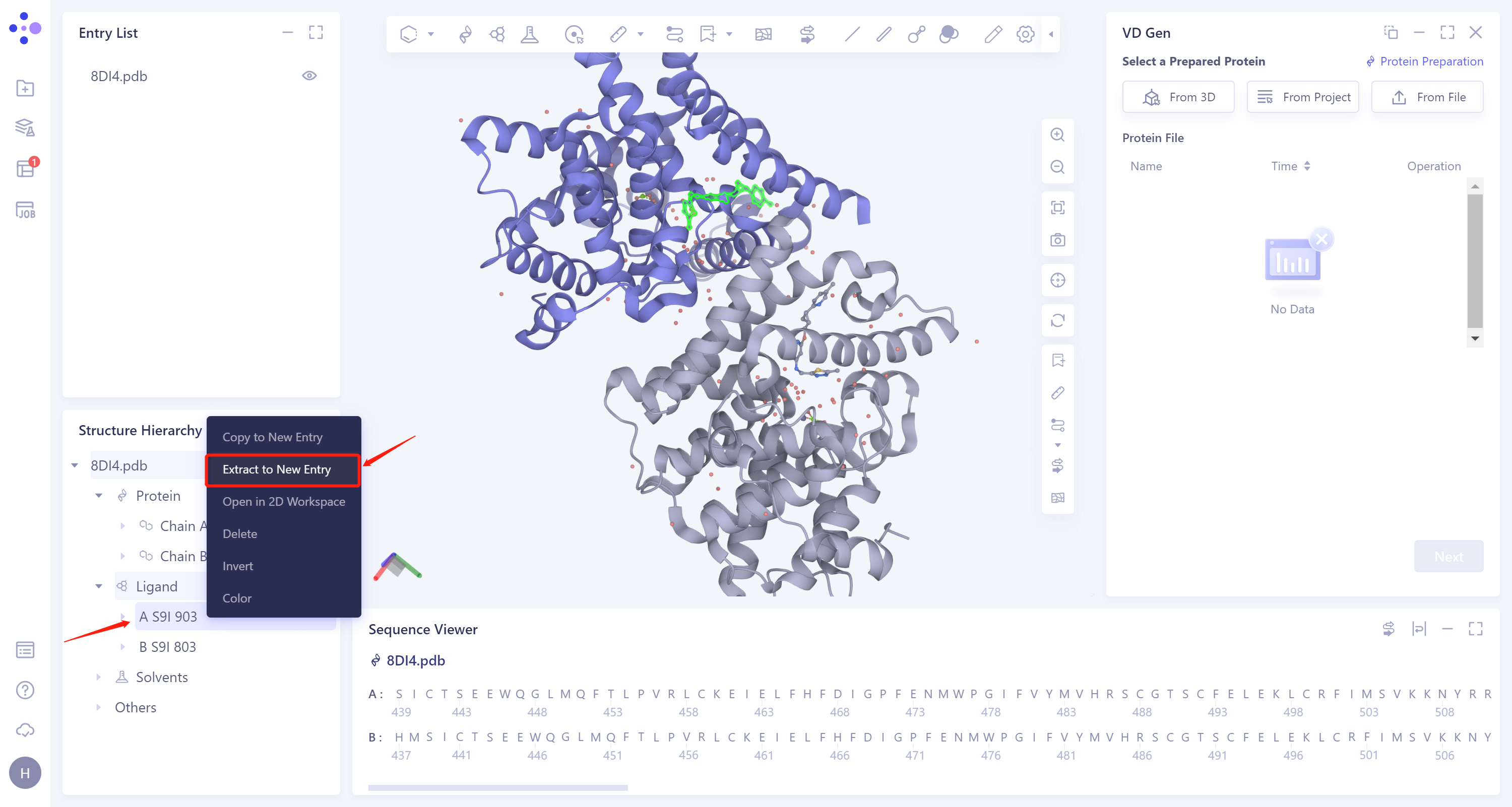
2.2. Extraction of ligand molecules
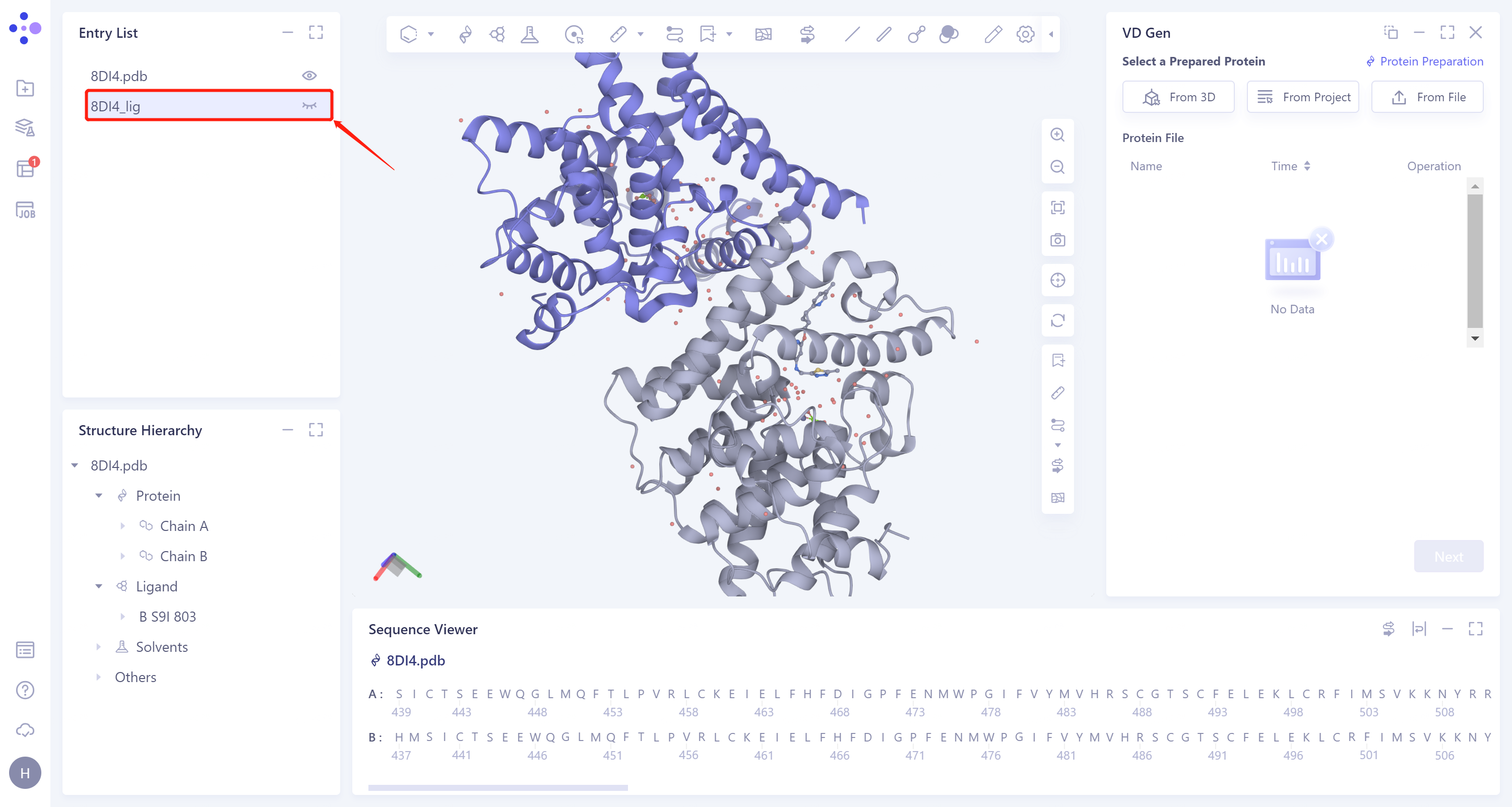
- Expand the 8DI4 structure, select the ligand molecule A S9I 903 on the A chain, right-click to select Extract to New Entry, and name the ligand 8DI4_lig.
 |  |
2.3. Protein structure input
- Select from 3D Workspace: Click the Select from 3D Workspace box → 'Select Structure' interface pops up → Select the A chain of 8DI4 in the Structure Hierarchy box on the left side of the interface → The name of the selected protein is displayed in the 'Select Structure' window. Click 'OK'.
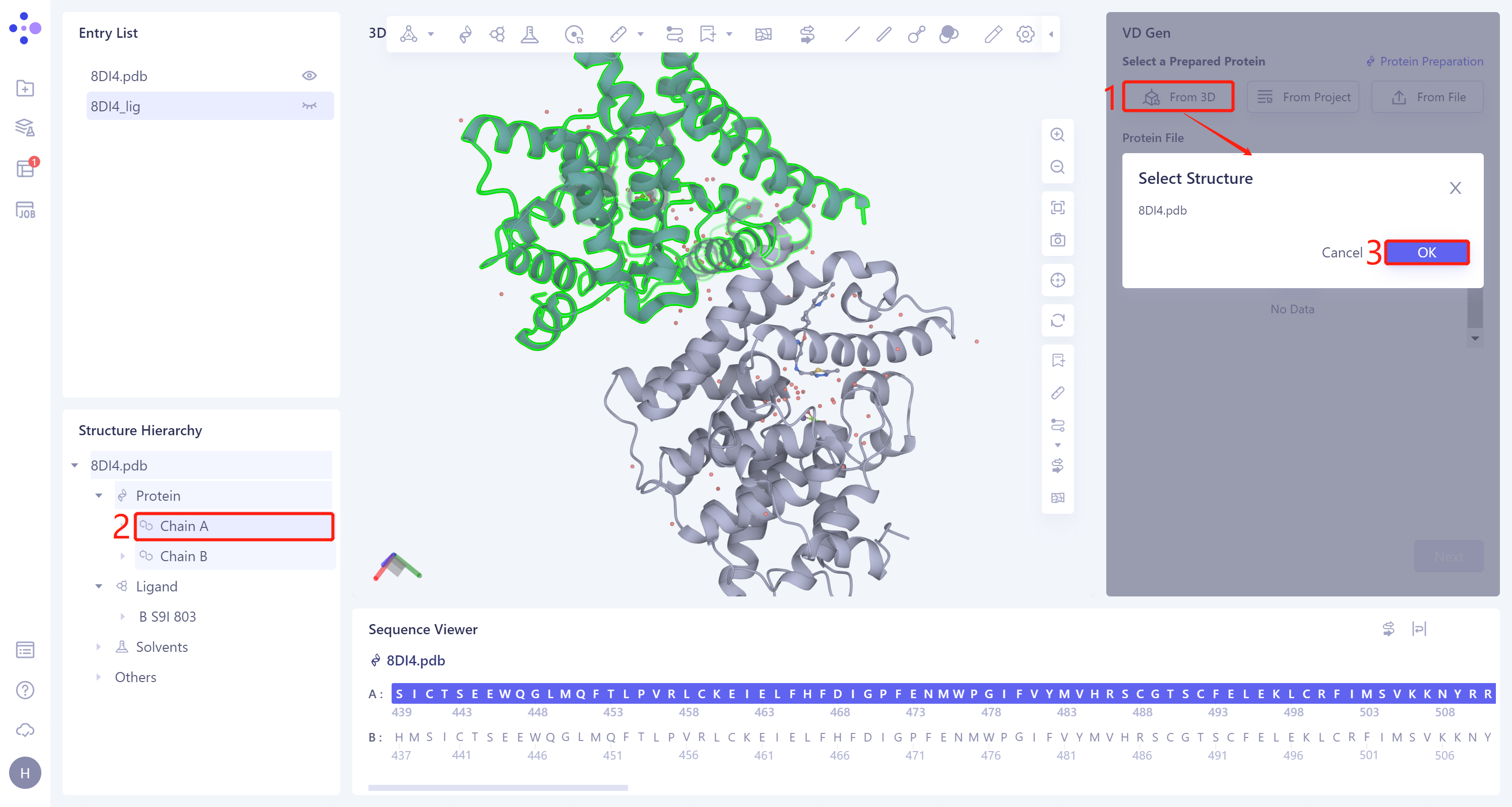
2.4 Reference Ligand File Input
- From 3D Workspace: Click the Select from '3D Workspace' interface → Select Ligands box pops up → Select 8DI4_lig in the Structure Hierarchy box on the left side of the interface → Select The Ligands box displays the name of the selected ligand. Click OK.
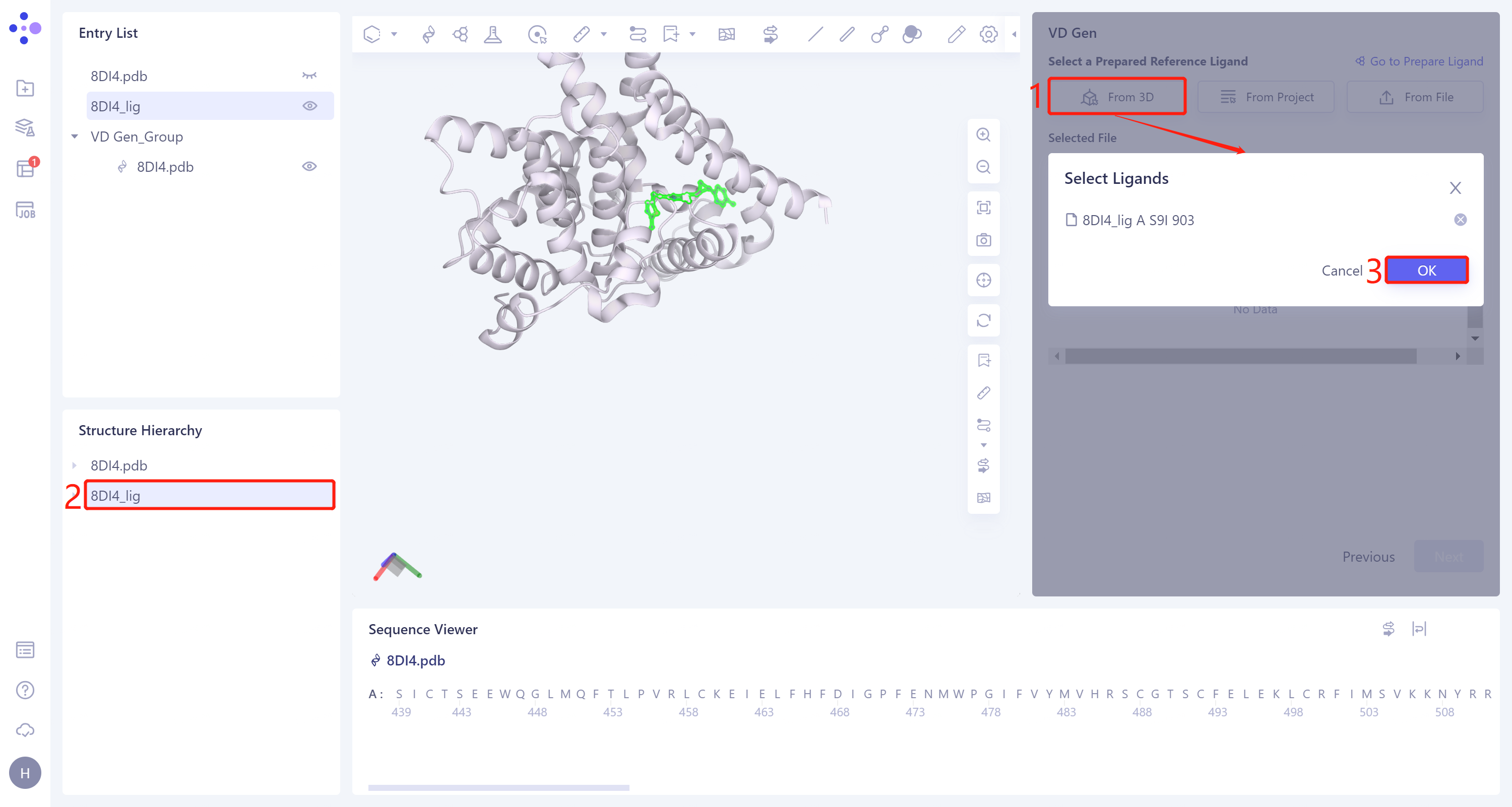
- Missing bond information of ligand molecules extracted from the.pdb file. Repair the missing bond information here. Click 'Yes'.
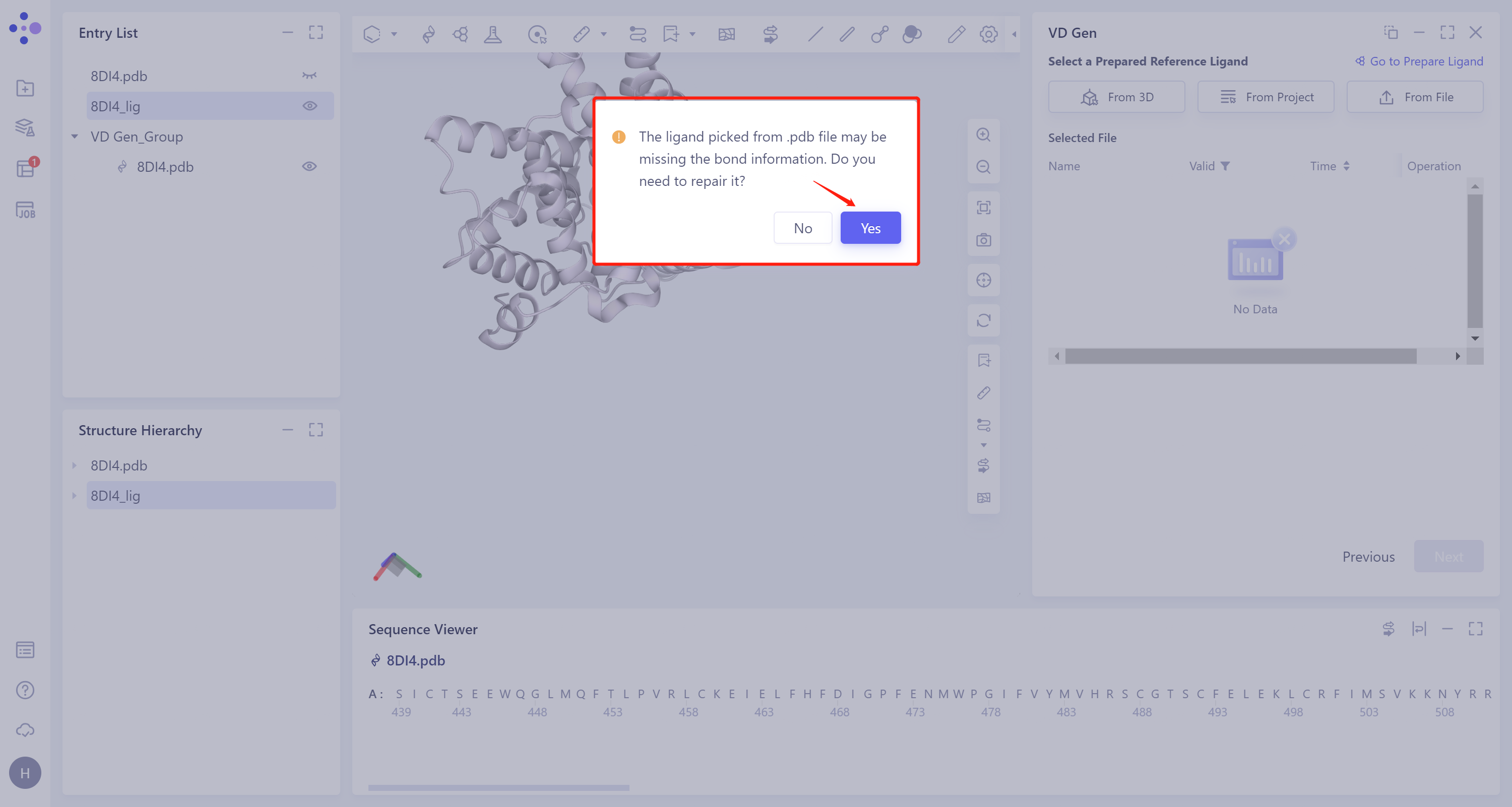
- The selected ligand will be hydrogenated and the force field will be checked. If the force field check of the ligand is not passed (here is Valid, that is, the force field check is passed), the Ligand Preparation is required (see the red box on the upper right of the figure for the shortcut entry). After the check is passed, click 'Next'.
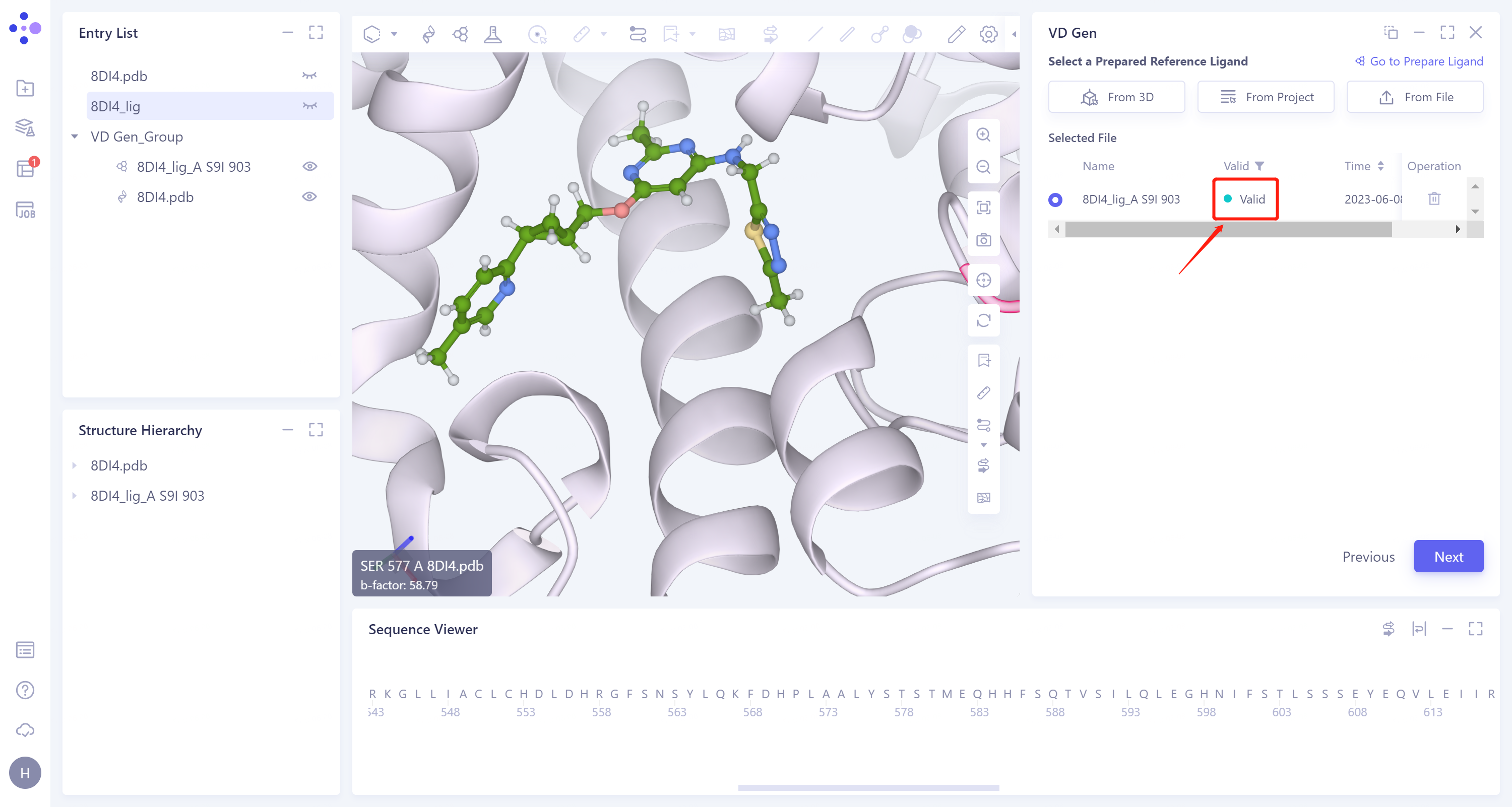
2.5 Molecular optimization
-
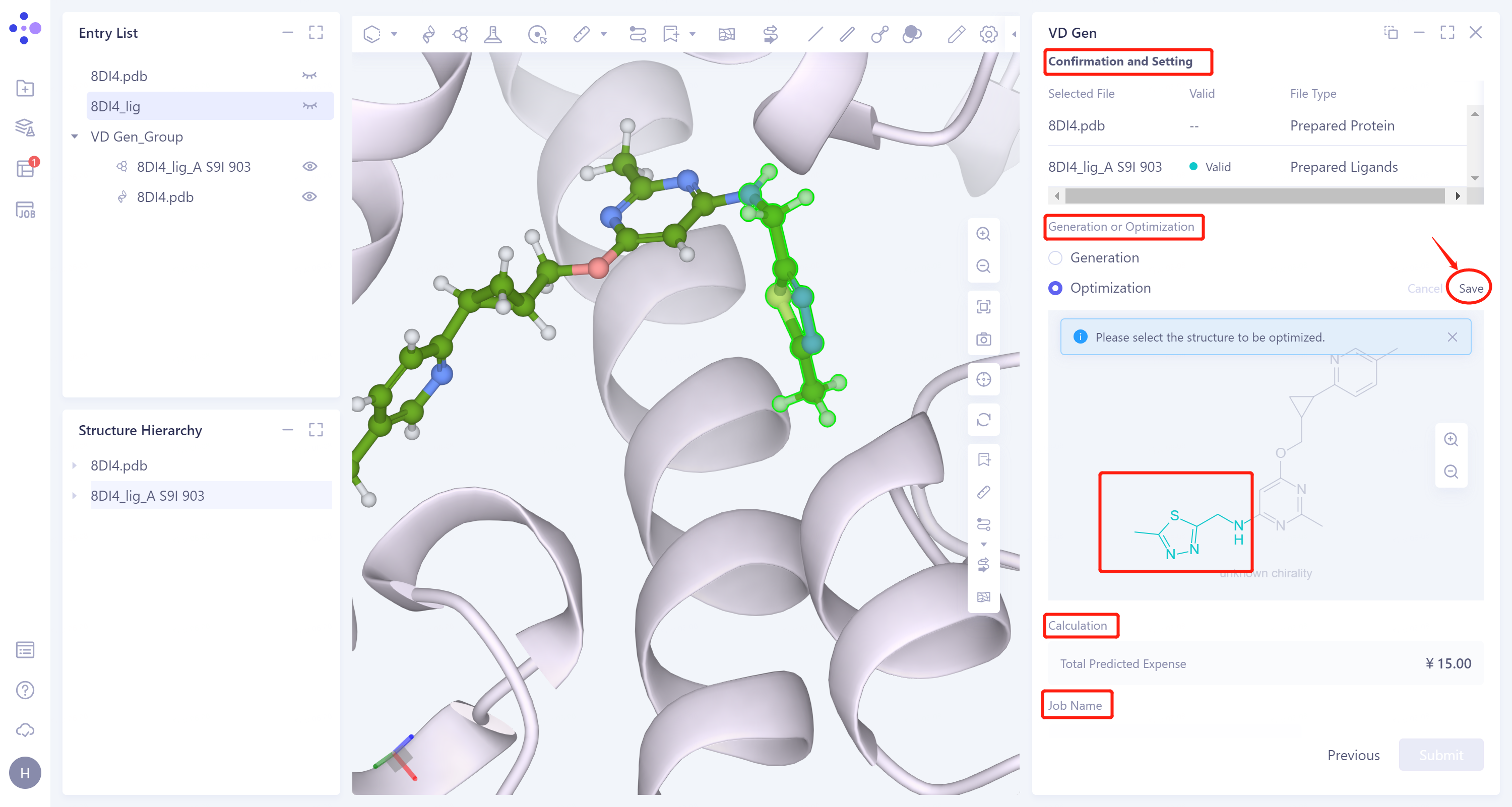
Confirm whether the input protein structure and reference ligand are correct at 'Confirmation and Setting'.
-
Generation or Optimization: Select 'Optimization' for molecular optimization, select the green part in the figure below, and click 'Save' in the upper right corner to optimize the segment.
-
Confirms the calculation cost at 'Calculation' .
-
The task is named 'VD Gen_Opt'.
3. Analysis of results
3.1 Entrance
-
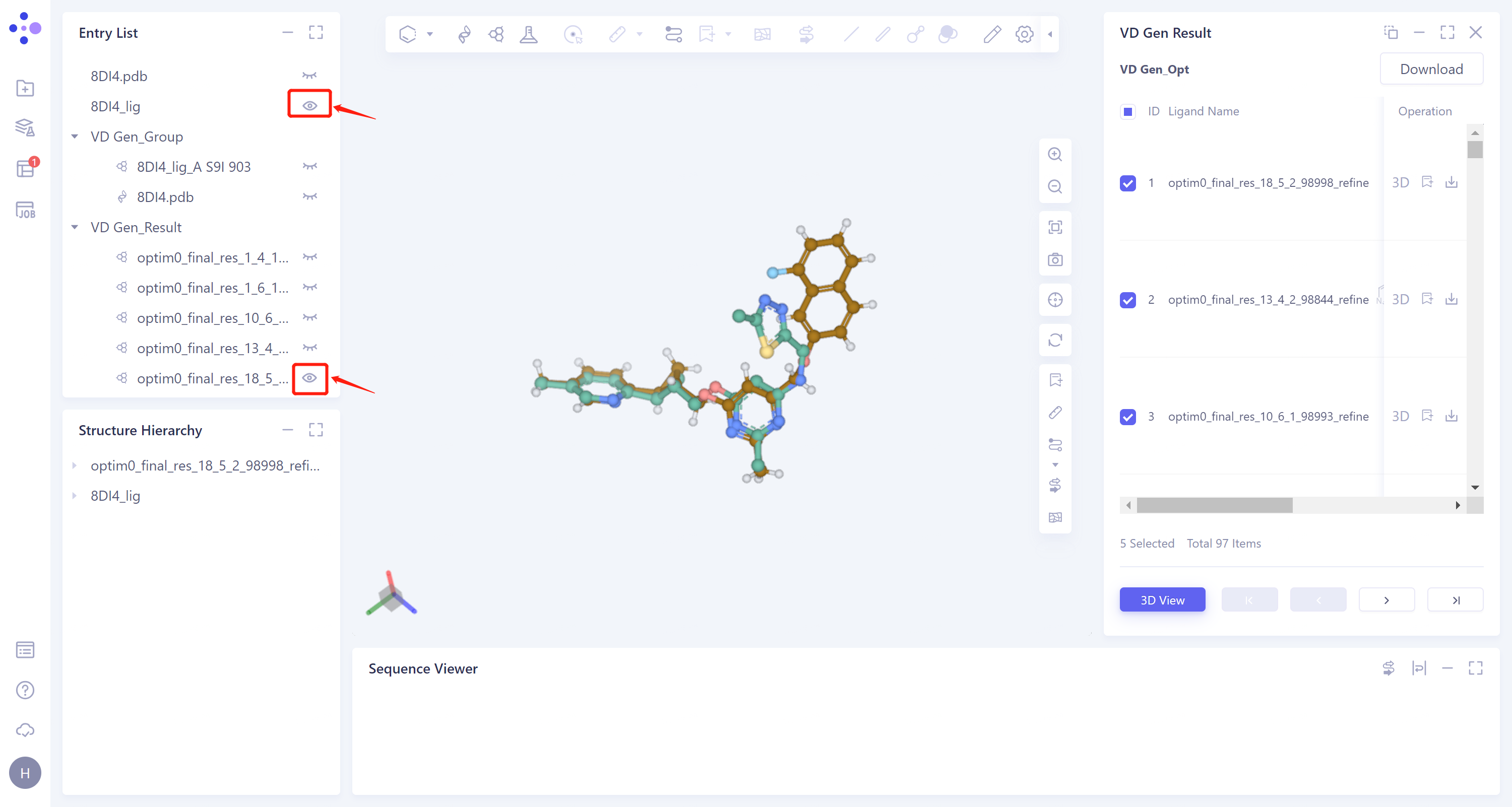
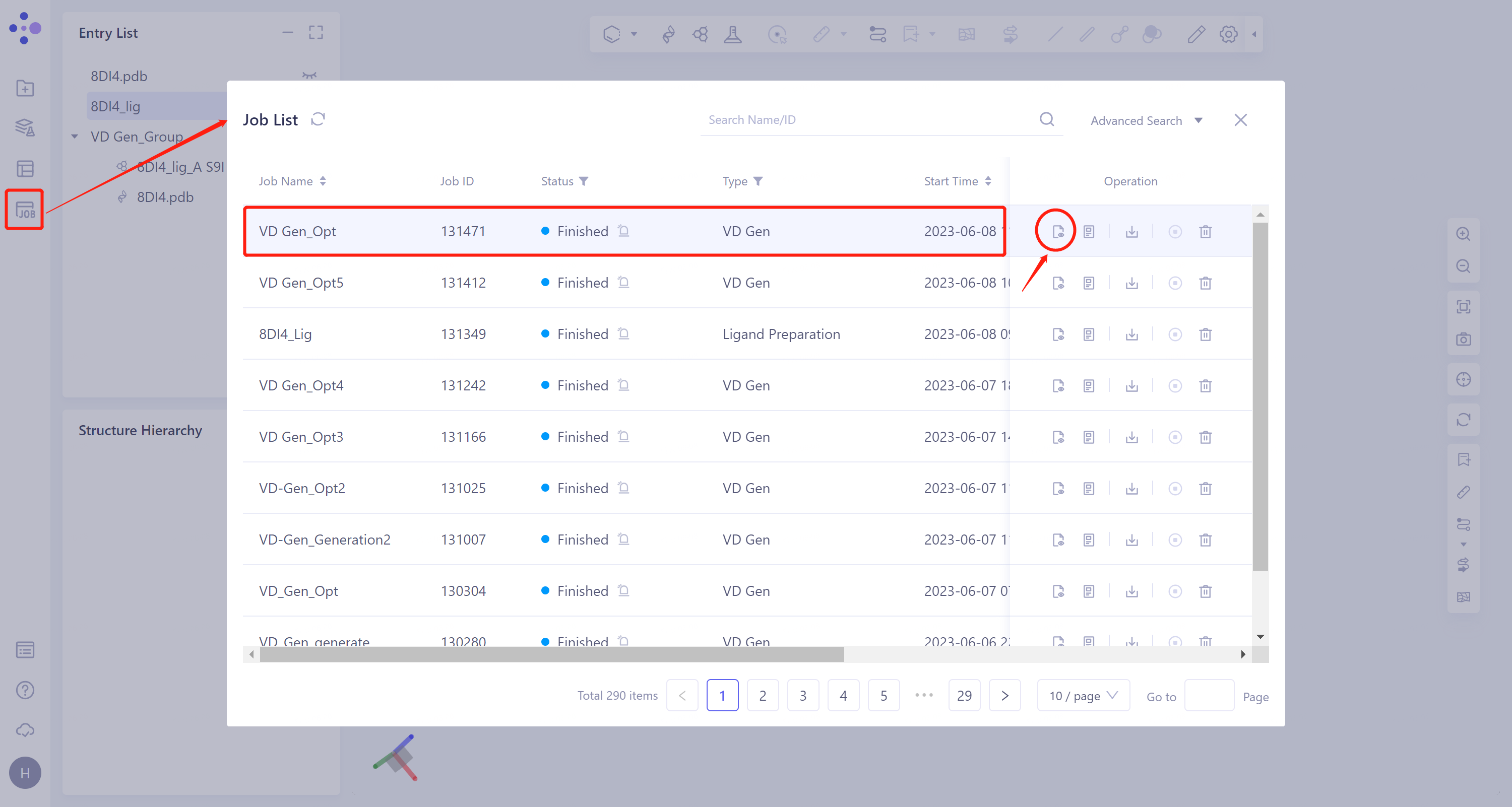
General menu bar on the left: Menu Job → Find the task 'VD Gen_Opt' → Click 'Show' in the 'Operation' column to view the task results.
-
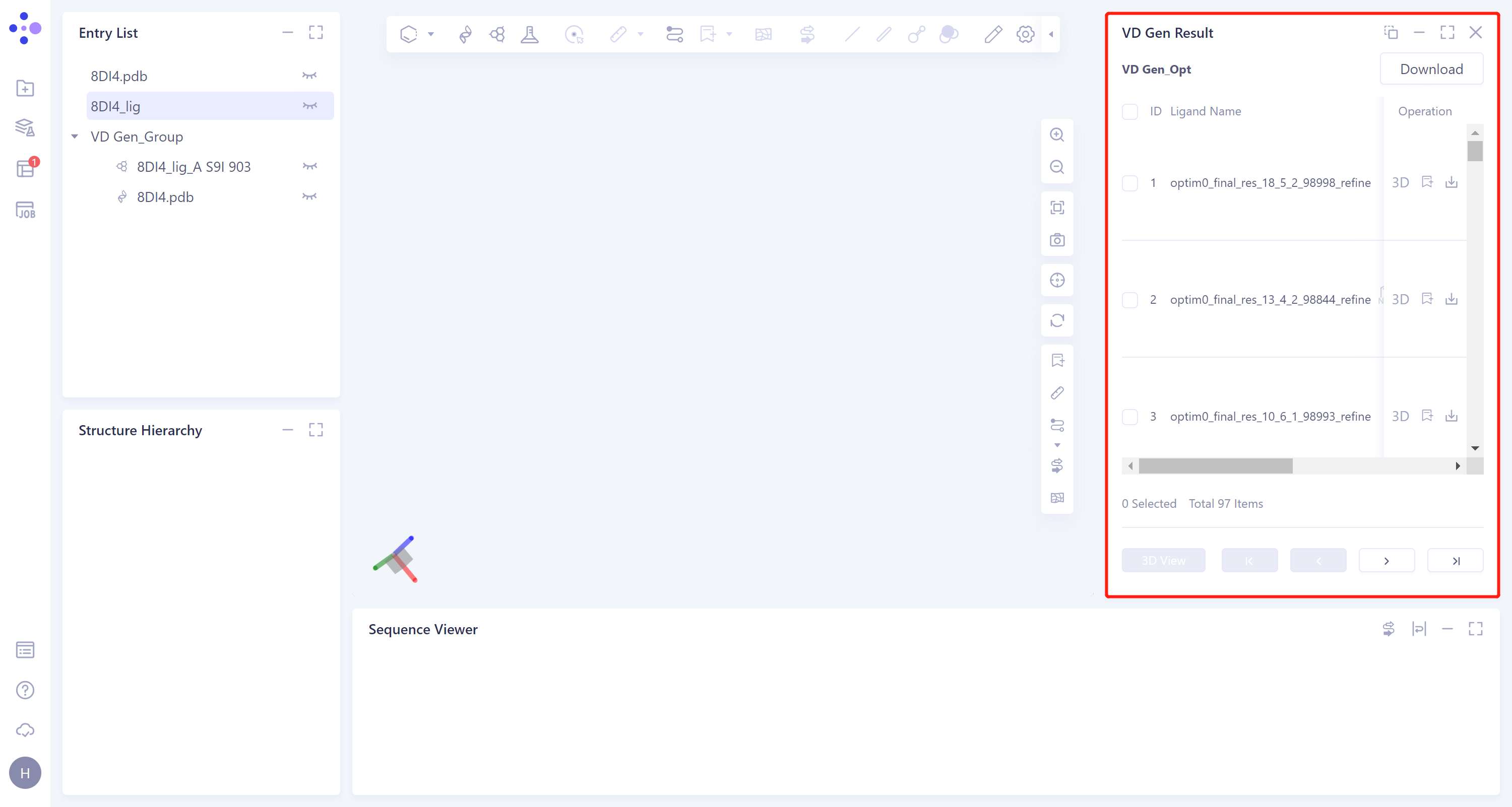
The result is shown as the right figure.
 |  |
3.Display the top five molecular conformations.
-
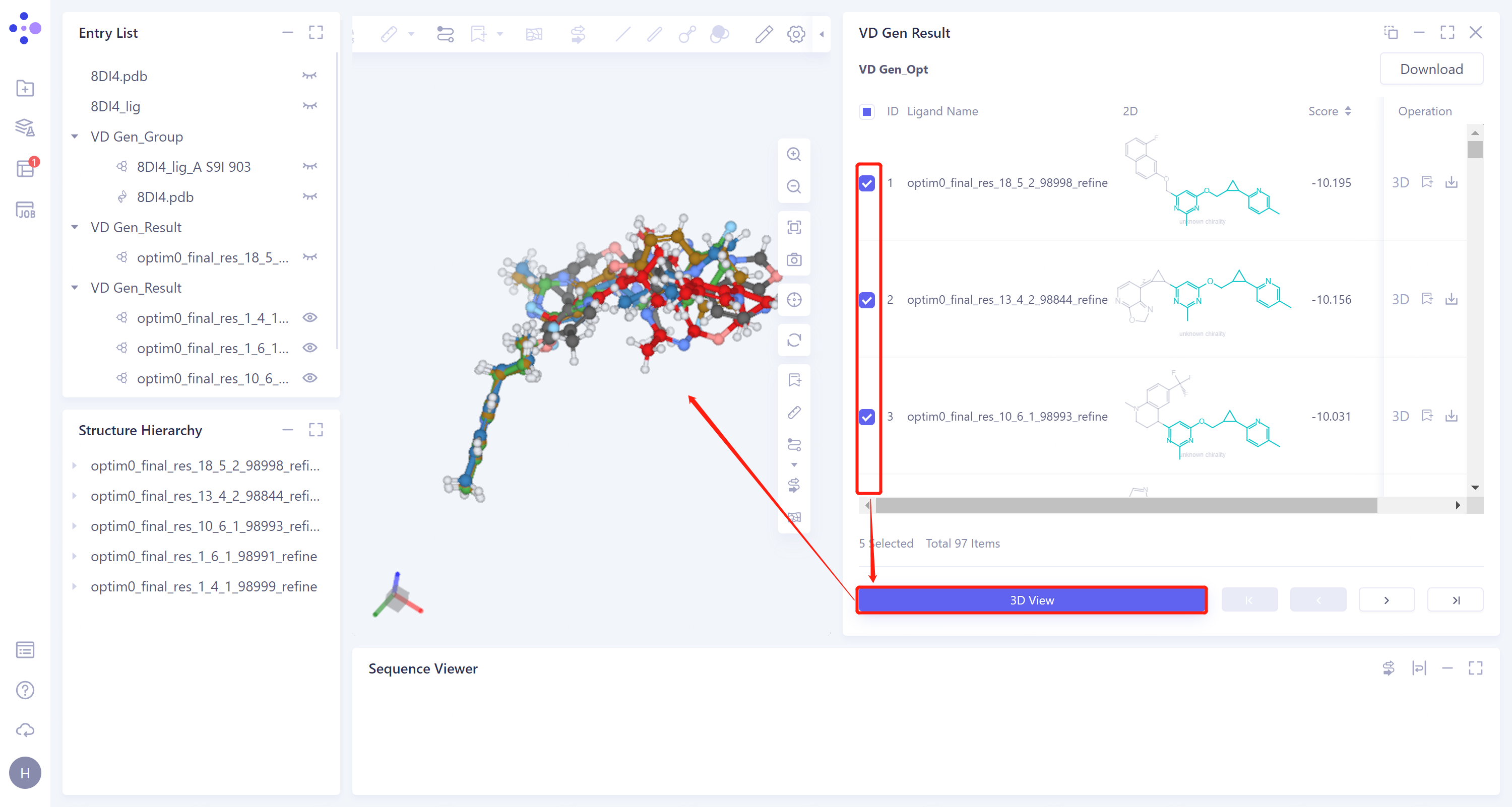
In the 'VD Gen Result' window, the name of the generated ligand molecule, the 2D image and the docking score of the ligands with the receptor are given in the list. The gray parts in the 2D image of the molecules are the molecular fragments generated by VD Gen.
-
From the 2D images, it can be seen that VD Gen has generated new molecular substructures based on the original molecular fragments.
-
Choose the top five molecules according to the docking scores in the check box, click '3D View' below, and these five molecules will be displayed in the 3D Workspace window.
3.3 Structural comparison
- In the Entry List window, 8DI4_lig is adjusted to the display state, and the result of VD Gen only displays the molecule with the highest score. The high degree of superposition of the two molecules indicates that the docking site of VD Gen is reasonable when using the generated molecule for docking, and the score can reflect the affinity of the molecule with the receptor to a certain extent.