Protein-Protein Interaction
Introduction
Many functions of proteins involve binding to other proteins, which are called protein-protein interactions (PPIs). PPIs control many cellular functions in biological signal transduction, transport, defense and so on. Therefore, understanding the mechanism of protein-protein interactions is essential for understanding disease mechanisms. In this tutorial, you will learn to use the Protein-Protein Interaction module of the Hermite ® platform to predict p53TAD-MDM2 [1] interactions and understand protein-protein binding patterns.
Protein-protein complex structure files used in this tutorial:
1. Create a project and import the structure
1.1 Log in to the system
- Login address: https://hermite.dp.tech

1.2 Create the project
- Once in the system, create a new project “p53TAD-MDM2 PPI”
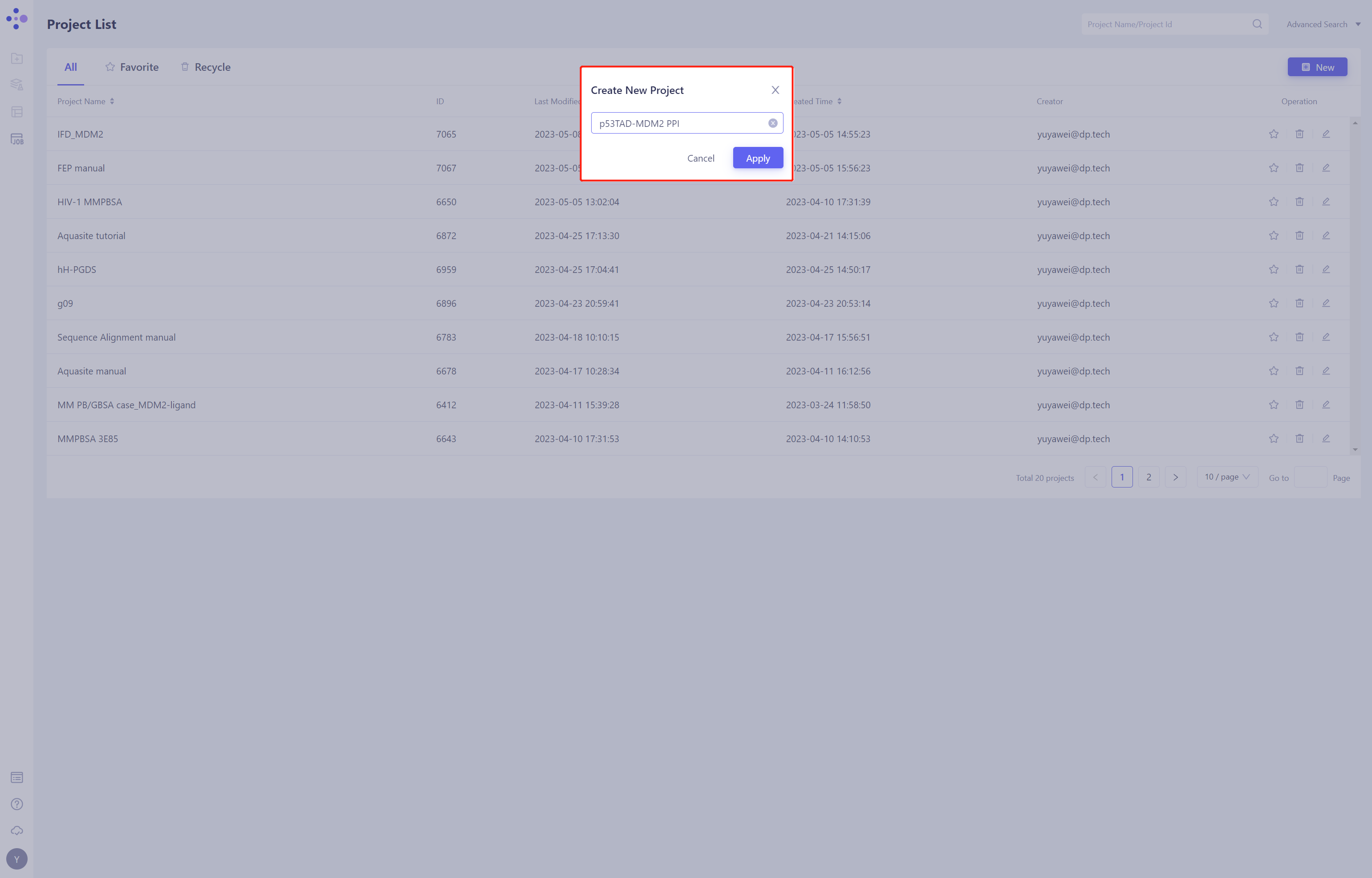
1.3 Import Protein Structure
- Left general menu bar Menu → File → Import Structure
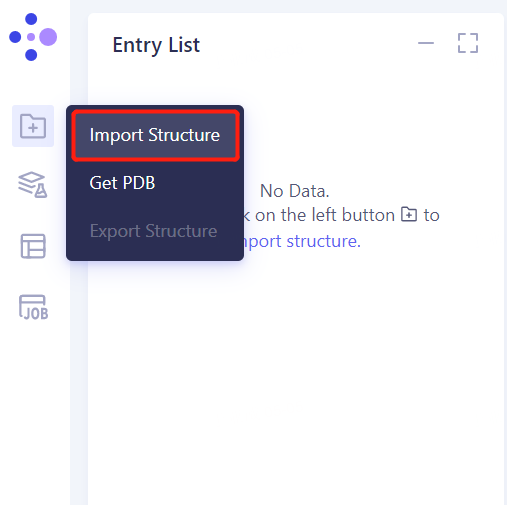
- Click Select Files, select the 1YCR _ Prep. PDB file, and click Import to import the protein structure
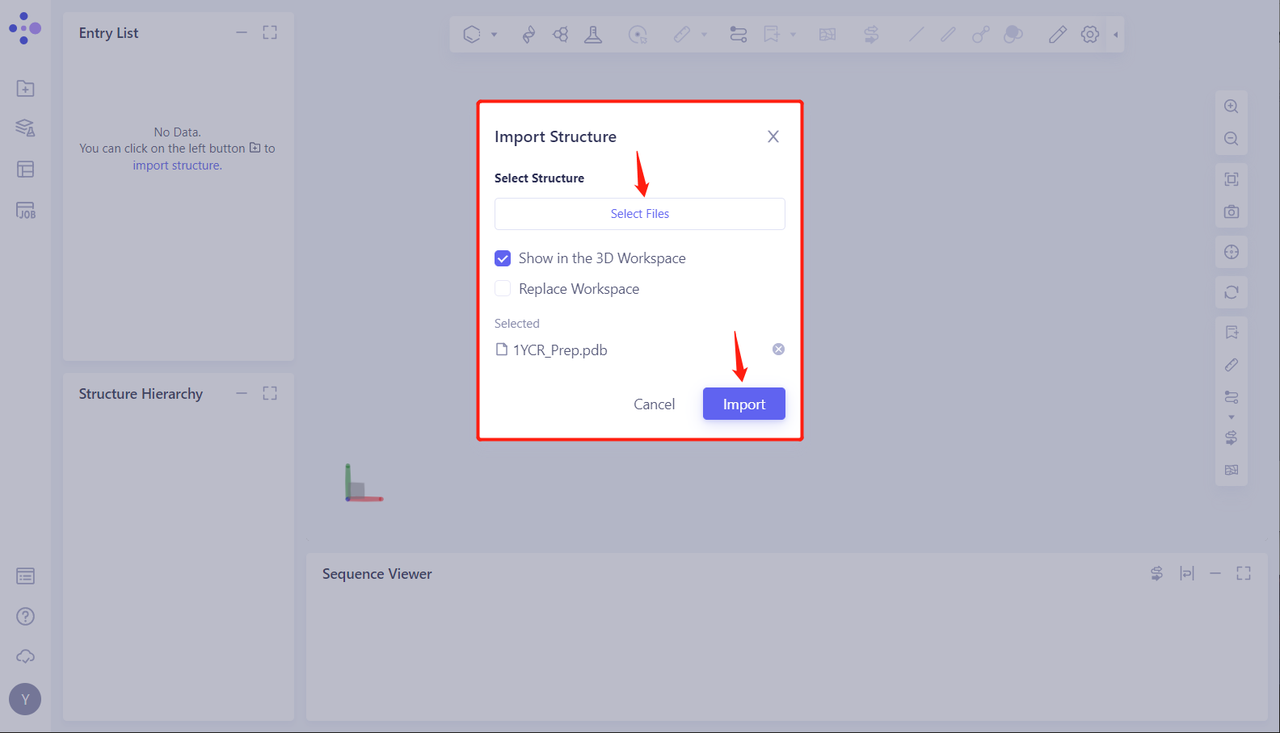
2. Creating a Protein-Protein Interaction Task
2.1 Left general menu bar Menu → Function → Biomacromolecule → Protein-Protein Interaction
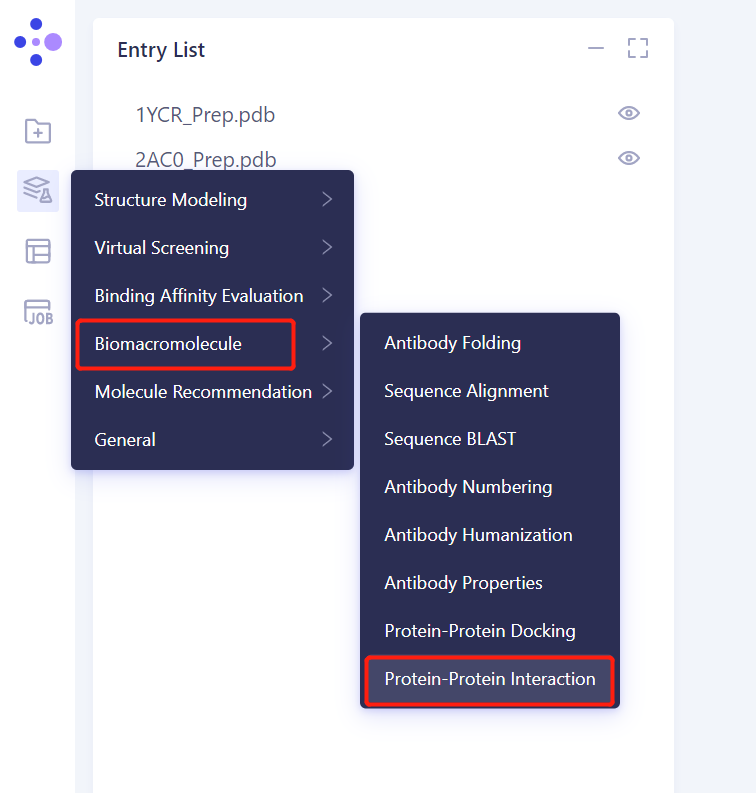
2.2 Select a Prepared Receptor Protein
- Select “1YCR _ Prep. PDB”-Protein-Chain A “from the Structure Hierarchy, and click OK
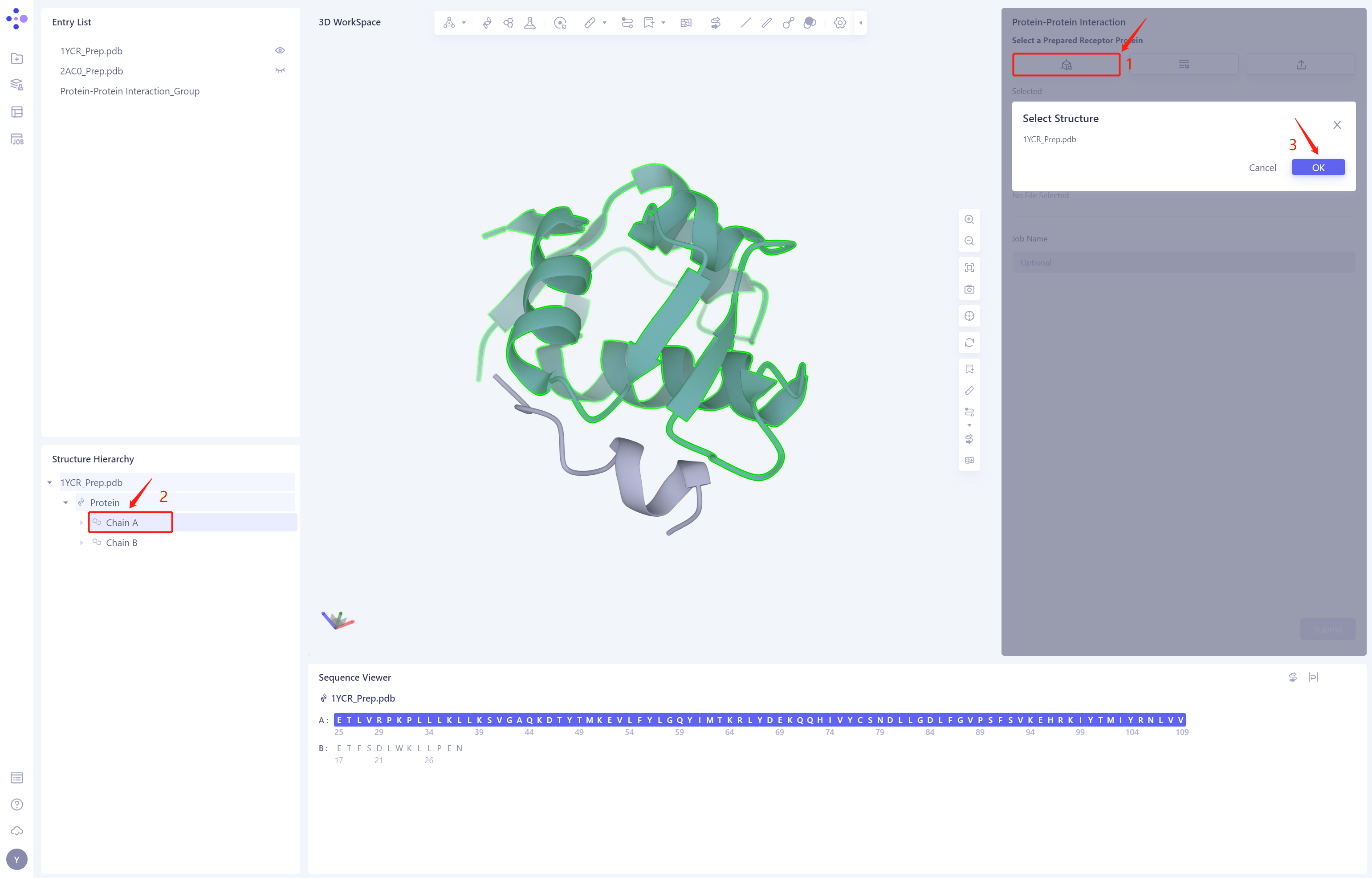
2.3 Select a Prepared Ligand Protein
- Select “1YCR _ Prep. PDB”-Protein-Chain B “from Structure Hierarchy and click OK
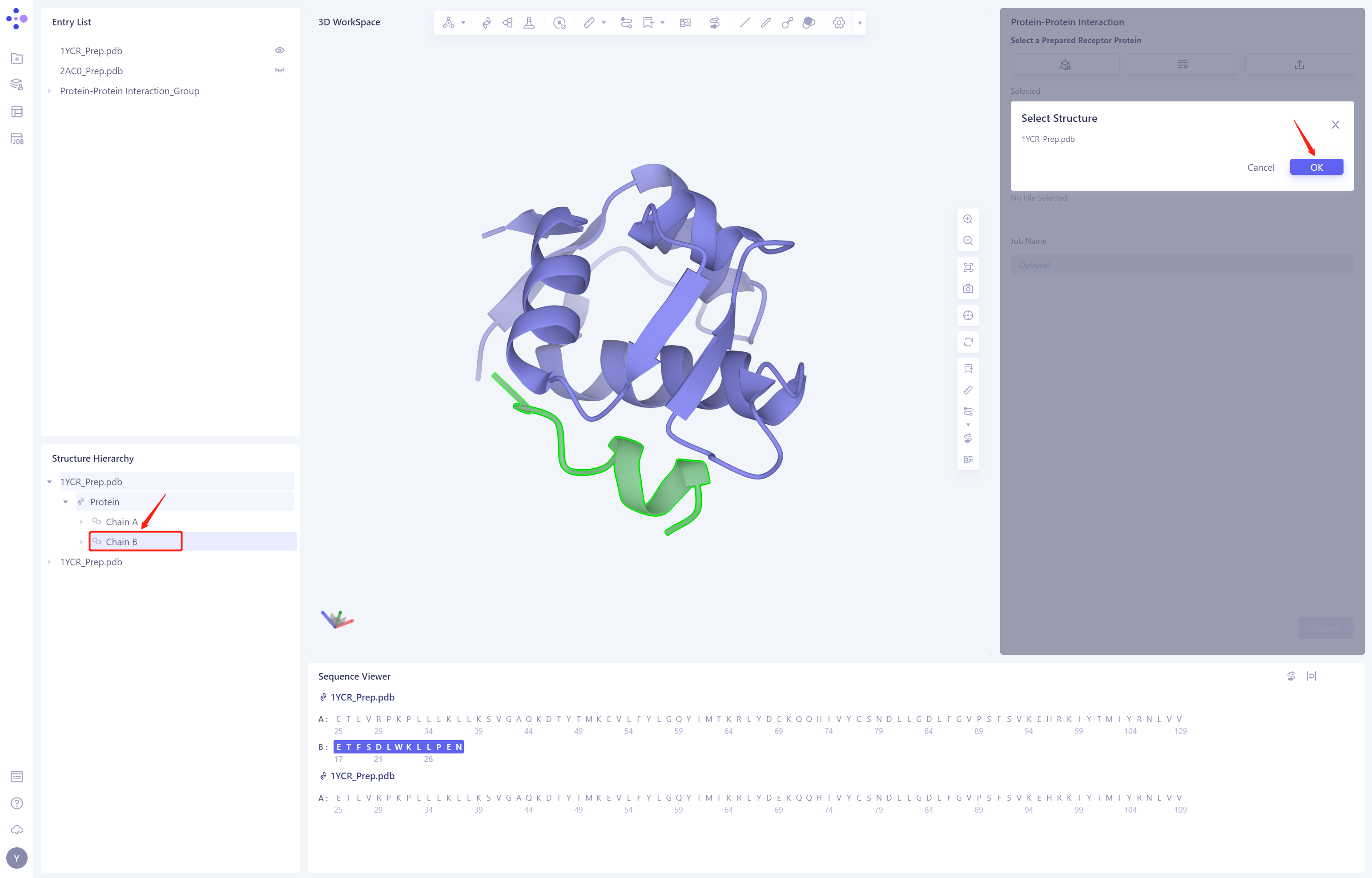
- The Job Name is “p53TAD-MDM2 PPI”. Click Submit to submit the job.
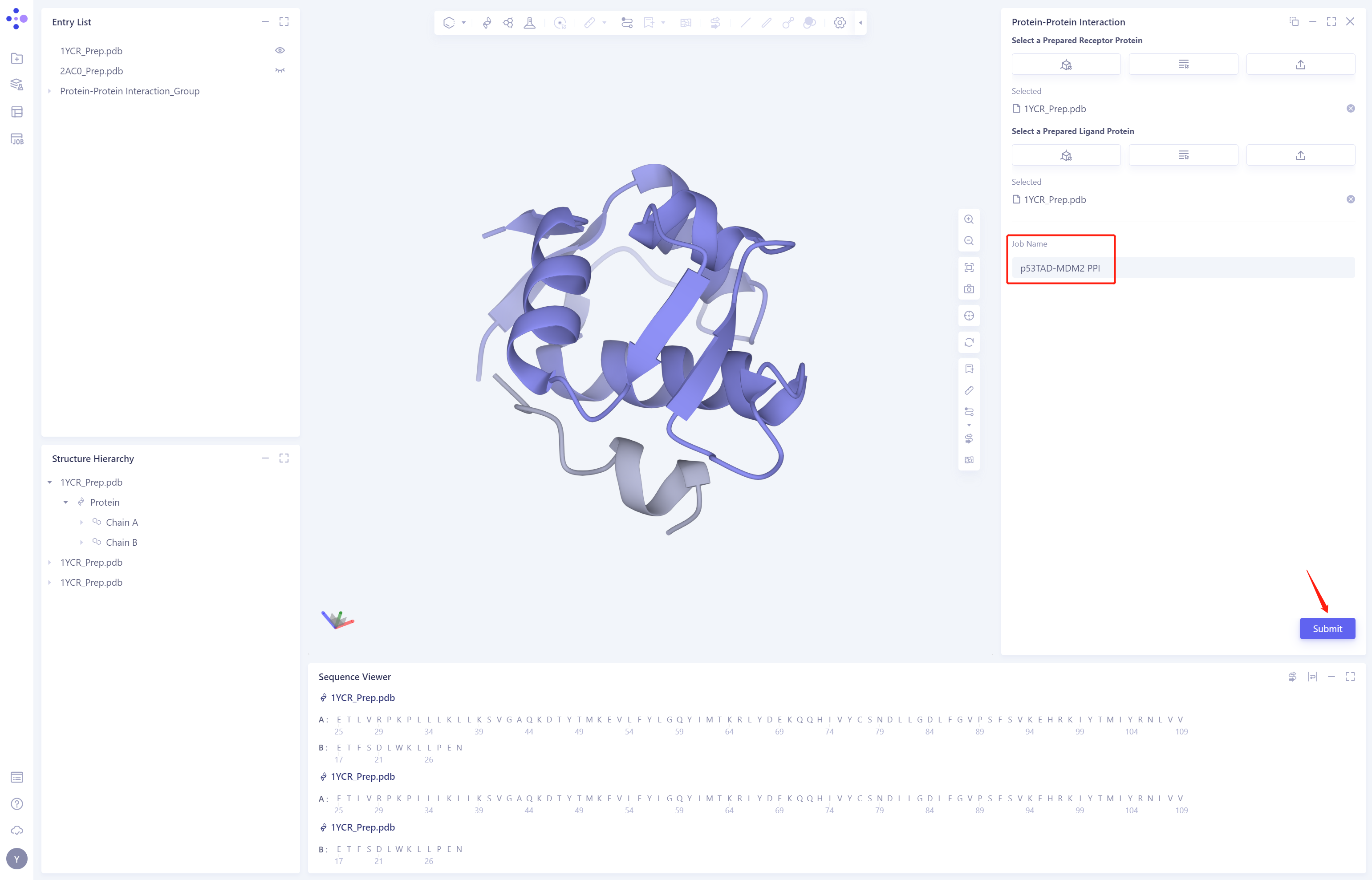
3. Analysis of results
3.1 Entrance
- Job → Job List in the left general menu bar, find the “p53TAD-MDM2 PPI” calculation task, and click “Show”
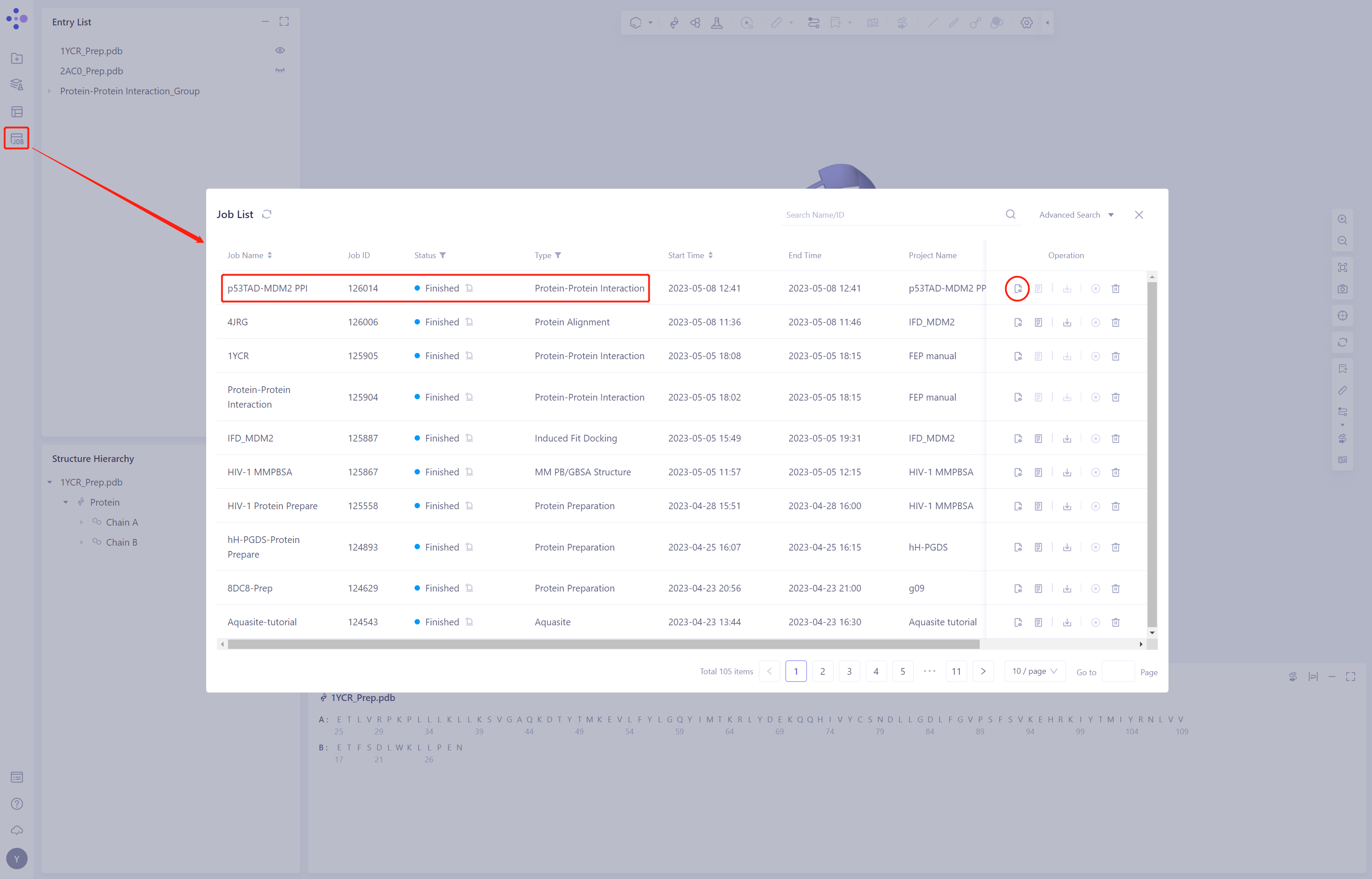
3.2 Results presentation
-
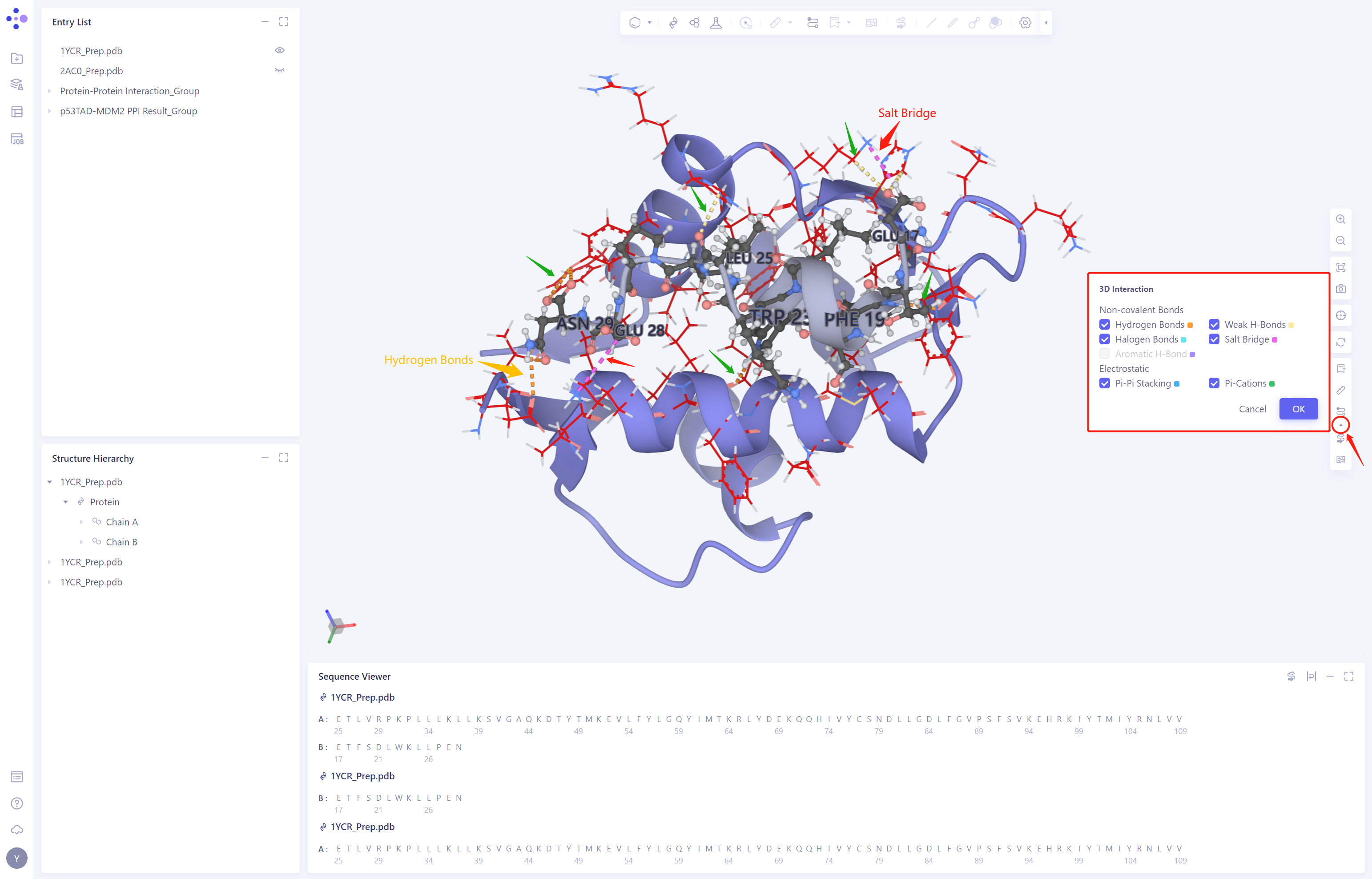
View the interaction between p53TAD (black) and MDM2 (blue) molecules according to the ID type of the 3D Interactions type.
-
Residues in p53TAD that are involved in intermolecular interactions are identified by residue name. It was found that there were mainly Hydrogen Bond and Salt bridge interactions between p53TAD and MDM2, which stabilized the protein-protein binding.
4. References
[1] Structure of the MDM2 Oncoprotein Bound to the p53 Tumor Suppressor Transactivation Domain. SCIENCE. 8 Nov 1996. Vol 274, Issue 5289. pp. 948-953.
[2] Structural Basis of DNA Recognition by p53 Tetramers. Molecular Cell 22, 741–753, June 23, 2006.