Antibody Properties
Introduction
In the process of antibody drug design, predicting the properties of antibodies is helpful for us to select the best antibody drug candidates and optimize the performance of antibody drugs. Antibody properties mainly include immunogenicity, stability, physical and chemical properties, and post-translational modification. Currently, with the availability of large data sets of antibody sequences, structures, and biophysical properties, the development of tools for antibody structure prediction and property calculation is possible.
Adalimumab (ADA) is the first fully humanized monoclonal antibody against Tumor Necrosis Factor (TNFα). It can specifically bind to soluble human TNF-α and block its interaction with cell surface TNF receptors p55 and p75, thus effectively blocking the inflammatory effect of TNF-α and improving the intestinal inflammatory activity of Crohn’s disease (CD). It is one of the main biological agents for the treatment of CD.
The Antibody Properties module of the Hermite ® platform provides the ability to predict antibody properties. Based on the user-friendly graphical task submission and result analysis interface, you can easily and quickly realize the prediction of biophysical properties based on antibody structure and generic principles.
In this tutorial, various antibody properties are predicted based on the sequence of Adalimumab, and the sequence information is as follows:
> Heavy Chain
EVQLVESGGGLVQPGRSLRLSCAASGFTFDDYAMHWVRQAPGKGLEWVSAITWNSGHIDYADSVEGRFTISRDNAKNSLYLQMNSLRAEDTAVYYCAKVSYLSTA SSLDYWGQGTLVTVSS
> Light Chain
DIQMTQSPSSLSASVGDRVTITCRASQGIRNYLAWYQQKPGKAPKLLIYAASTLQSGVPSRFSGSGSGTDFTLTISSLQPEDVATYYCQRYNRAPYTFGQGTKVEIK
1. Create Antibody Properties Task
1.1 Create a task
- The left general menu bar → Function → Biomacromolecule → Antibody Properties.
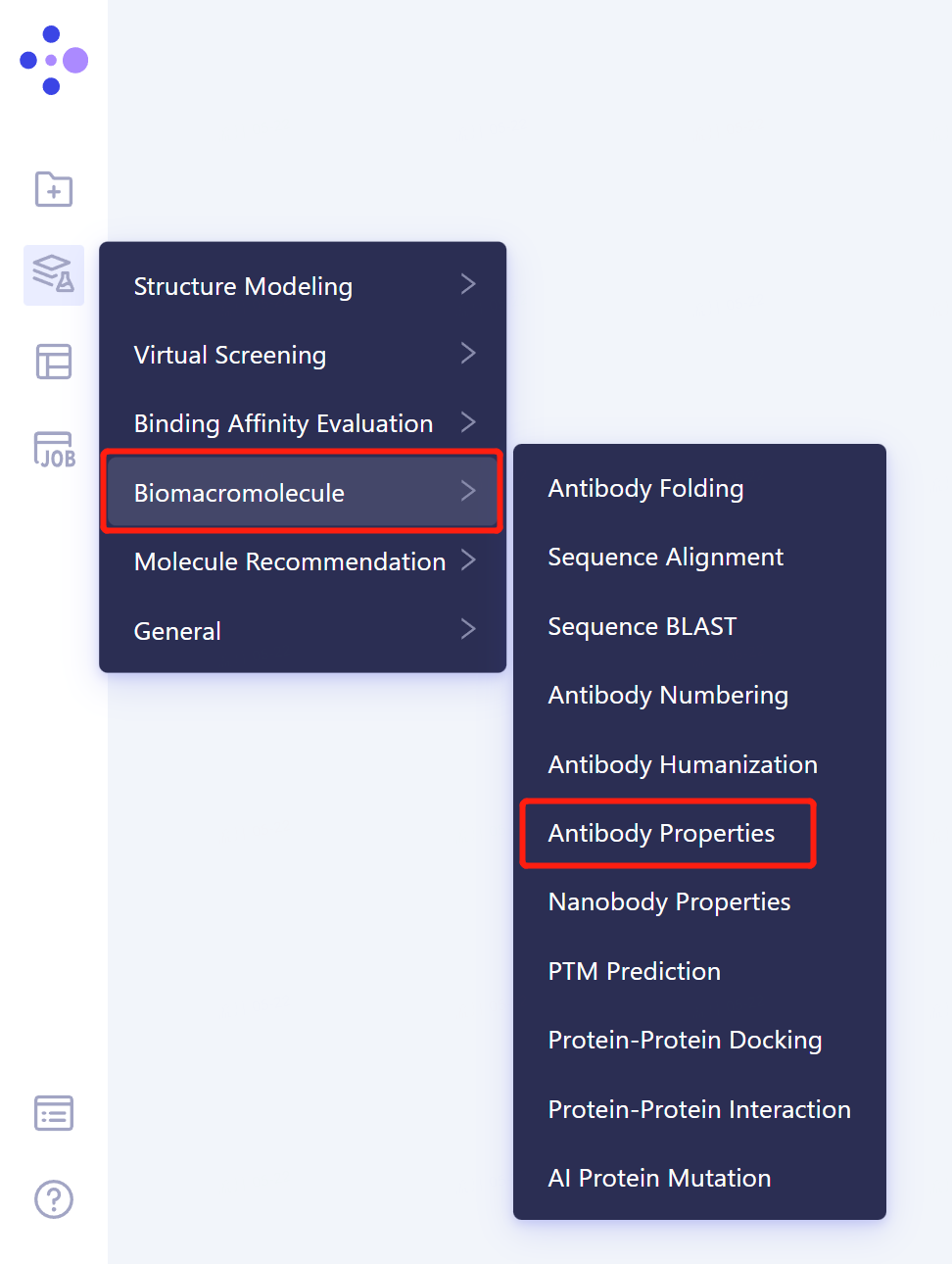
- The operation box of Antibody Properties (shown in the red box) appears on the right side, and the overall interface is as follows:
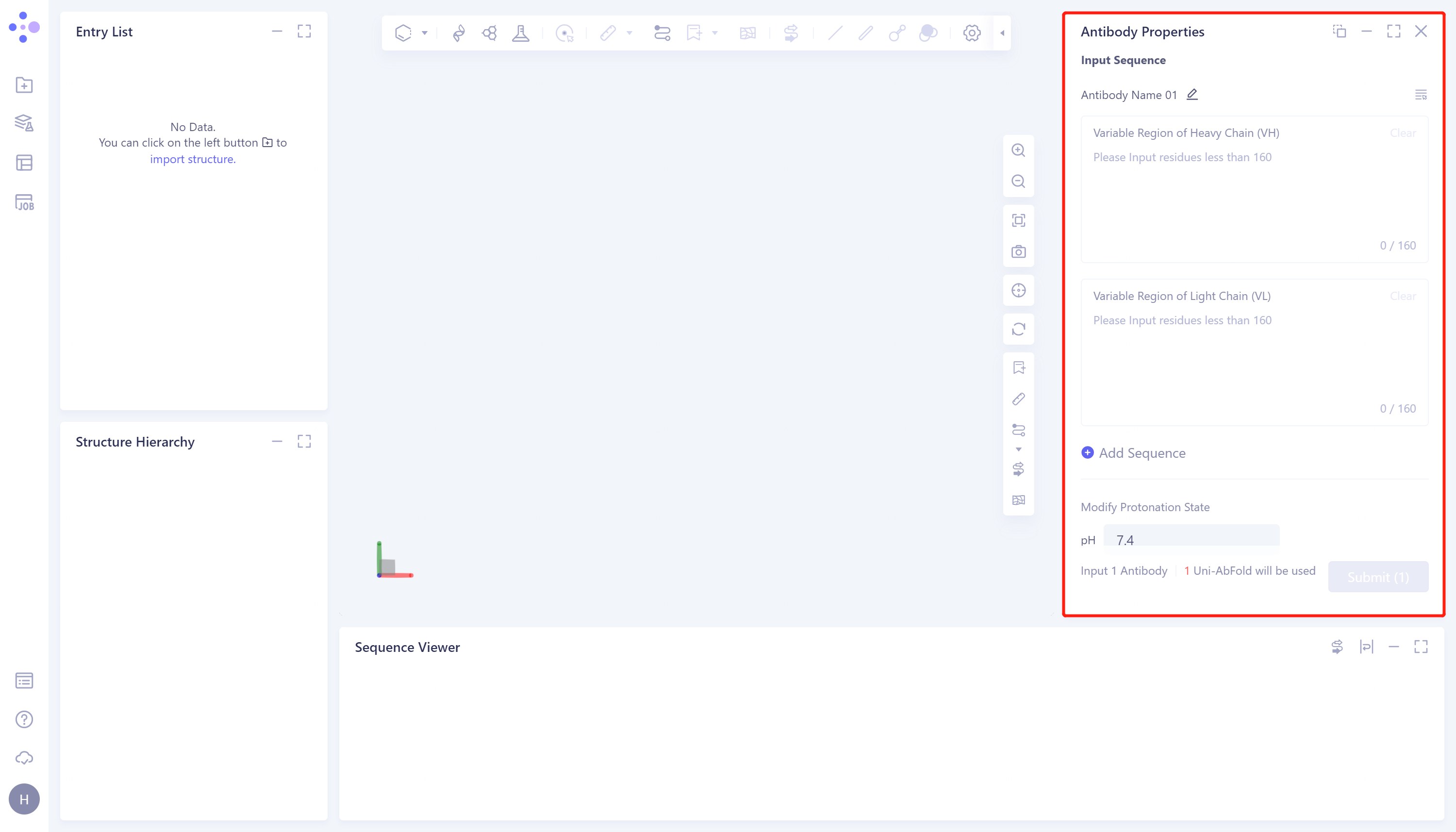
1.2 Input antibody sequences
-
Input Sequence: Input the sequence of the variable region of the antibody, which is directly input here.
-
Enter the variable region sequence of the antibody heavy chain in the sequence box under Variable Region of Heavy Chain (VH);
-
Enter the variable region sequence of the antibody light chain in the sequence box under Variable Region of Light Chain (VL);
-
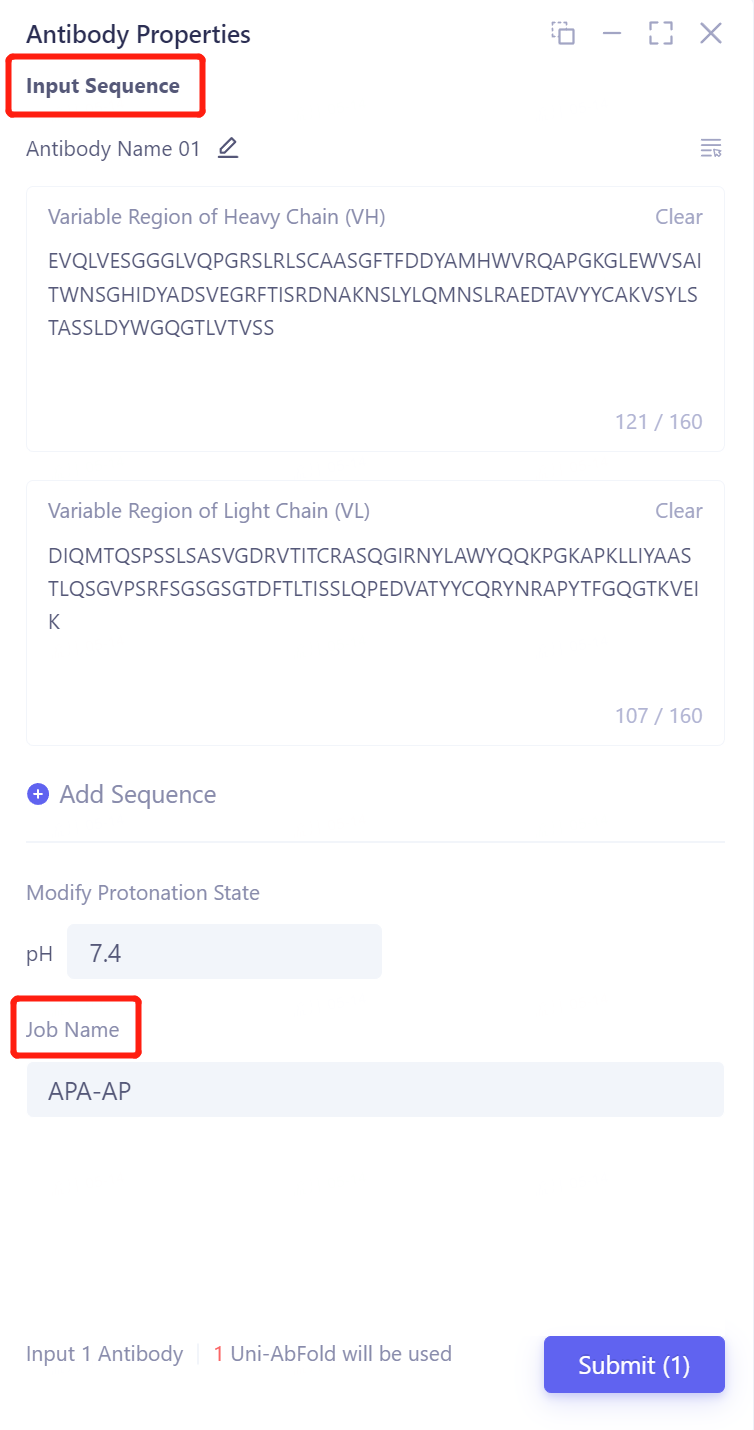
-
Note: The variable region sequence length of the imported antibody light chain/heavy chain should be less than 160.
-
Modify Protonation State: The protonation state of the antibody is changed by adjusting the pH. A default value of 7.4 is used here.
-
The job is named "ADA-AP" at the "Job Name".
-
Click "Submit" to submit the task.
2. Analysis of results
2.1 Entrance
-
General menu bar on the left: Job → Search for the required task.
- The task can be found by searching for the Job Name, or by filtering the Job Type.
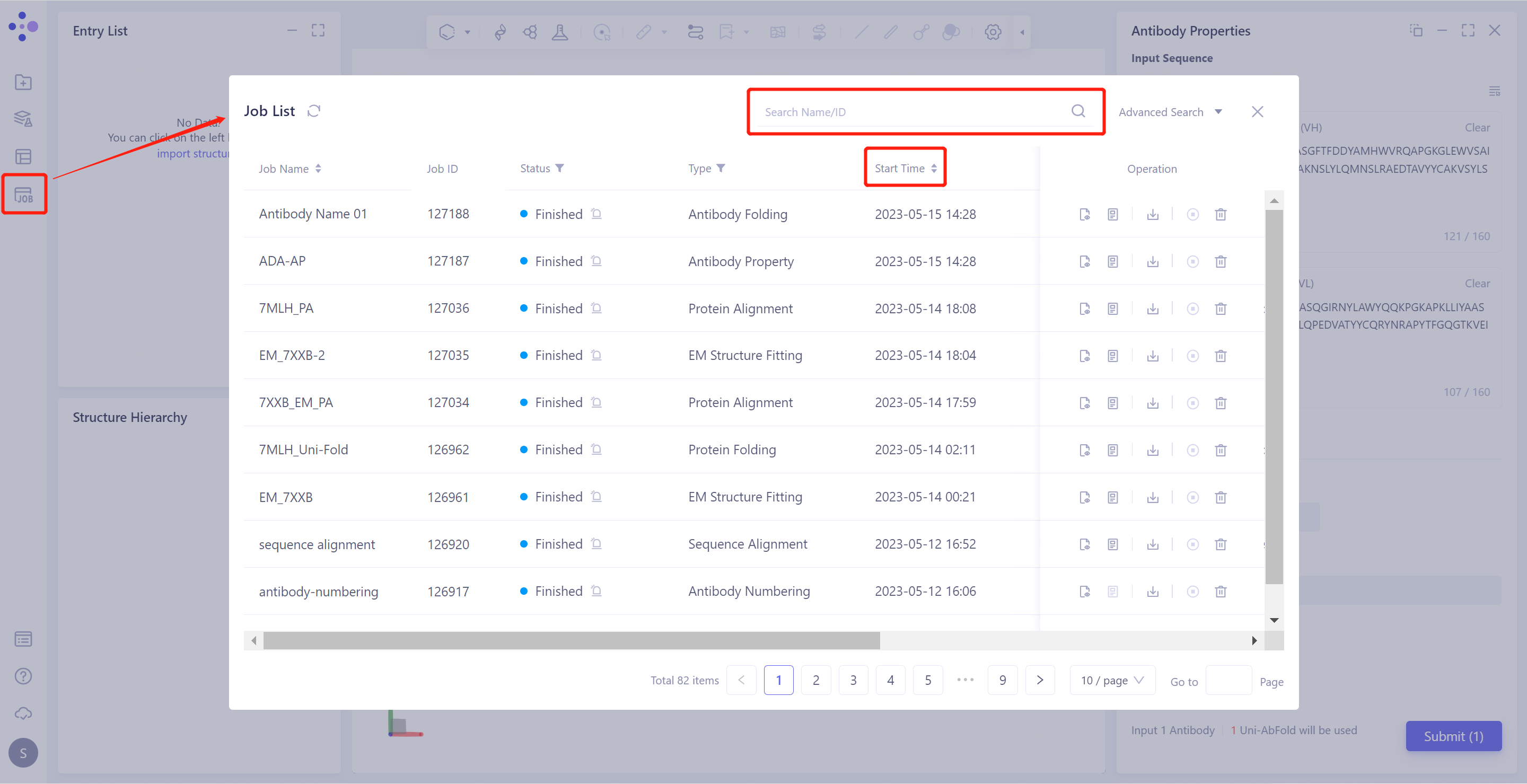
2.2 Results presentation
2.2.1 Result interface
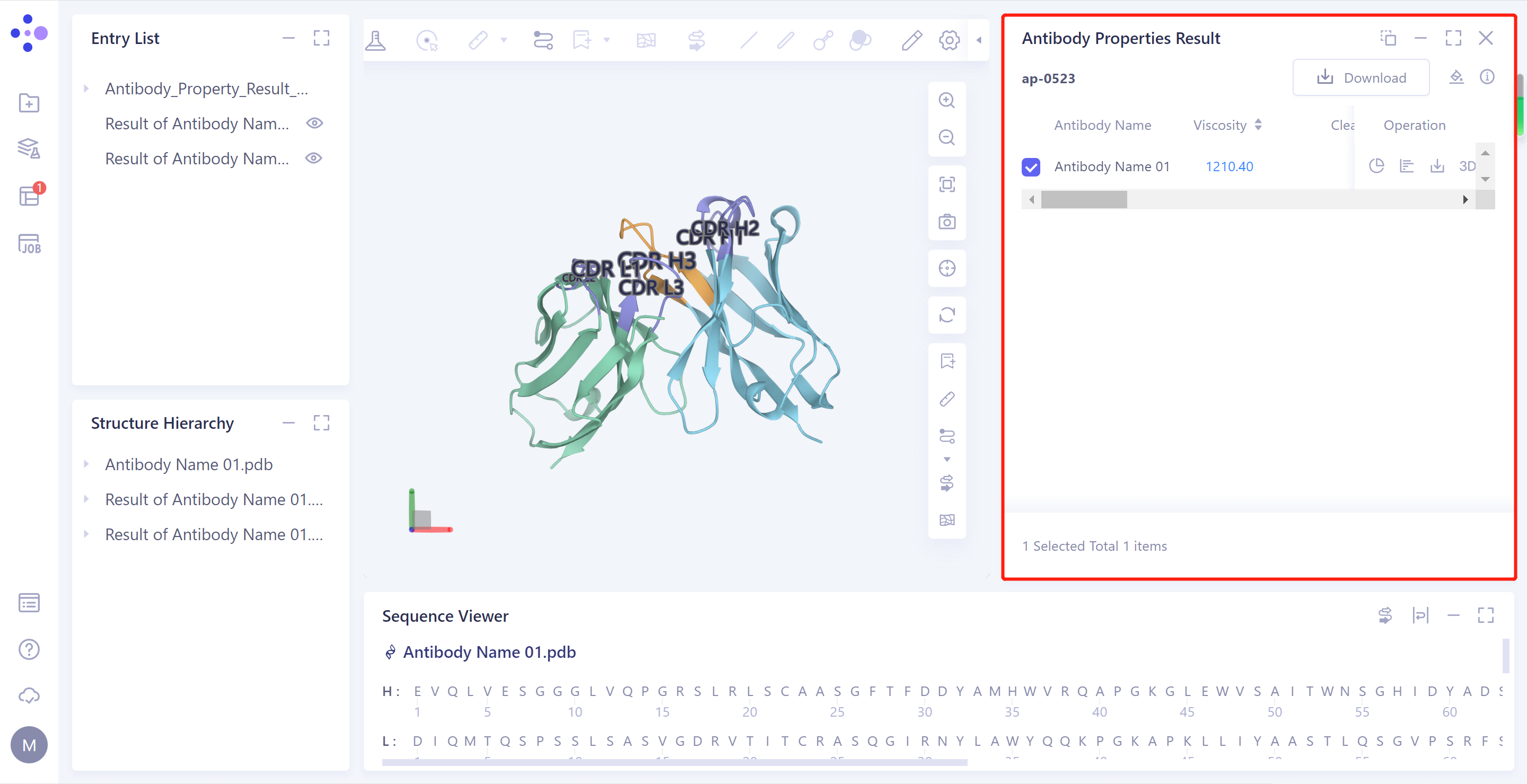
- Select the task to be viewed and click "Show" in the Operation column to display the result of the task, as shown in the figure below.
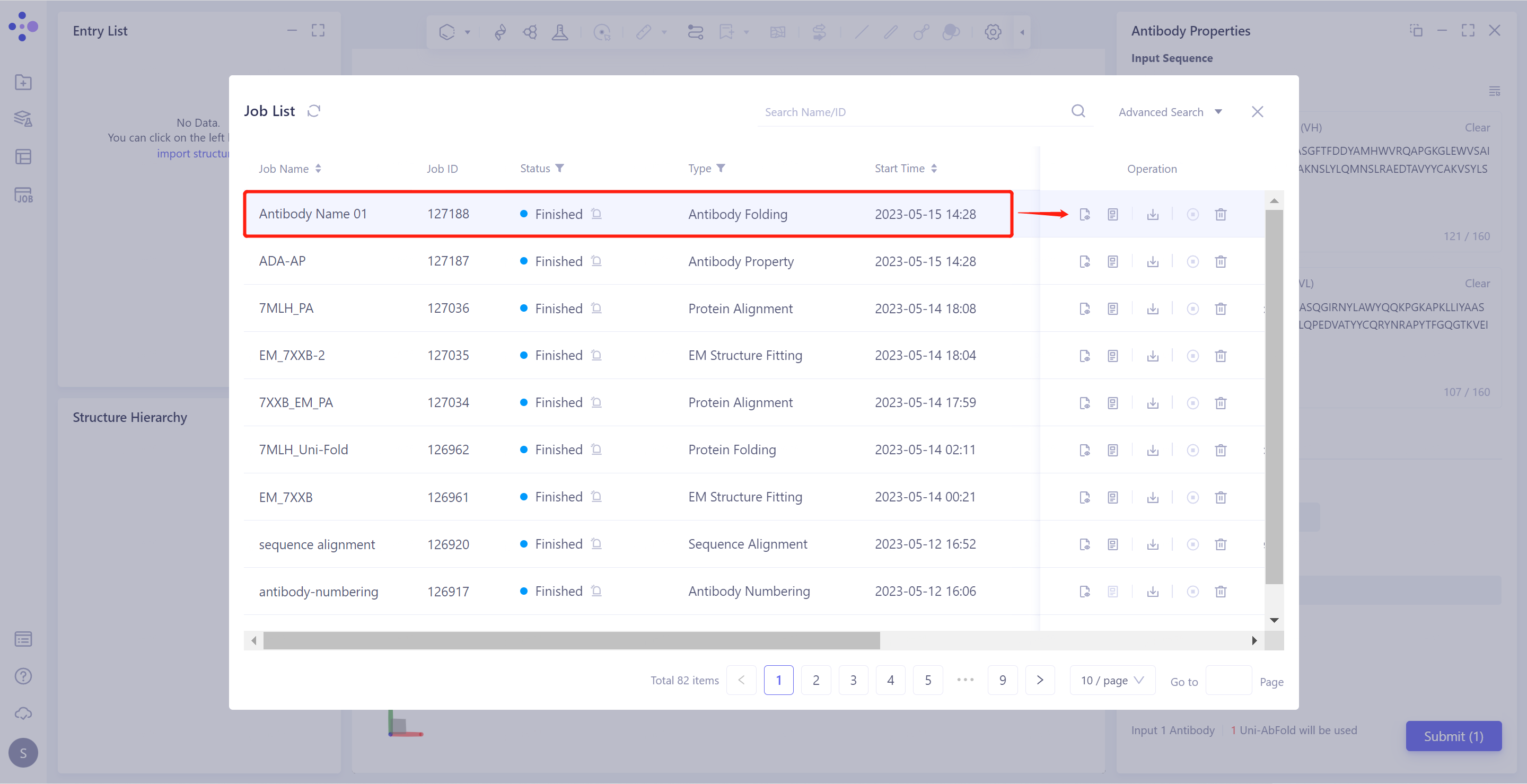
2.2.2 Explanation of results
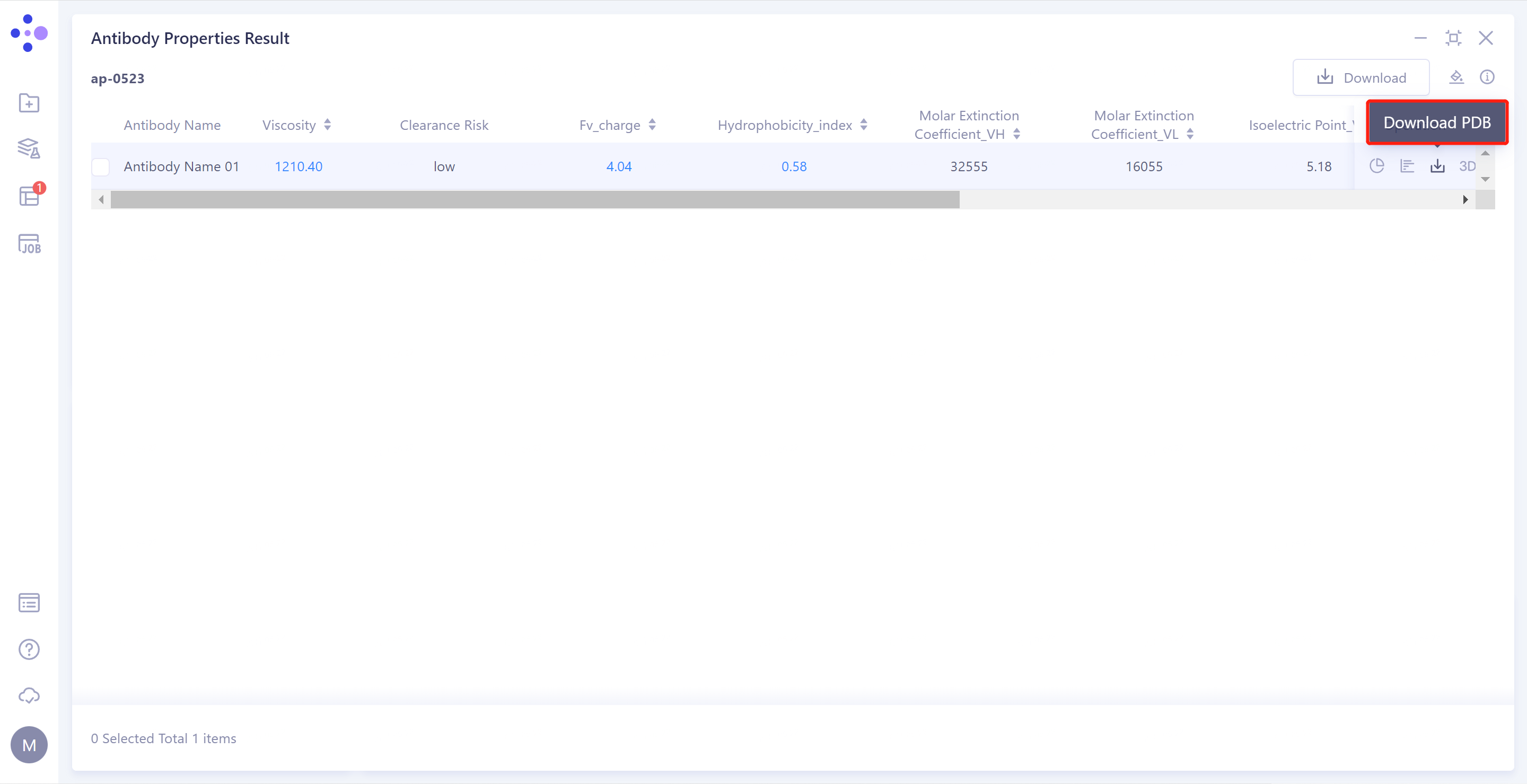
1)Antibody Properties Result: Antibody property prediction results are given in the list: Viscosity, Clearance Risk (Fv_charge and Hydrophobicity_index), Molar Extinction Coefficient_(VH/VL), Isoelectric Point_(VH/VL), Aggregation and TAP (PSH Score, PPC Score, PNC Score and FvCSP Score)
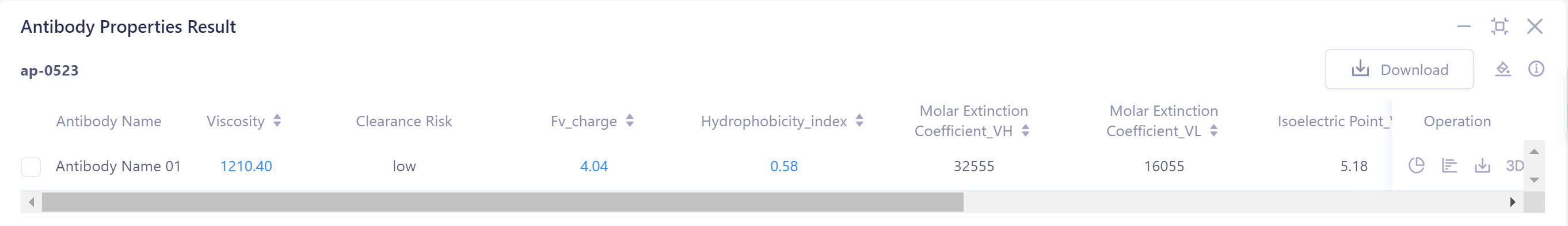
- The result can be judged by the threshold description in the upper right corner.
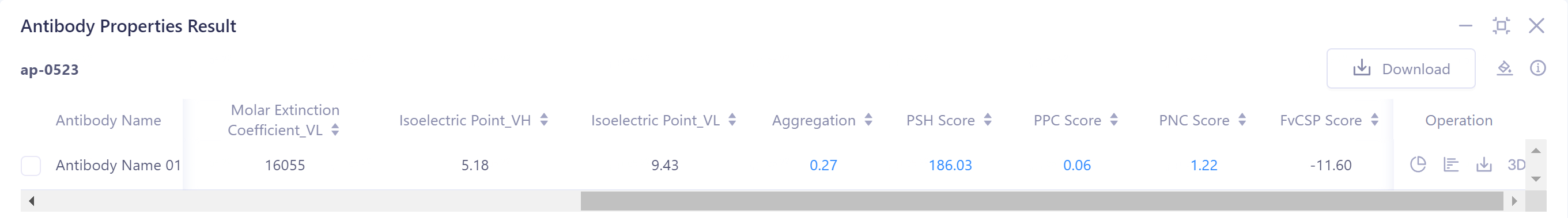
2)Download: Download the predicted structure file of the selected antibody, the table file of antibody properties, and the residue decomposition file (record the Aggregation, PSH Score, PPC Score, PNC Score, and FvCSP Score values for each residue on the antibody sequence).
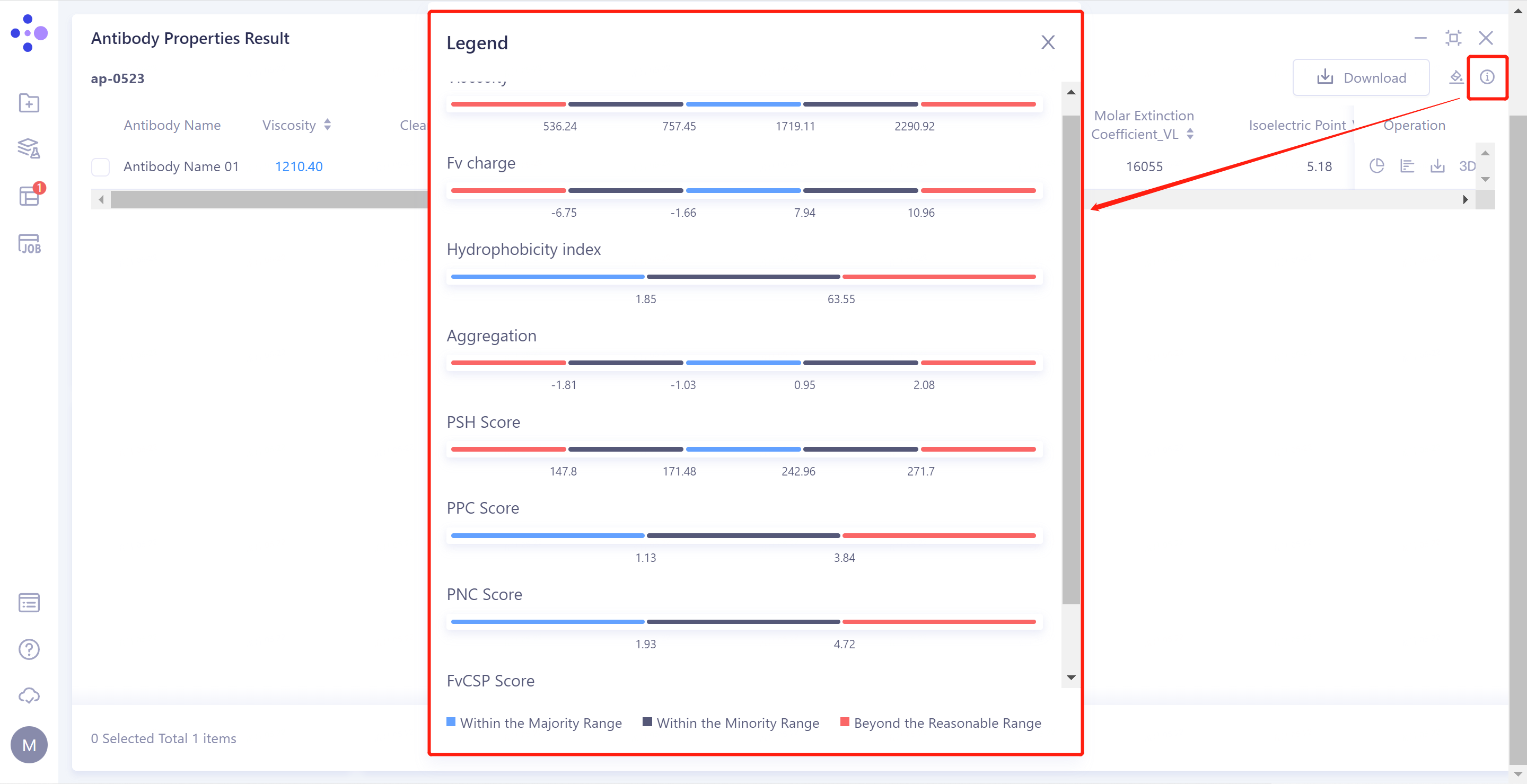
3)Operation :
- Decomposition: Click to display the list view, and record the values of Aggregation, PSH Score, PPC Score, PNC Score and FvCSP Score of each residue on the antibody sequence.
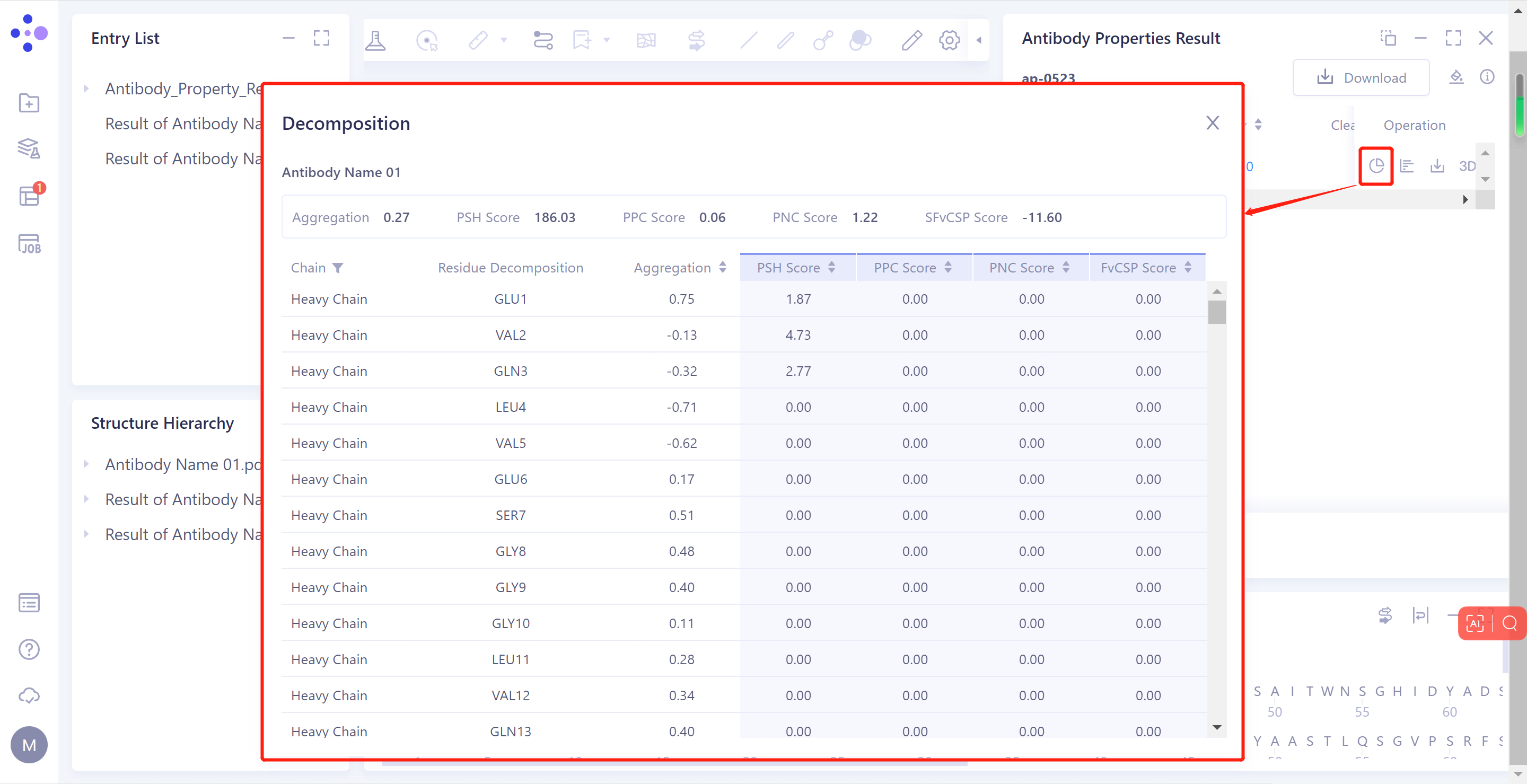
- Distribution: Click to display the chart, and record the distribution of the Aggregation, PSH Score, PPC Score, PNC Score and FvCSP Score of the antibody in the antibody database.
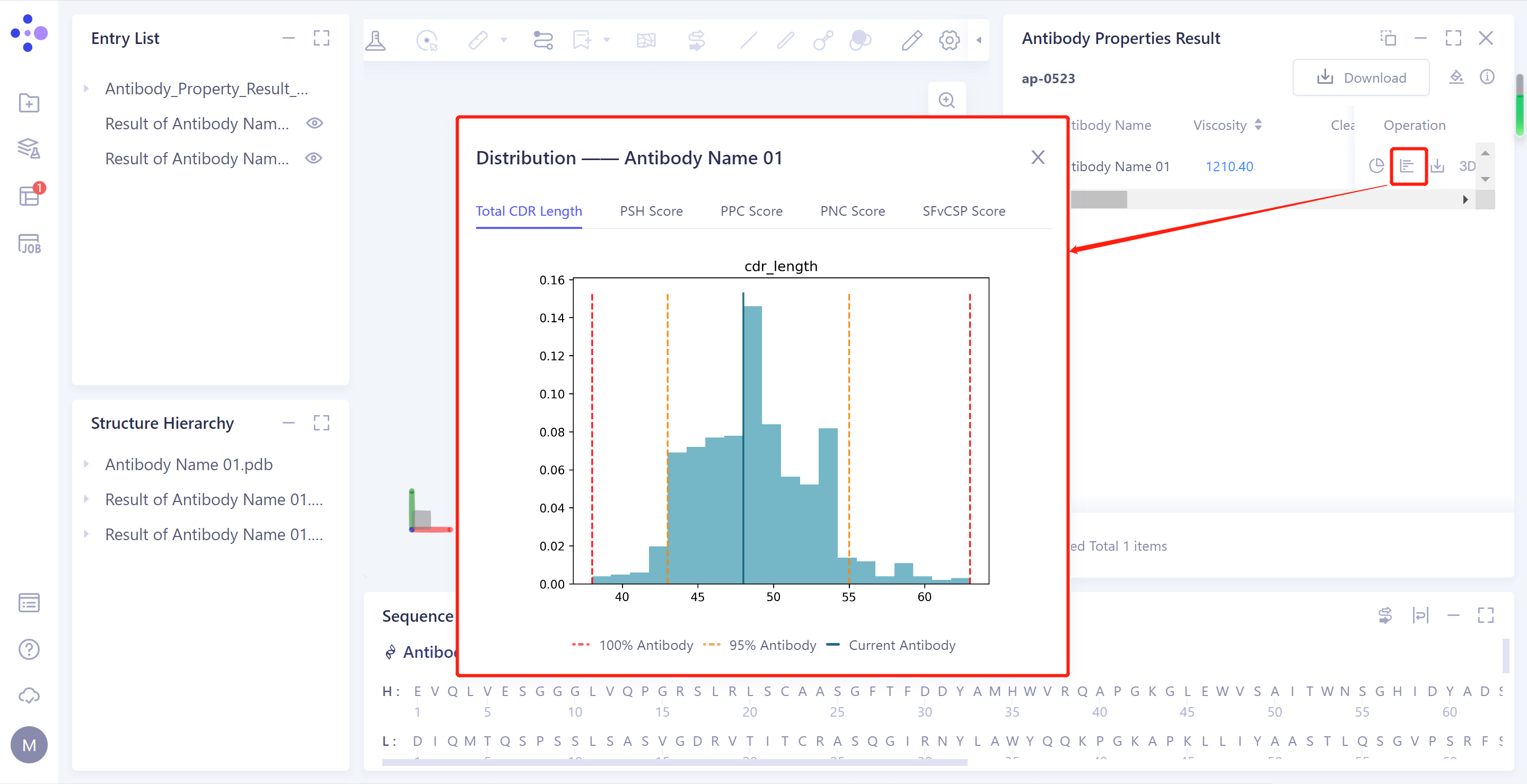
- Download PDB: Download the predicted structure file for the selected antibody.
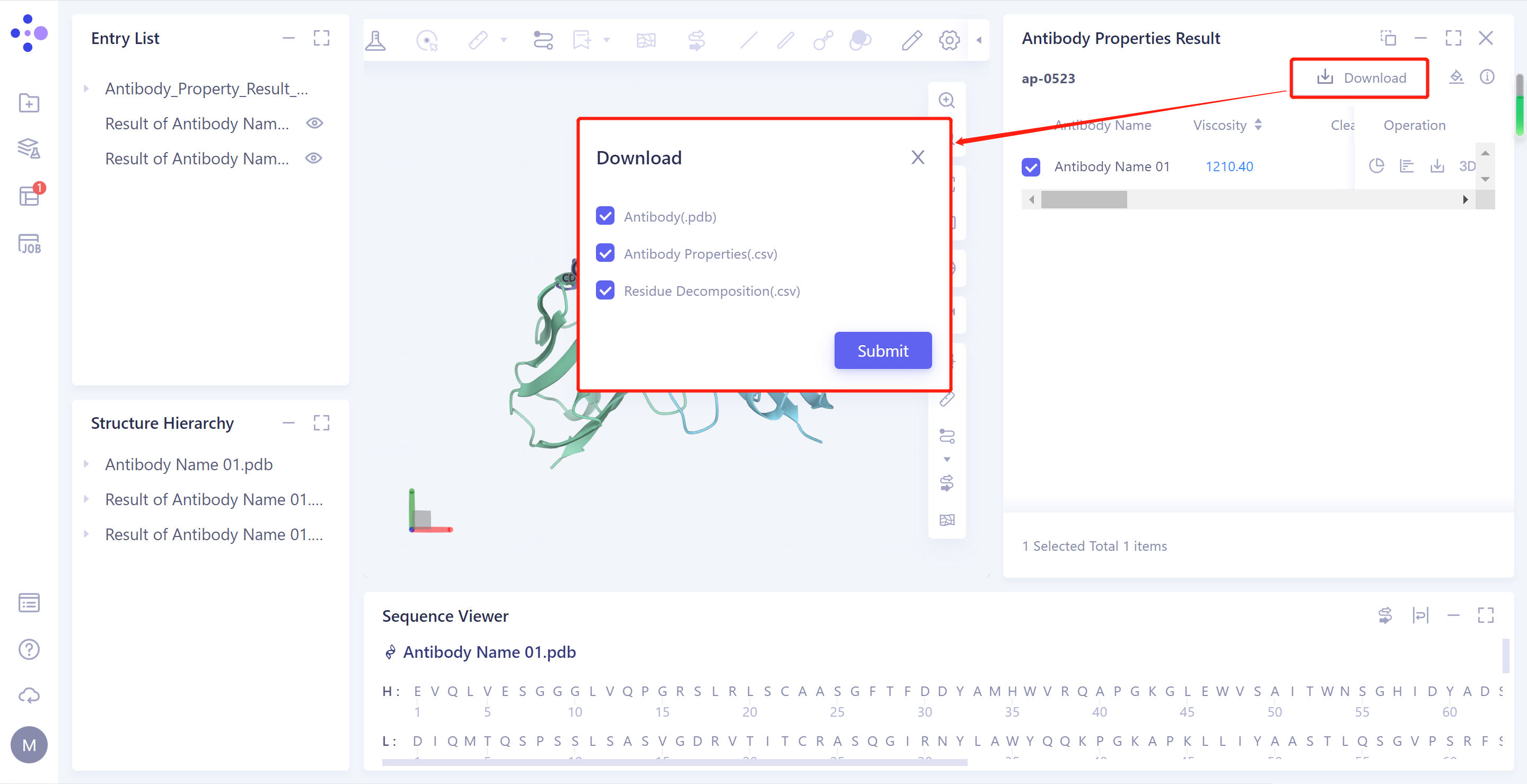
- The 3D: antibody structure is displayed in the 3D Workspace window.
2.3 Analysis of prediction results
-
Viscosity: The prediction result is 984.29, indicating that the viscosity is appropriate.
-
Clearance Risk: The prediction result is low, indicating a low risk of antibody clearance.
-
Molar Extinction Coefficient_(VH/VL): The molar extinction coefficients reflecting the heavy/light chains of the antibody are 32555 and 16055, respectively.
-
Isoelectric Point_(VH/VL): indicated that the isoelectric points of the heavy/light chains of the antibody were 5.18 and 9.43, respectively.
-
Aggregation: The prediction result is 0.27, indicating that there is no solubility difference caused by aggregation effect.
-
Decomposition: The contribution of each residue in the variable region of an antibody to aggregation can be demonstrated.
-
In the TAP prediction result, in the "Distribution", the values of the four properties of the antibody to be tested are distributed in the statistical interval, indicating that the properties of the antibody to be tested are reasonable.
2.3.1 Parameter description
-
Viscosity: indicates the viscosity of the antibody
-
750 ~ 1800: the viscosity of antibody is appropriate;
-
530 ~ 750 and 800 ~ 2200: the antibody viscosity is acceptable;
-
Less than 530 or more than 2200: the antibody viscosity is unacceptable and needs to be modified.
-
-
Clearance Risk: refers to the antibody clearance rate, which is determined by the combination of Fv_charge and Hydrophobicity_index.
- Clearance Risk: refers to the antibody clearance rate, which is determined by the combination of Fv_charge and Hydrophobicity_index.
-
Clearance Risk can be classified as low, medium, or high risk. When the risk is high, the antibody needs to be modified, and when the risk is medium, the antibody modification is recommended.
-
Fv_charge: When pH = 5.5, the net charge of antibody variable region (Fv) is appropriate when Fv_charge is between 0 and 6.2.
-
Hydrophobicity_index: when pH = 5.5, the hydrophobicity index of antibody LC1 + LC3 + HC3 ≤ 4.
-
-
Molar Extinction Coefficient_(VH/VL): Molar extinction coefficient of heavy chain/light chain
-
Isoelectric Point_(VH/VL): isoelectric point of heavy chain/light chain
- Molar Extinction Coefficient_(VH/VL): Molar extinction coefficient of heavy chain/light chain
- Isoelectric Point_(VH/VL): isoelectric point of heavy chain/light chain
-
Aggregation: indicates antibody aggregation
-
When it is greater than 0, there is no problem of poor solubility caused by aggregation effect;
-
-1 ~ 0, there may be some solubility problems caused by aggregation effect, but it is not serious;
-
When it is less than -1, the solubility problem caused by aggregation effect may be more serious, and it is recommended to modify it.
-
Note: The contribution of each residue in the antibody variable region to the aggregation can be shown by the Decomposition under the Operation on the right.
-
-
The TAP is composed of four indexes, and records the statistical distribution of these indexes of the antibody in the antibody drug database. Generally speaking, when the values of these four properties of the antibody to be tested are distributed in the statistical interval, it is considered that the properties of the antibody to be tested are reasonable. The distribution of the antibody under these four properties can be displayed more intuitively through the Distribution operation under the Operation on the right side;
-
PSH Score: surface hydrophobicity index near CDR;
-
PPC Score: positive charge indicator near CDR;
-
PNC Score: negative charge indicator near CDR;
-
FvCSP Score: Charge symmetry parameter of antibody variable region structure.
-
Note: This function supports multiple antibody property prediction.
-
3. References
[1] Cooke H A, Arndt J, Quan C, Shapiro R I, Wen D, Foley S, Vecchi M M, Preyer M. EFab domain substitution as a solution to the light-chain pairing problem of bispecific antibodies [J]. mAbs, 2018, 10: 1248-1259.