Compound Clustering
Introduction
Compound clustering refers to the aggregation of compounds with similar chemical structures and properties. In the fields of chemistry, drug discovery and biology, compound clustering is widely used in tasks such as data mining , classification, pattern recognition and activity prediction.
The Compound Clustering module of the Hermite platform provides the compound clustering function, which realizes compound clustering based on the widely recognized K-Means algorithm and DBSCAN algorithm. At the same time, it provides a massive compound library built into the Hermite platform, which helps you expand the chemical space of compound clustering.
This tutorial is based on the Compound Clustering module of the Hermite platform for clustering 30 compounds into 5 classes.
The documents required for this tutorial are as follows:
1. Usage
1.1 Entrance
The left general menu bar Function → Molecular Recommendation → Compound Clustering .
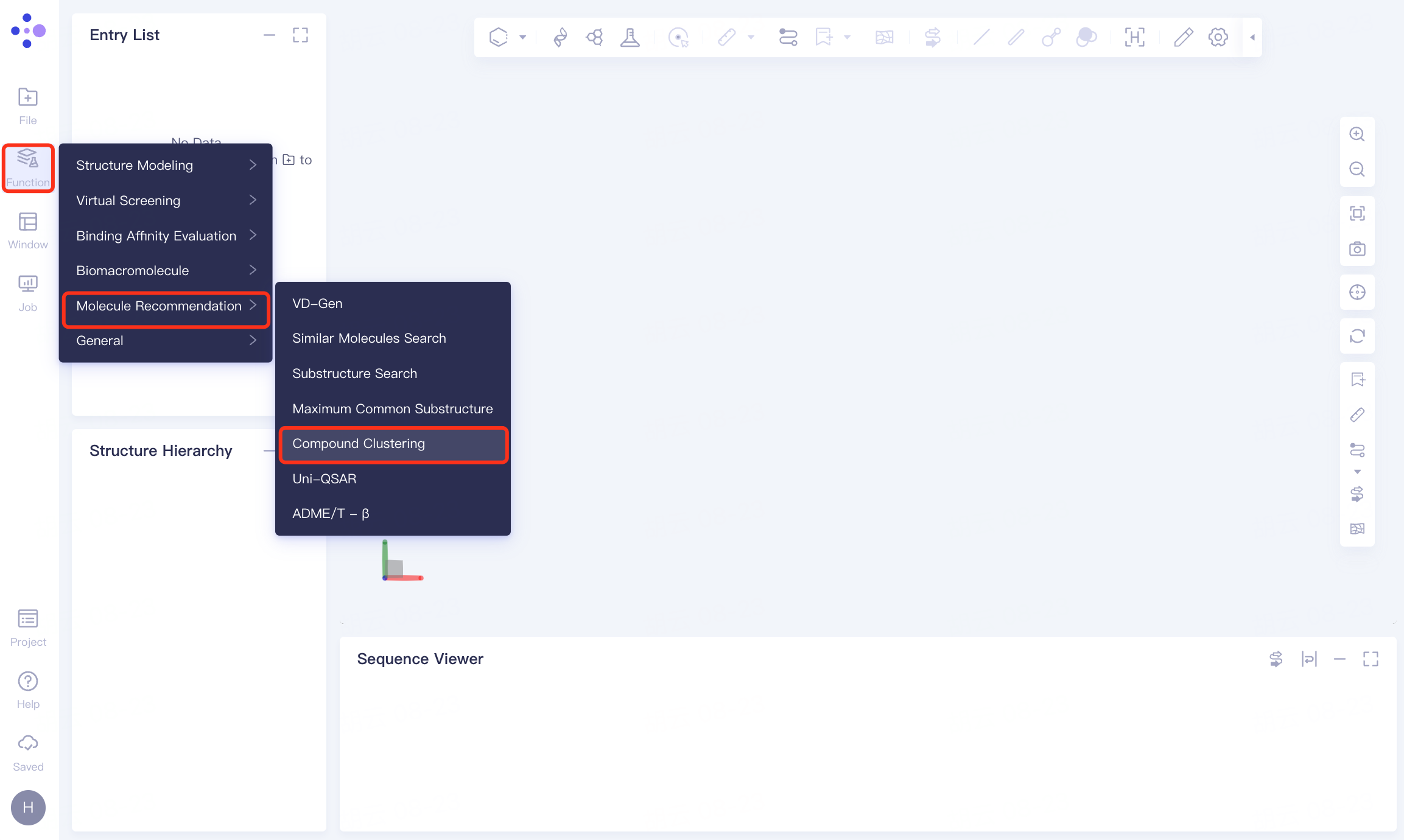
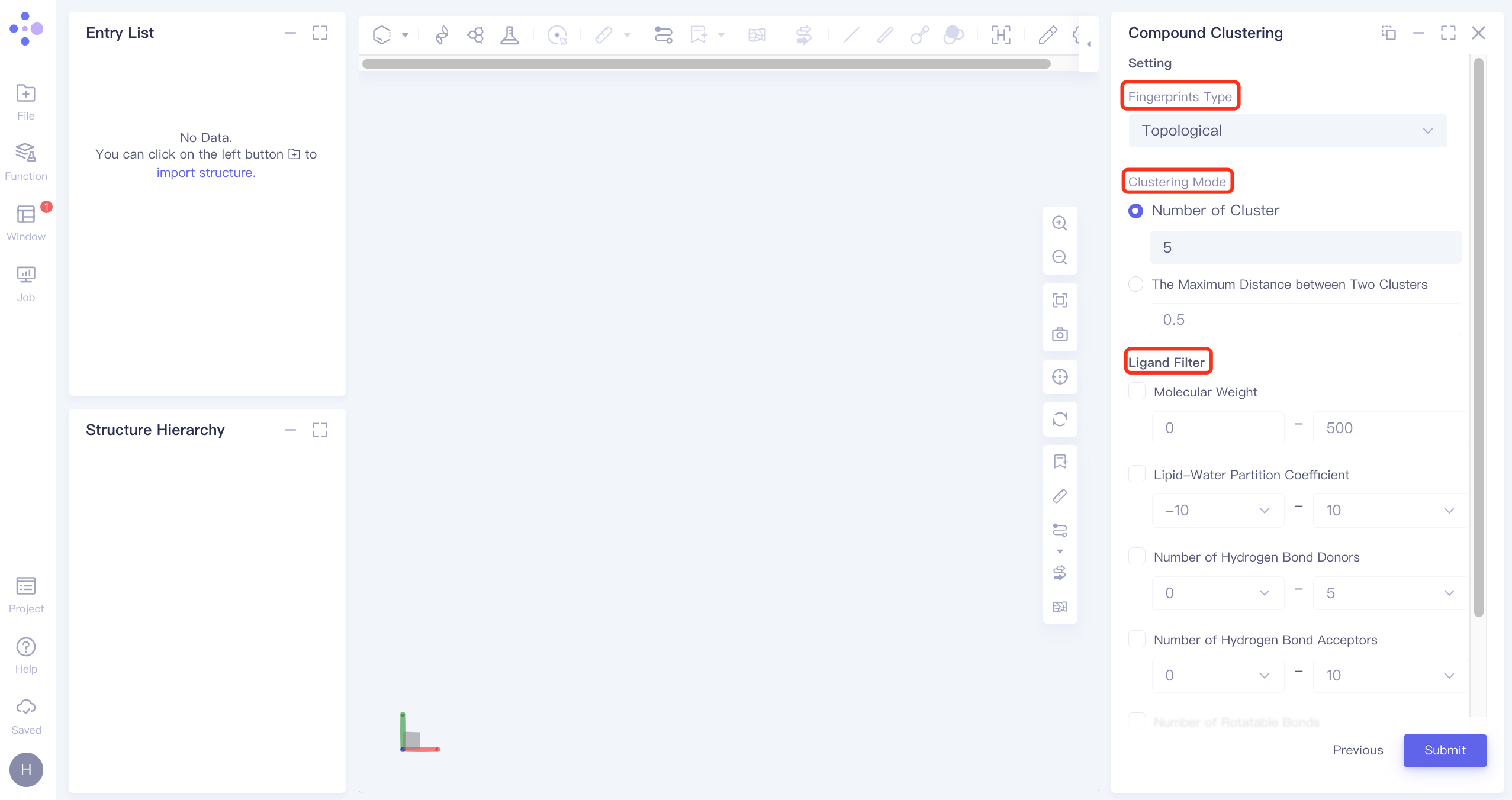
The operation box of Compound Clustering appears on the right (shown in the red box), and the overall interface is as follows:
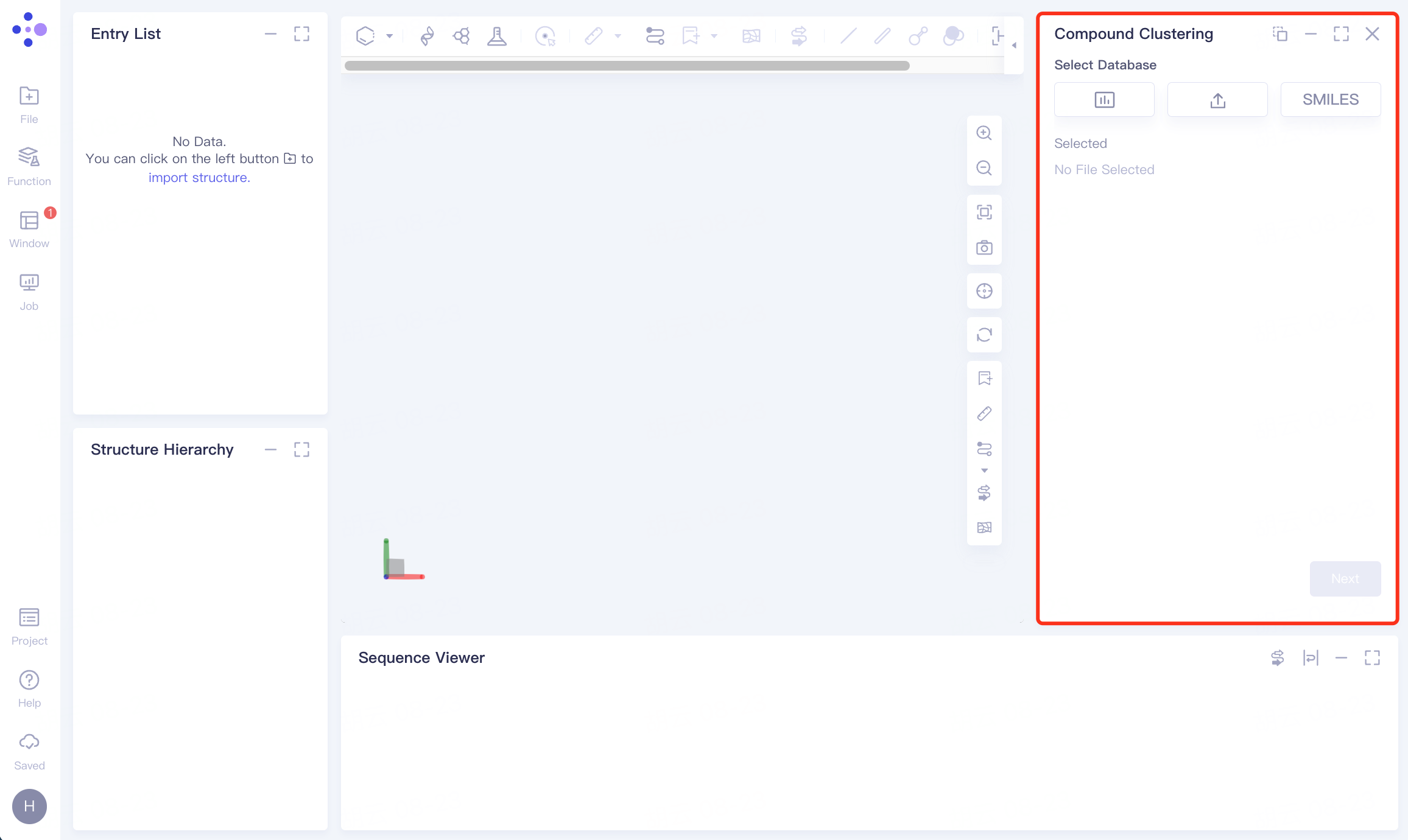
1.2 Upload molecular dataset
Select File
Click the "Select File" checkbox → Select CompoundClustering.smi file from the local folder and upload it.
Click "Next" to enter the next operation.
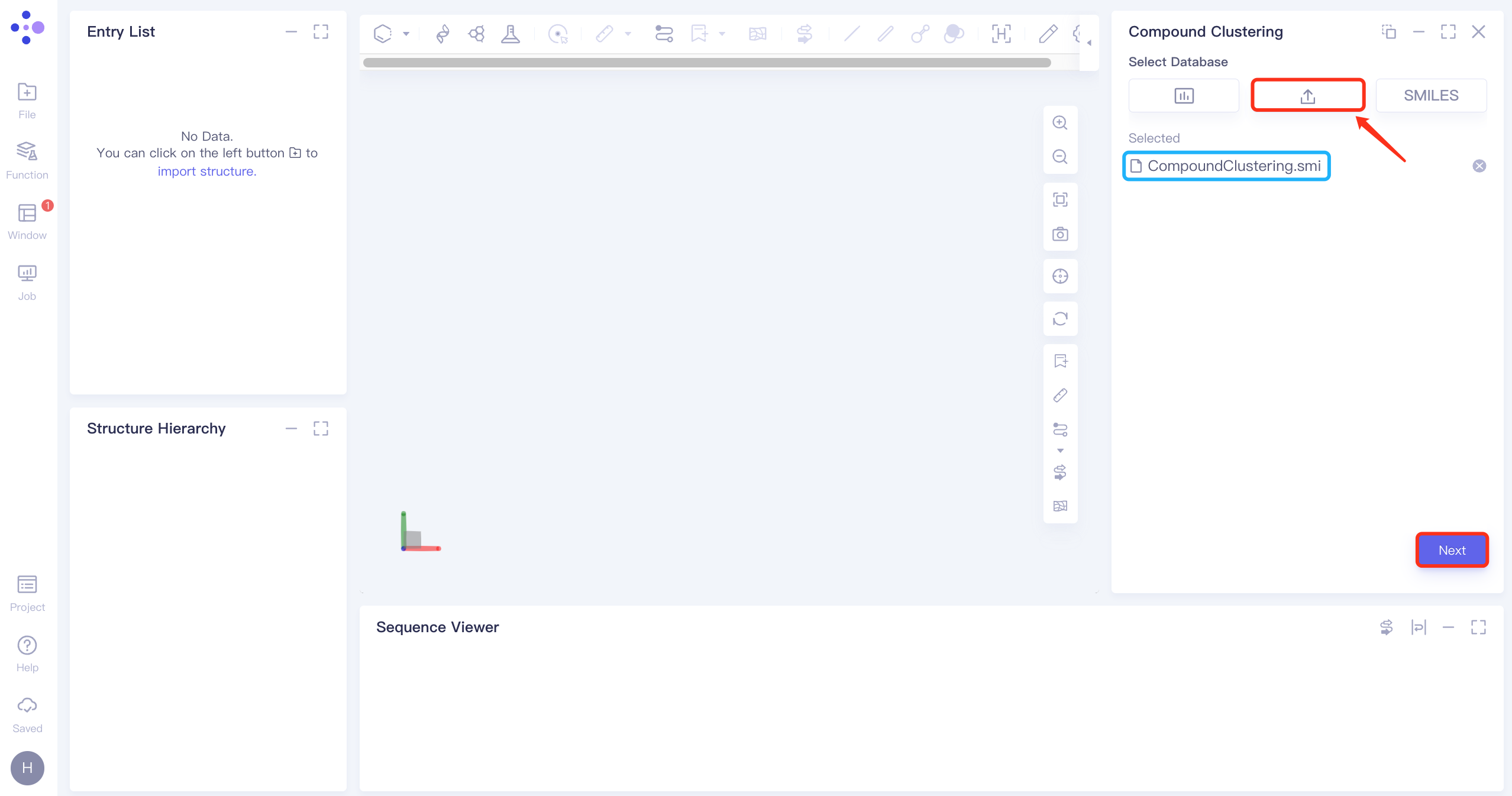
1.3 Settings
Fingerprint Type selects "Topological", and molecular features are extracted by RDKit's topological fingerprint;
Clustering Mode ( Clustering Mode) Select Number of Clusters and set the number of classifications to 5;
Ligand Filter is not set, do not filter molecules;
Name the task "30compounds" at Job Name.
Click "Submit" to submit the task.
2. Results analysis
2.1 Entrance
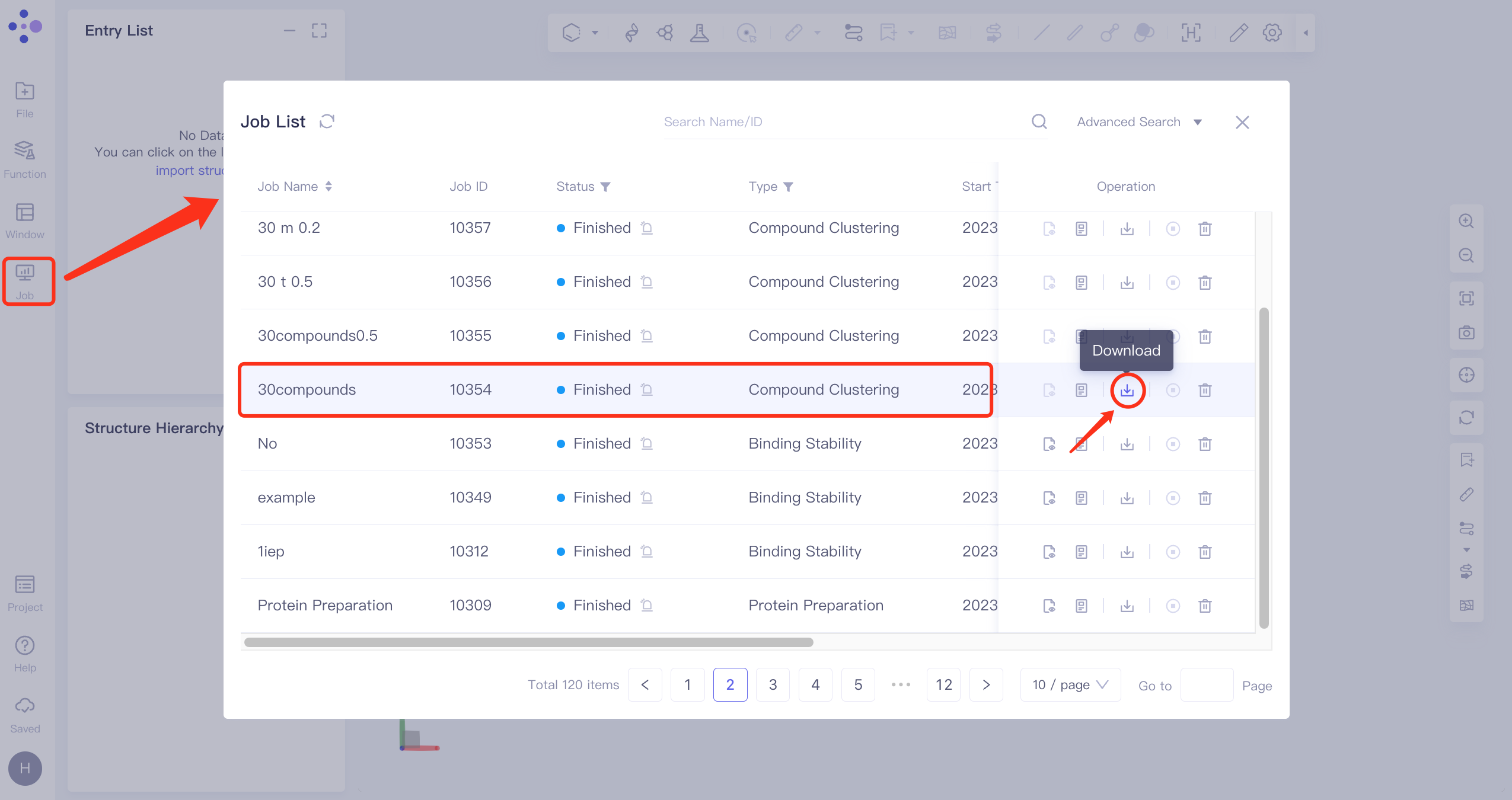
In the left general menu bar Job → find the task 30compounds → click Download in the Operation column to download the result of the task.
2.2 Check the results
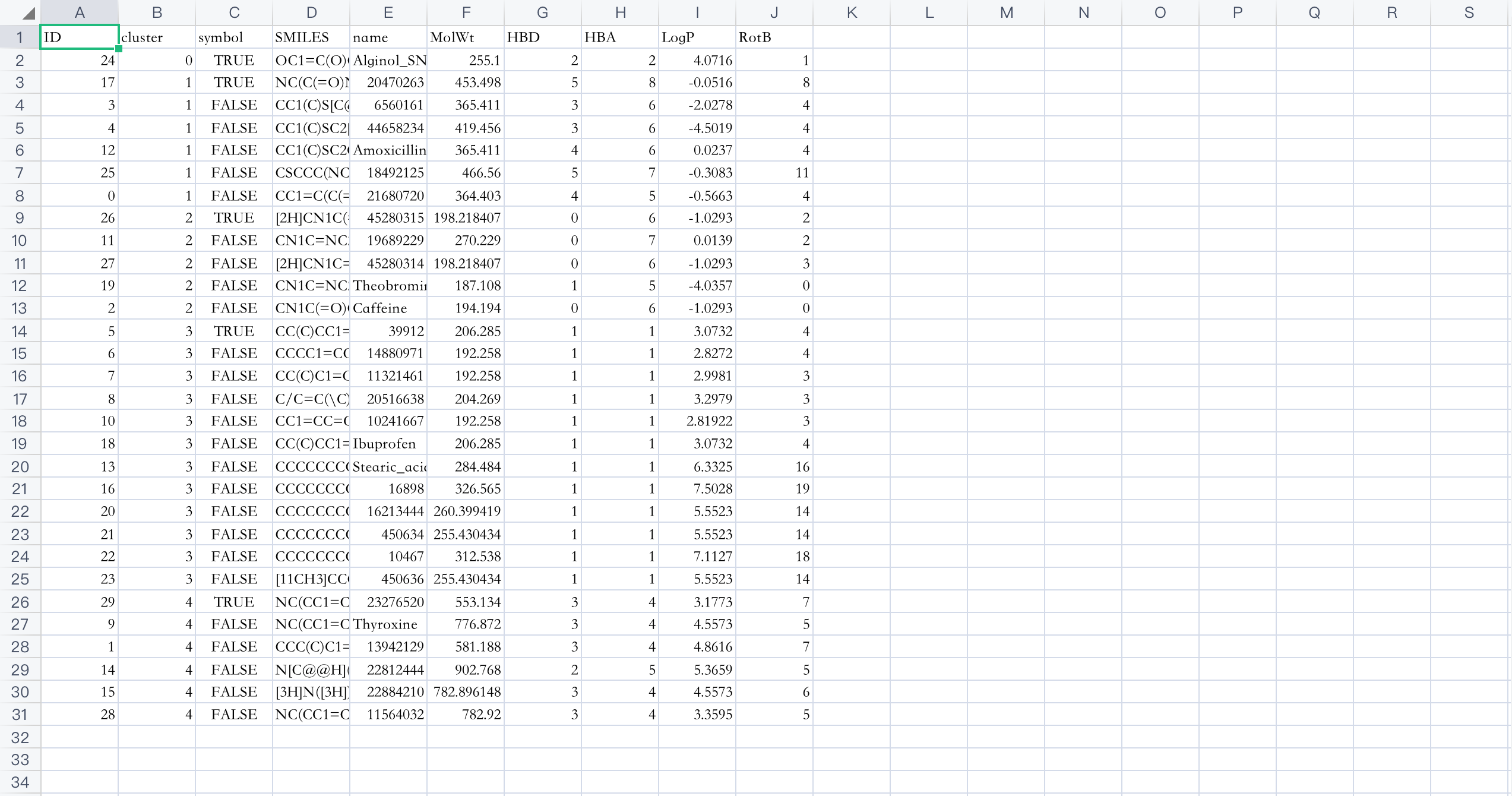
Download and get a .csv file, open the file to see the results:
According to the number labeling in the Cluster column, it can be seen that these 30 compounds are correctly divided into 5 categories. In addition, the calculated MolWt (molecular weight), HBD (number of hydrogen bond donors), HBA (number of hydrogen bond acceptors), LogP (lipid-water partition coefficient) and RotB (number of rotatable bonds) can help us screen the drug-like properties of compounds.