FEP Protein Mutation
Introduction
Proteins are one of the most important molecules in living organisms. They play various roles in life, including catalyzing chemical reactions, transmitting signals, and supporting cell structures. The structure and function of a protein are usually determined by its amino acid sequence, and mutations can change the amino acid sequence, thus affecting the structure and function of the protein. Thermal stability is one of the important indicators of protein stability, which has a great impact on the application and industrial production of proteins. Therefore, it is very important to predict the thermal stability of proteins after mutation.
The FEP Protein Mutation module of Hermite ® platform combines free energy perturbation theory, molecular dynamics simulation and high-performance computing, which can accurately and quickly predict the changes of protein thermal stability after mutation, and help researchers quickly evaluate the impact of protein mutation on its thermal stability. To guide the design and optimization of protein structure and function
In this tutorial, we will learn to use the FEP Protein Mutation module of the Hermite ® platform to accurately and rapidly predict the change in the binding free energy of various amino acid mutations in Tenascin [1] [2]. It helps researchers to quickly assess the impact of protein mutations on their thermal stability and guide the design and optimization of protein structure and function.
1. Create a project and import the structure
1.1 Log in to the system
- Login address: https://hermite.dp.tech

1.2 Create the project
- Once in the system, create a new project FEP Protein Mutation ”.
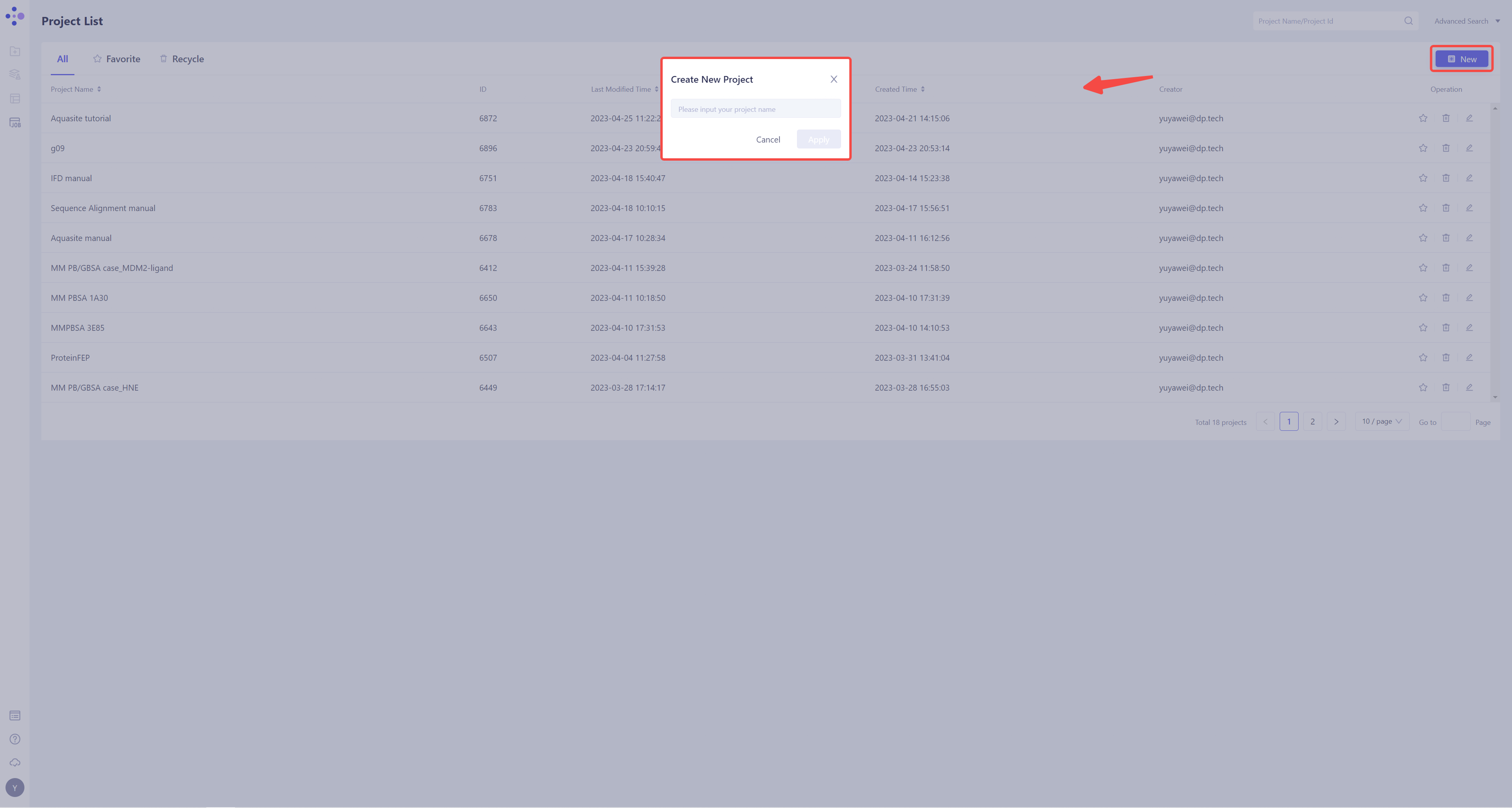
1.3 Import Protein Structure
- Left general menu bar Menu → File → Import Structure
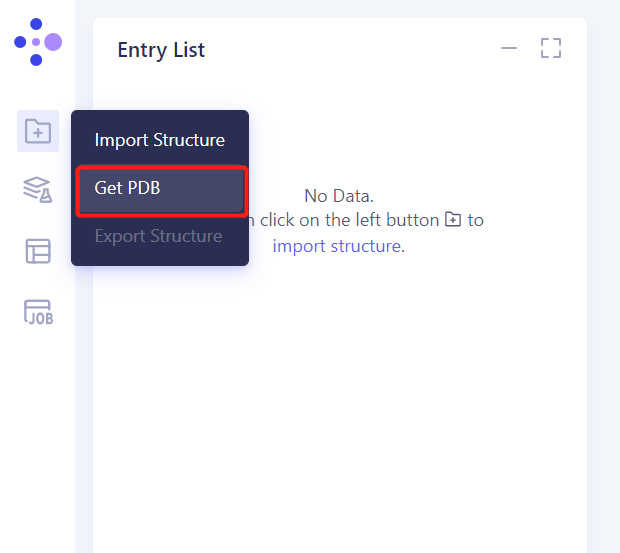
- Click Get PDB, input 1 TEN file, and click Import to import the protein structure
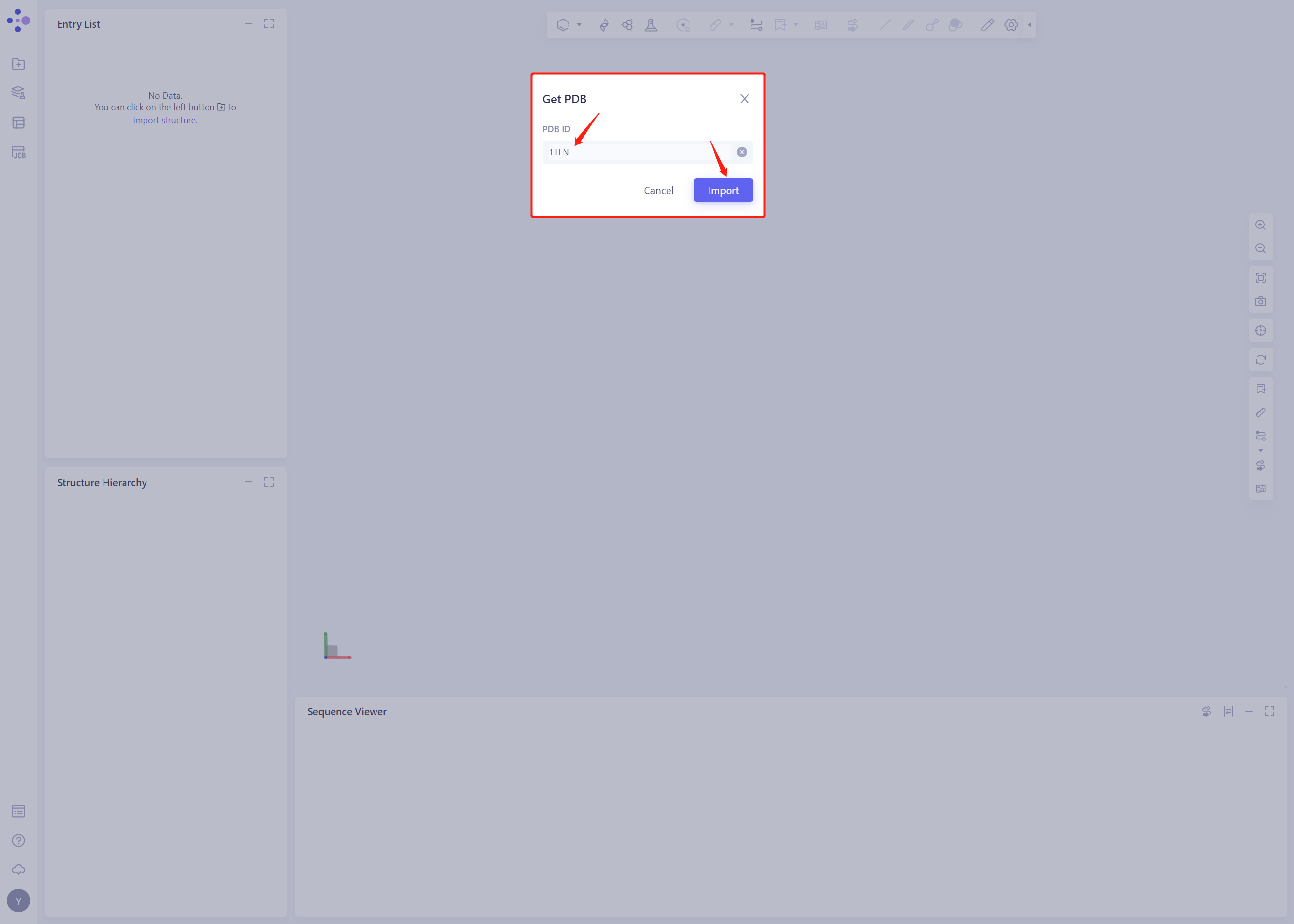
2. System preparation
2.1 Prepare the protein structure
2.1.1 Select Structure
- Left general menu bar Function → Structure Modeling → Protein Preparation
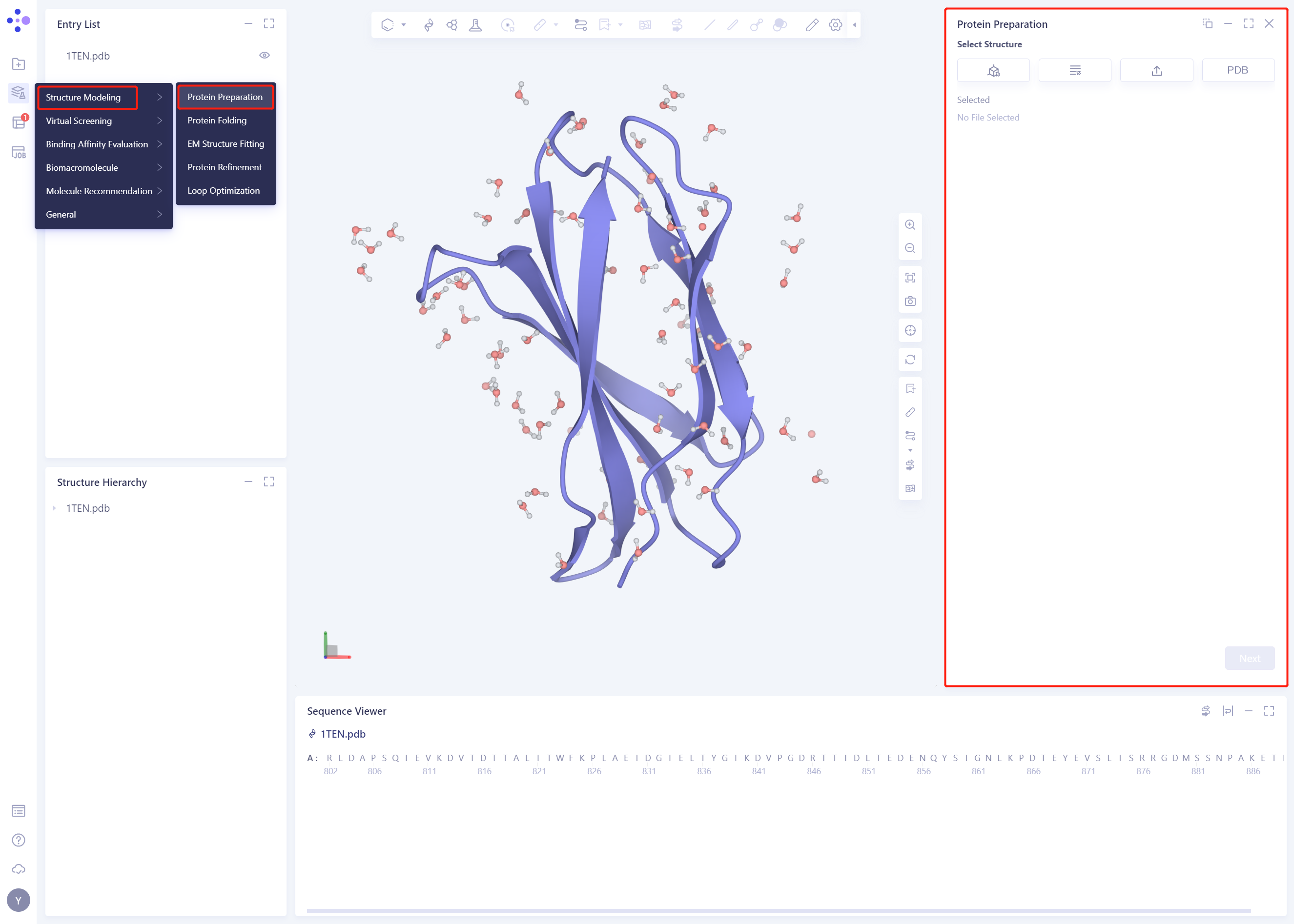
- Select Structure from 3D Works pace: Select Protein in Structure Hierarchy and click Ok
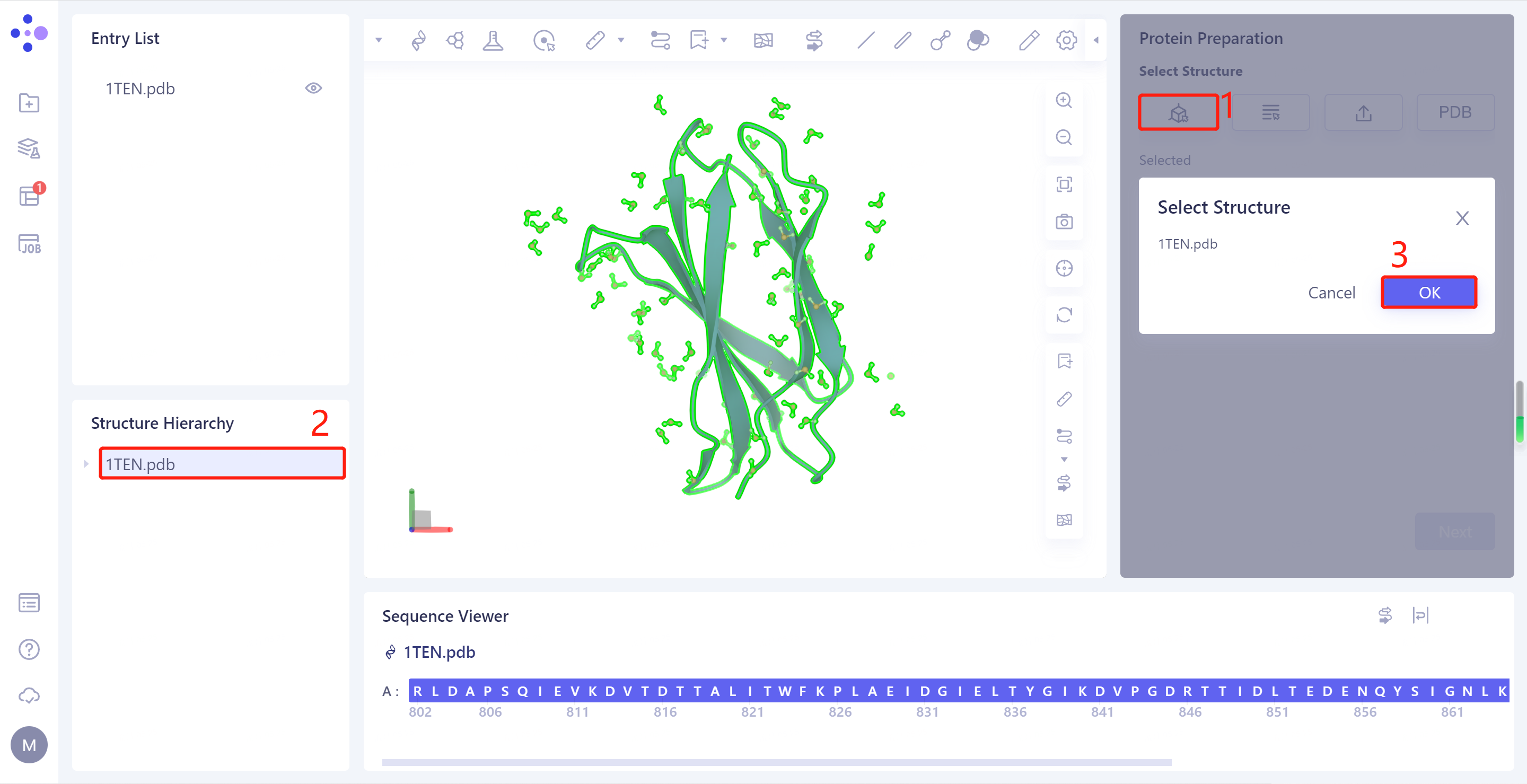
- 1TEN.pdb is loaded to the Protein Preparation parameter setting panel, click Next
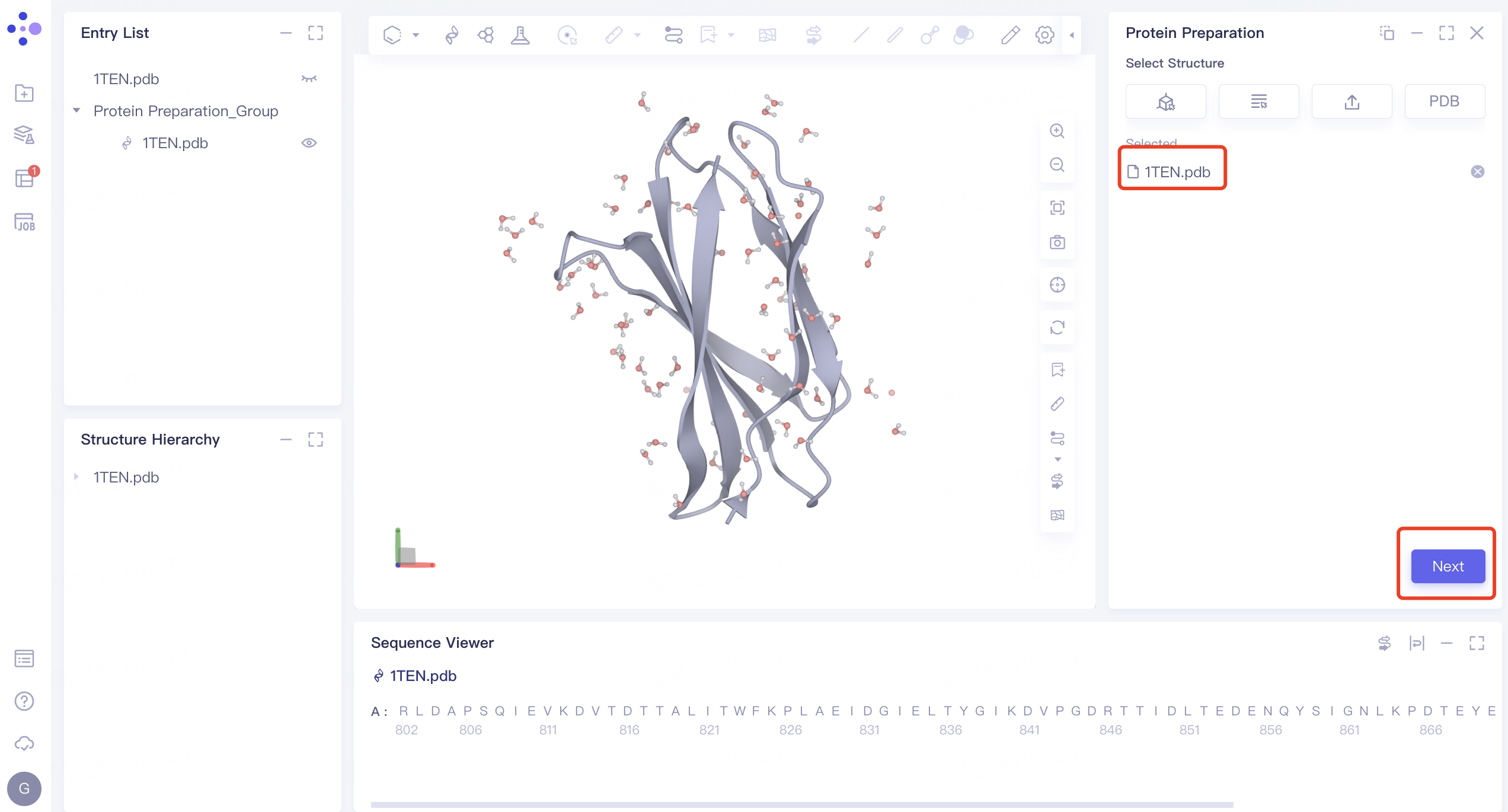
2.1.2 Alternative Position:
- Select the default amino acid position and click Next
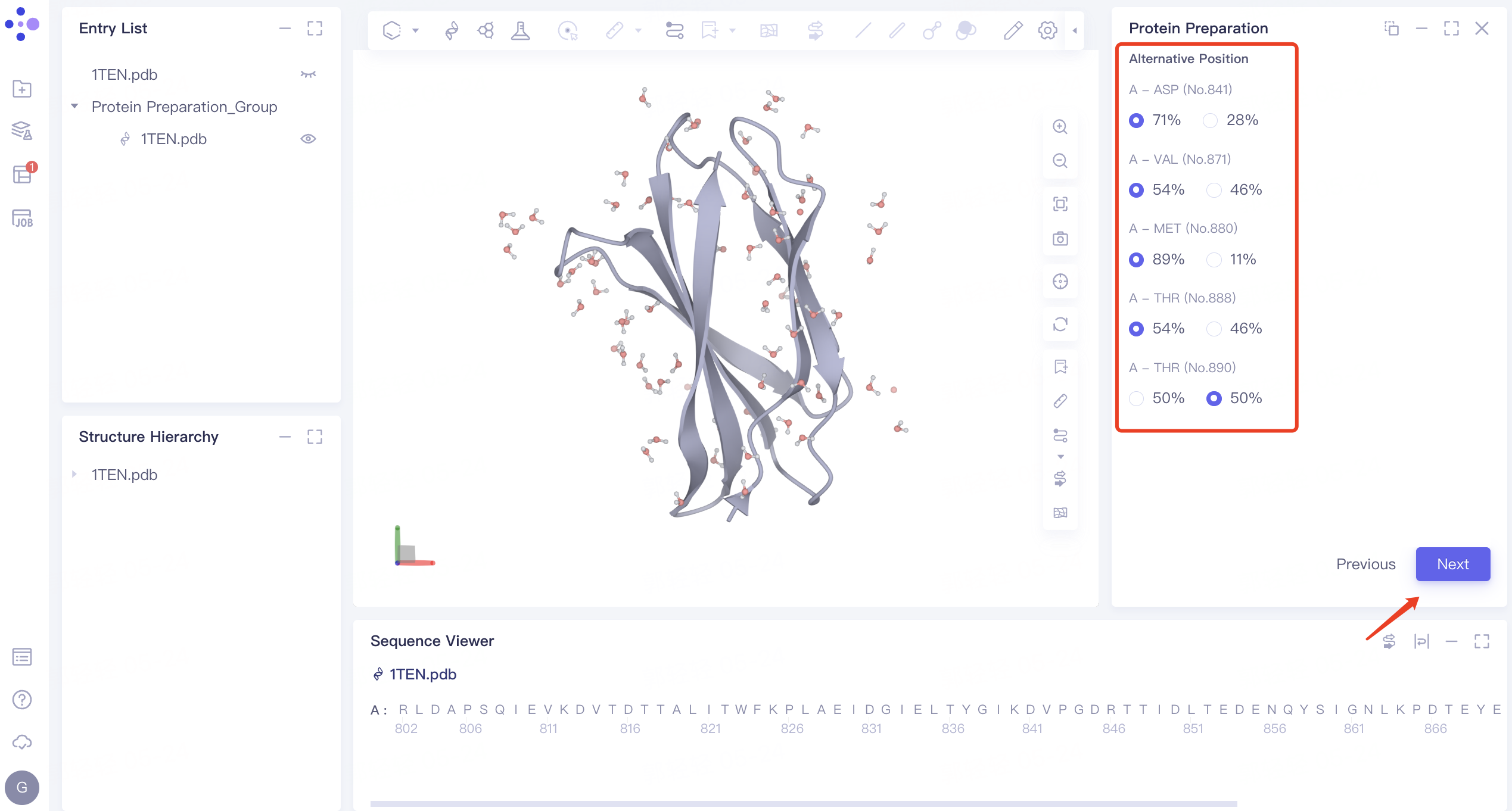
2.1.3 Select Polymer、Other Groups to Keep:
- Select the A-chain protein and A-chain water, and click Next
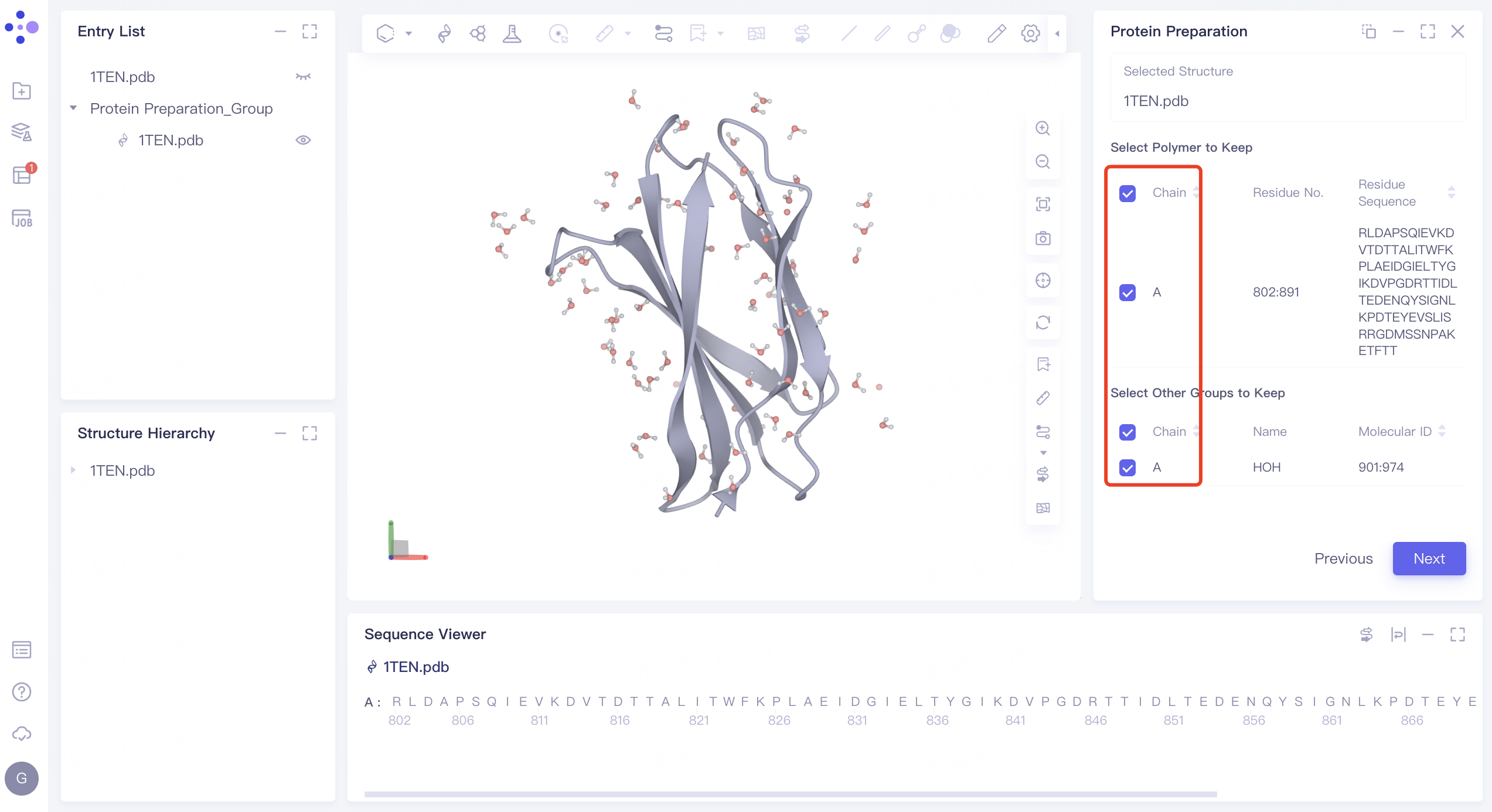
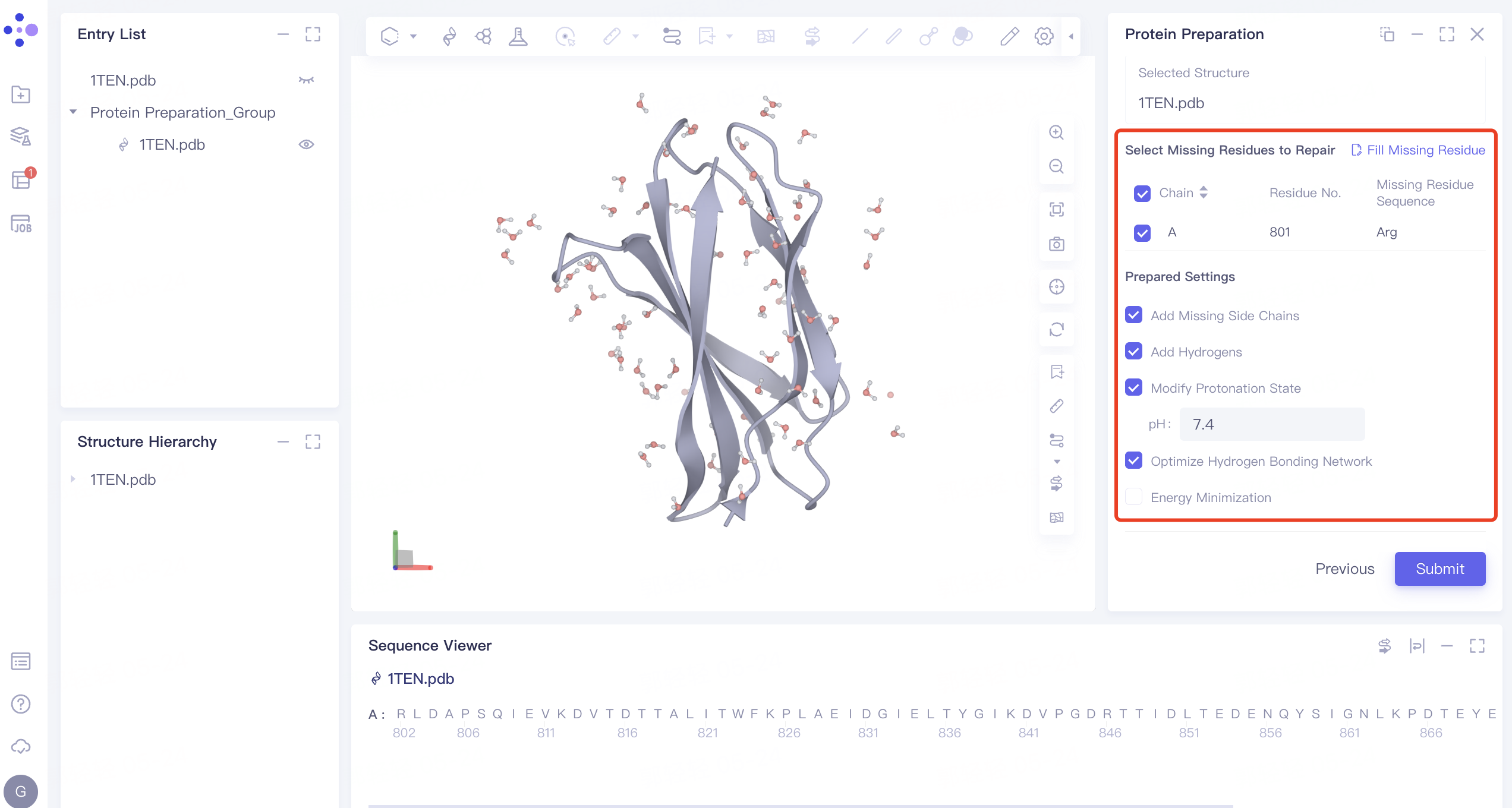
2.1.4 Select Missing Residues to Repair
- Check A-chain Missing Residues
2.1.5 Prepared Settings
- Uncheck “Energy Minimization” ”, do not optimize the energy of the crystal structure, and keep the other parameters as default
2.1.6 Name the Job and Submit the Task
- The Job Name is “1ten _ Prepare”. Click “Submit” to submit the job
2.1.7 View Protein Preparation Results
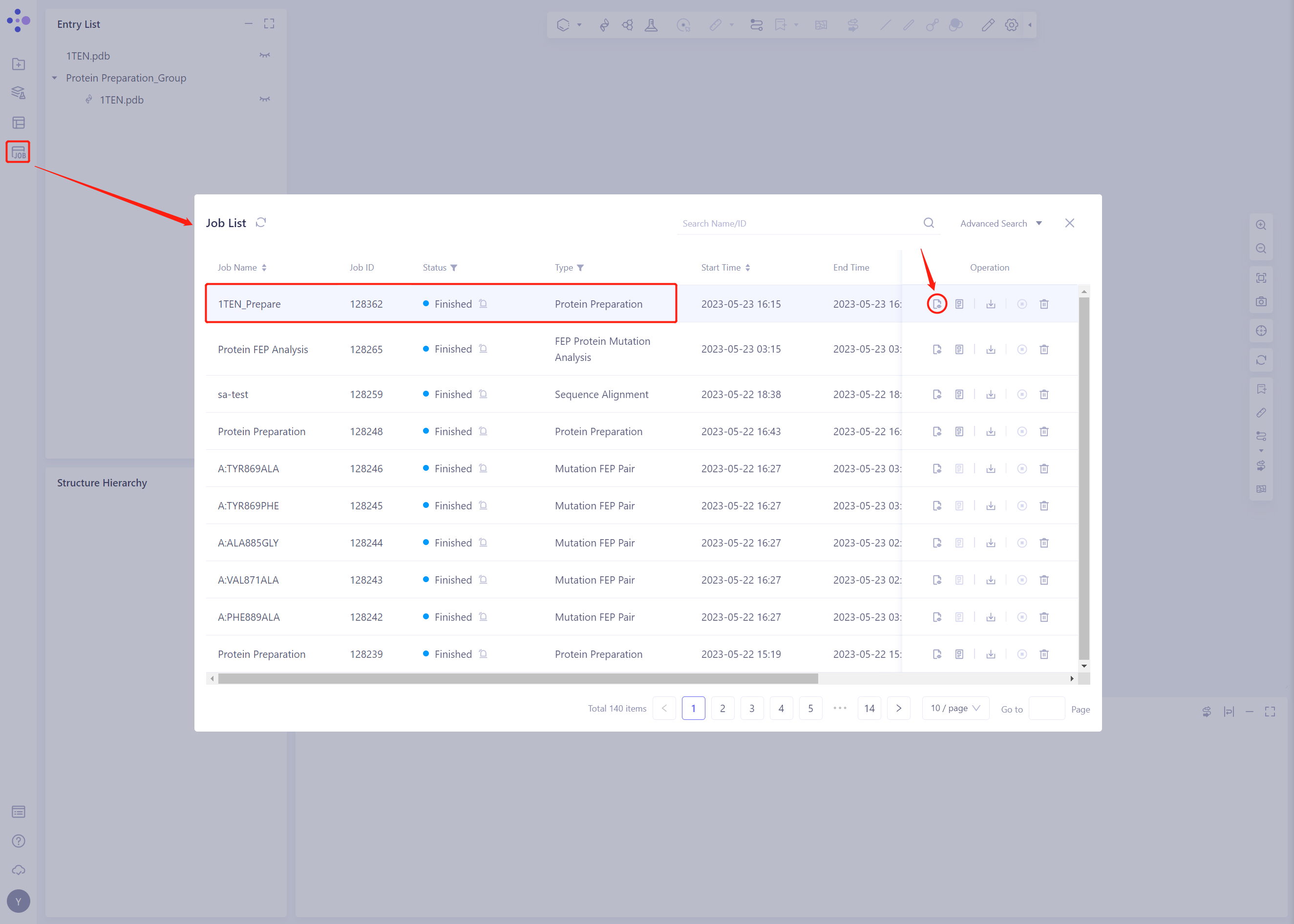
- Protein Preparation computing tasks are generally completed in a dozen seconds to a few minutes. After the task is completed, click Jobs to view the corresponding task;
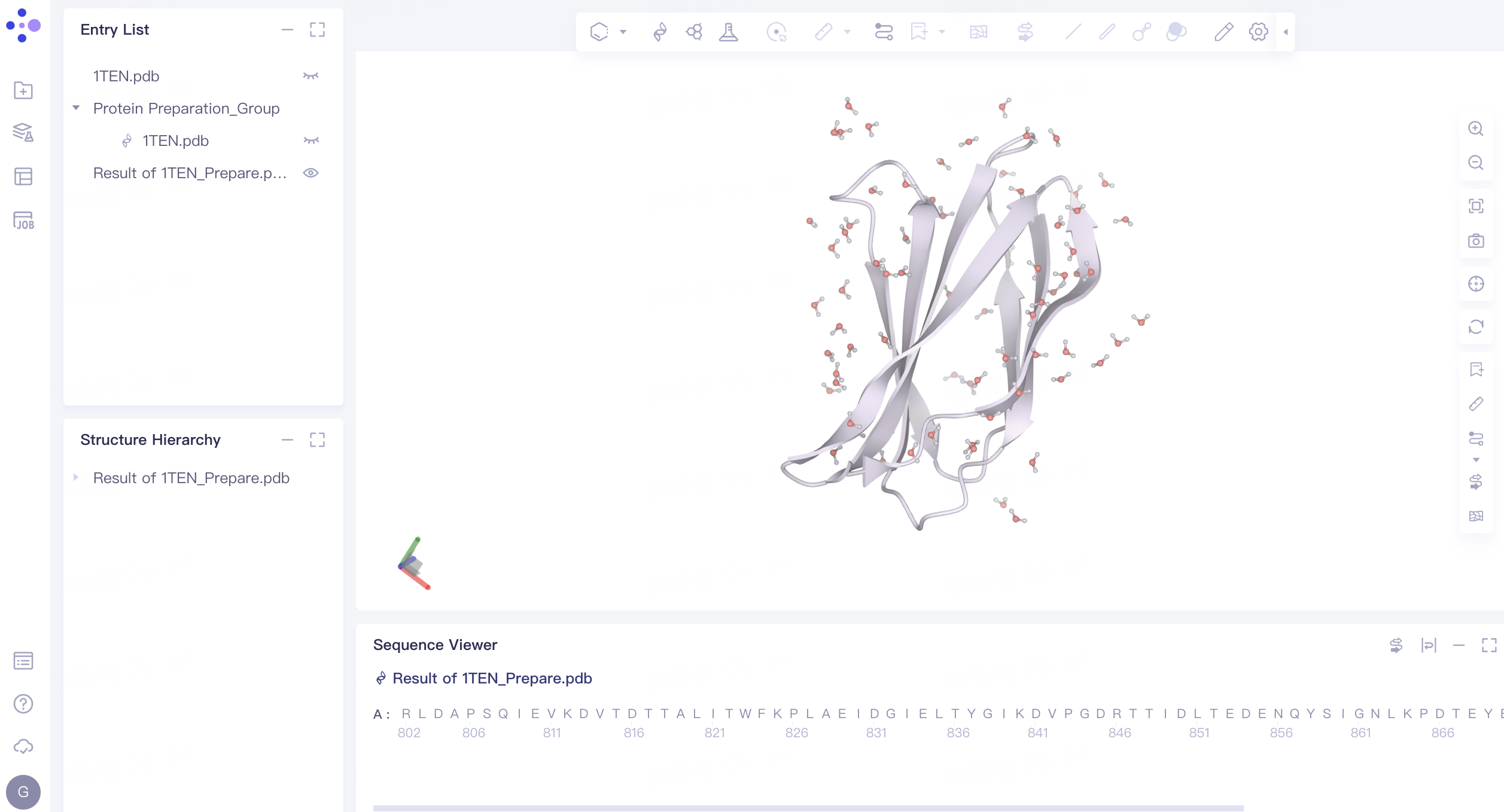
- Click the “show” button to display the prepared protein structure in 3D Workspace
3. Create FEP Protein Mutation Task
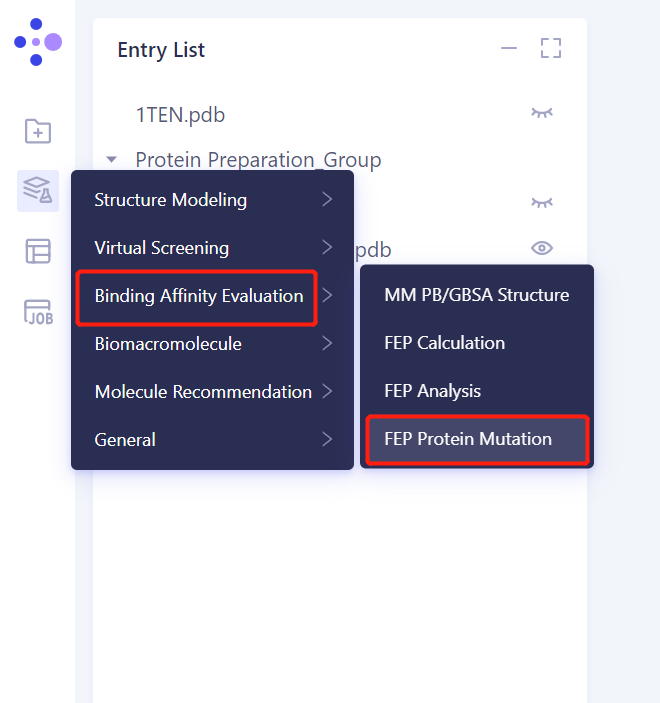
3.1. Left general menu bar Menu → Function → Binding Affinity Evaluation → FEP Protein Mutation
3.2 Select Prepared Protein From Project
- Select “1ten _ PDB _ prepared” “as the prepared protein structure and click OK
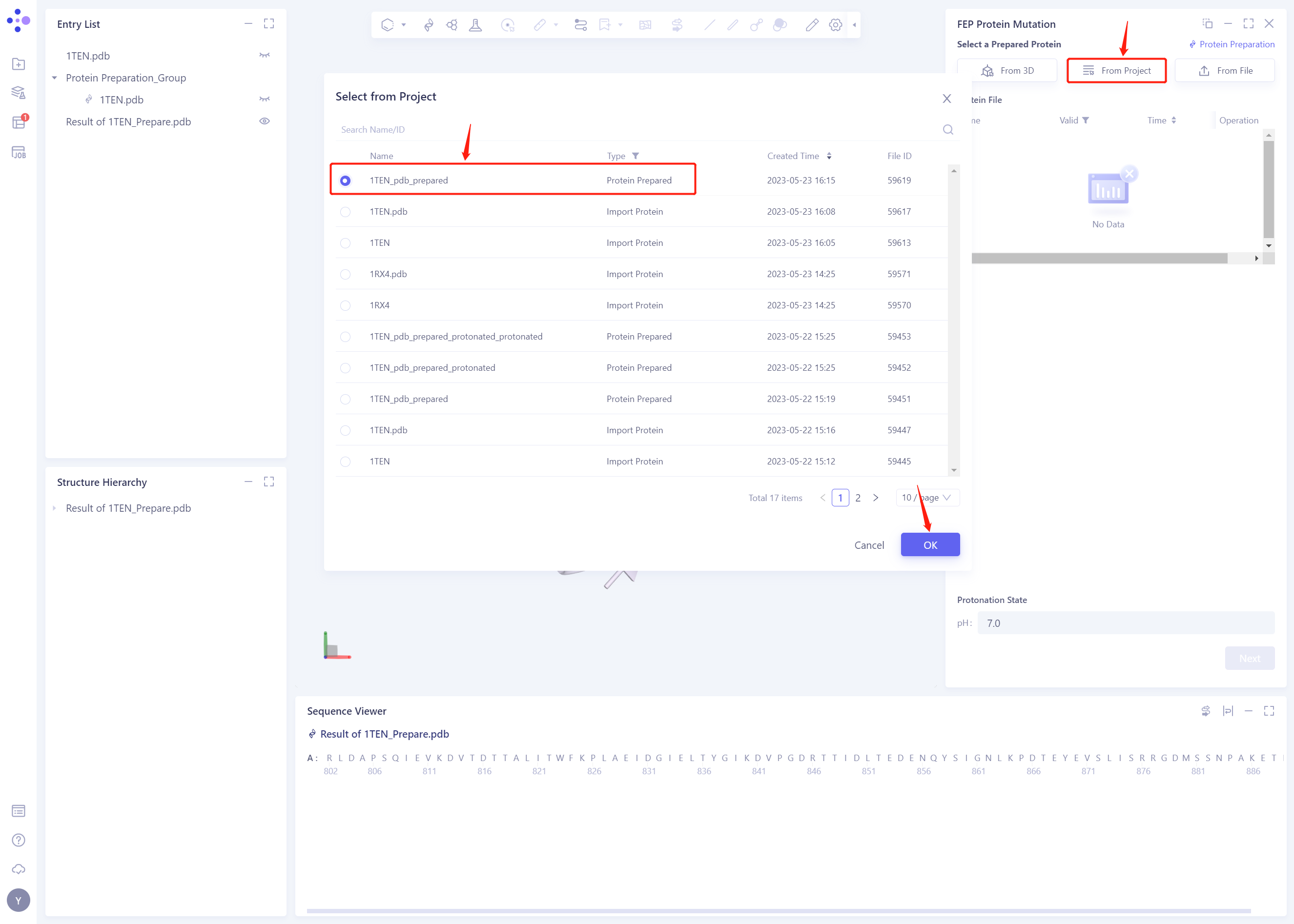
- After clicking OK, the system will automatically check whether the input protein meets the calculation requirements, and the status is “Processing”.
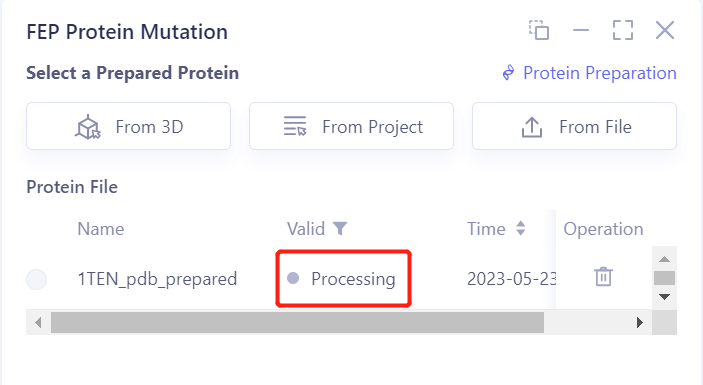
-
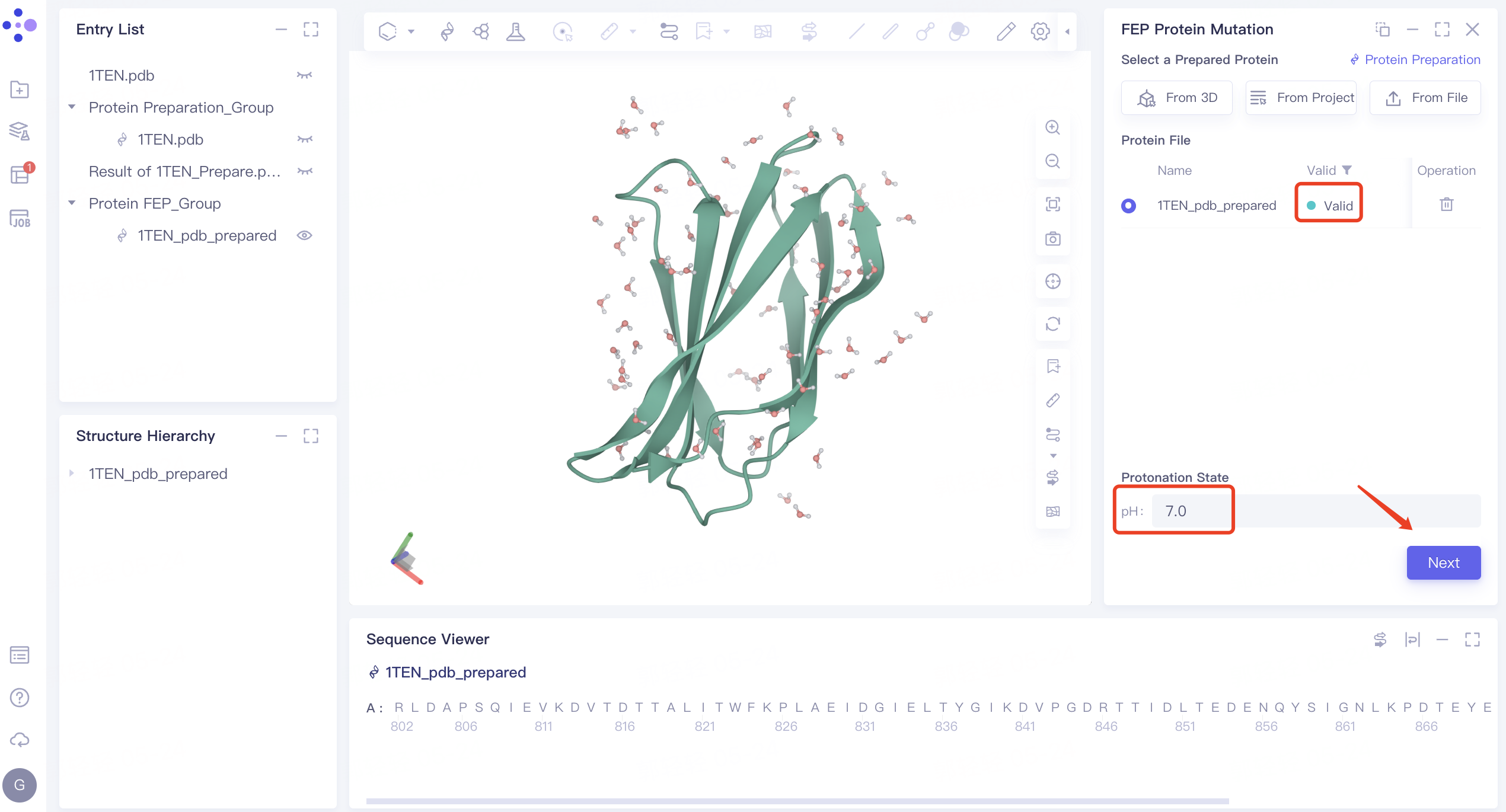
In less than 1 minute, the system will judge that the protein is “Valid”.
-
Set the protonation status of protein amino acids. The default value of pH is 7.0. Do not modify it here. Click Next.
3.3 Select Mutation Site
-
Chain ID: select “Chain A”
-
Residue Number: Enter the residue number according to the following amino acid mutation list (residues can be selected in Sequencce Viwer)
-
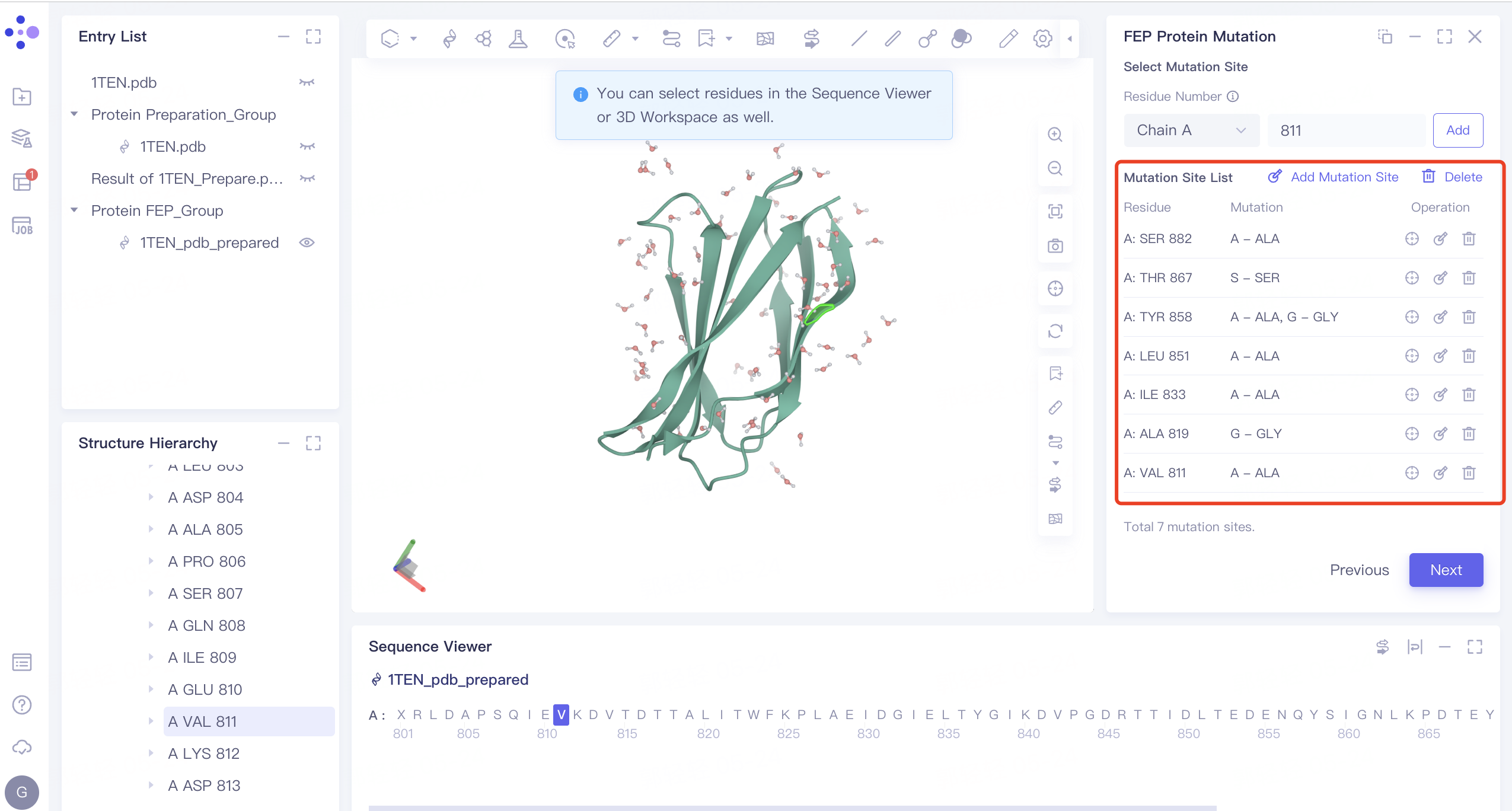
Mutation Site List: Select the type of residue to mutate to based on the following list of amino acid mutations
-
After the mutation setting is completed, as shown in the following figure, click Next to proceed to the next step
- Amino acid mutation information list:
| Residue Number | Wild Type | Mutation | ddG_Exp |
| 811 | V | A | 1.36 |
| 819 | A | G | 2.8 |
| 833 | I | A | 3.28 |
| 851 | L | A | 2.99 |
| 858 | Y | A | 1.38 |
| 858 | Y | G | 2.76 |
| 867 | T | S | 1.22 |
| 882 | S | A | 2.28 |
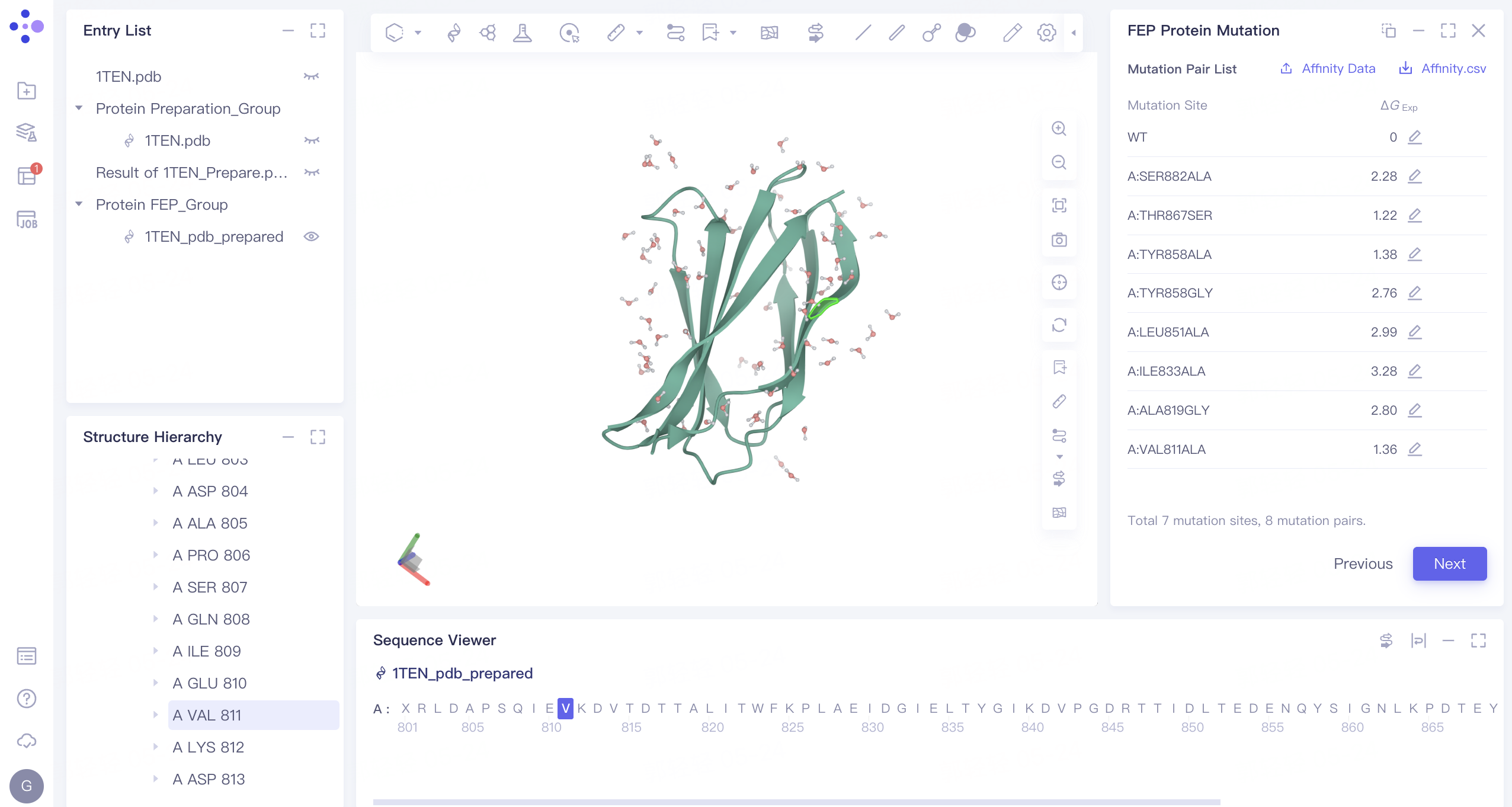
3.4 Mutation Pair List
-
According to the amino acid mutation information list, manually input the free energy change experimental value of amino acid mutation. (The experimental free energy of the WT system was set to 0, and the experimental free energy value of each mutant was obtained according to the experimental free energy difference.)
-
Click Next to proceed to the next step
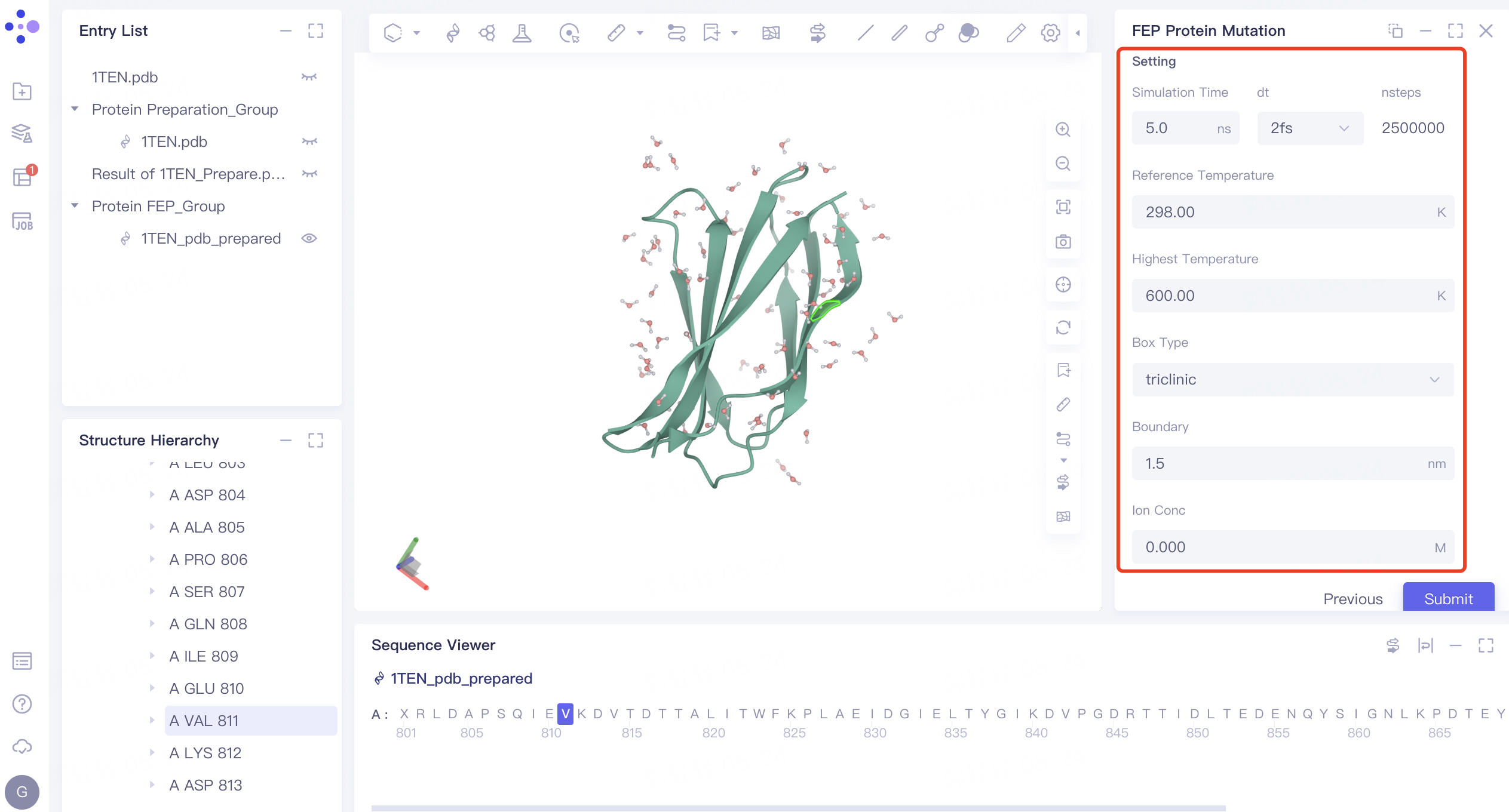
3.5 Setting
- The parameter settings remain at the default values. Click Submit to submit the task
4. Analysis of results
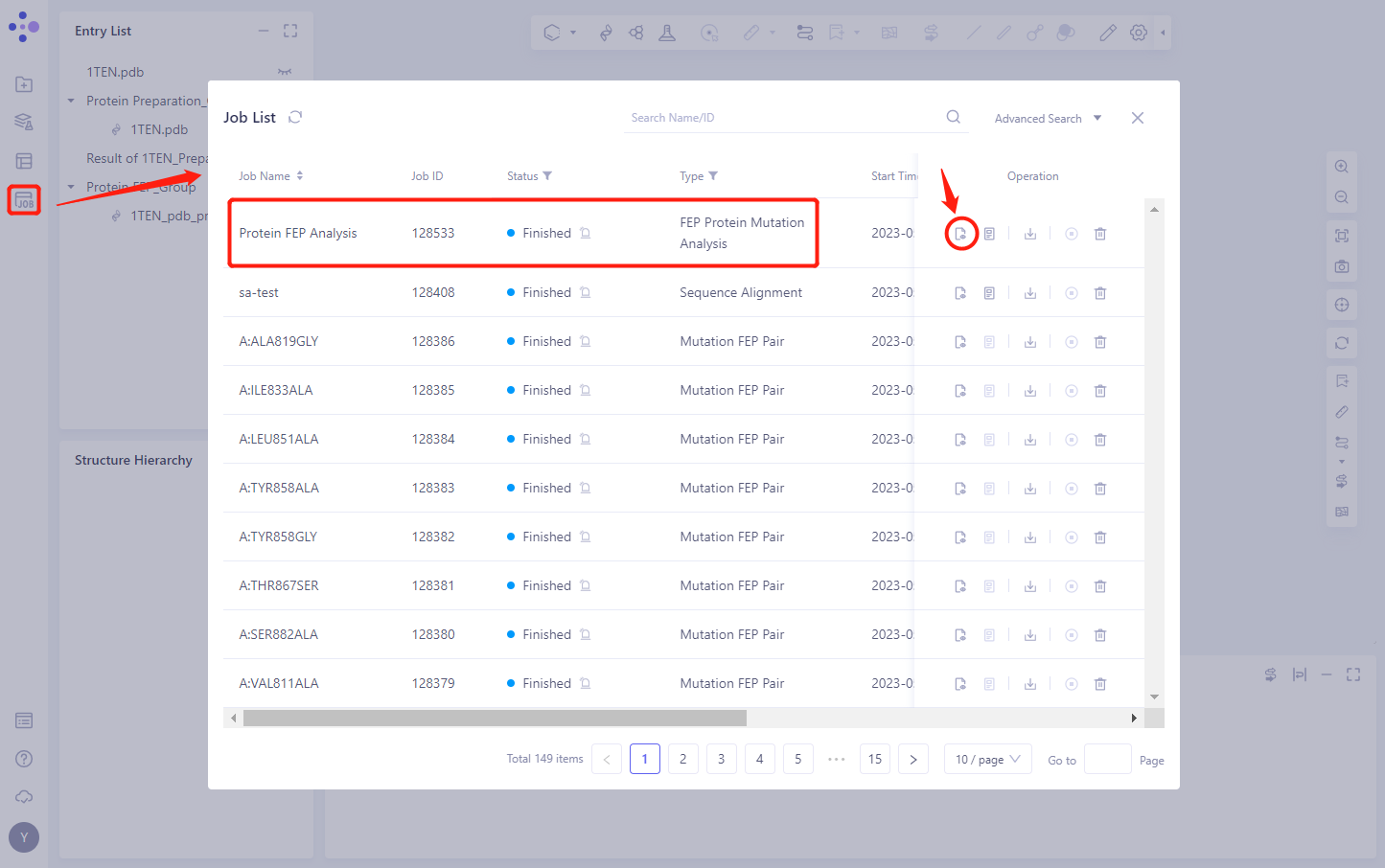
4.1 Entrance
- In the left general menu bar Job → Job List, find the “Protein FEP Analysis” calculation task “and click the” Show “button
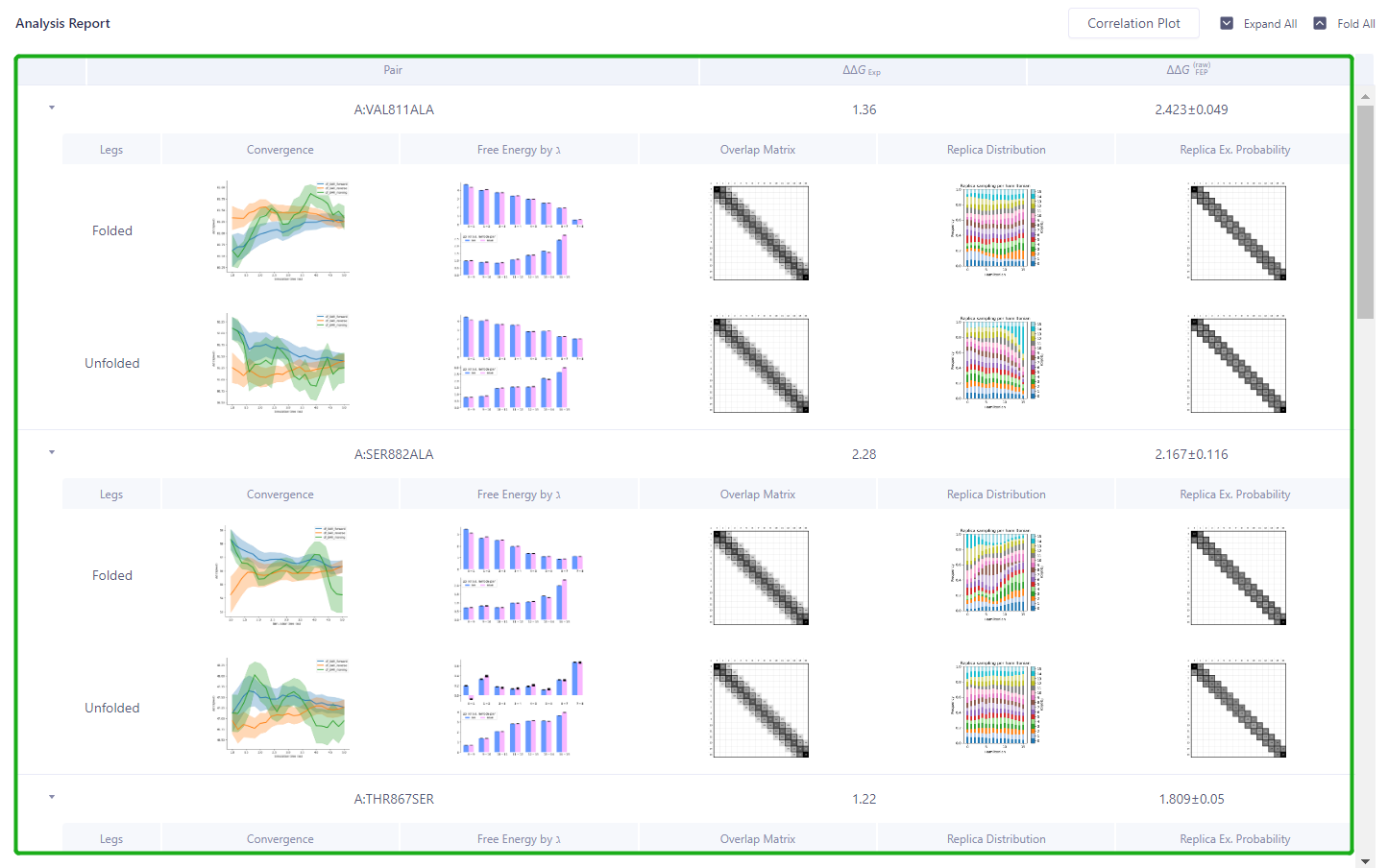
4.2 Results presentation
-
The FEP Protein Analysis result interface will pop up automatically
-
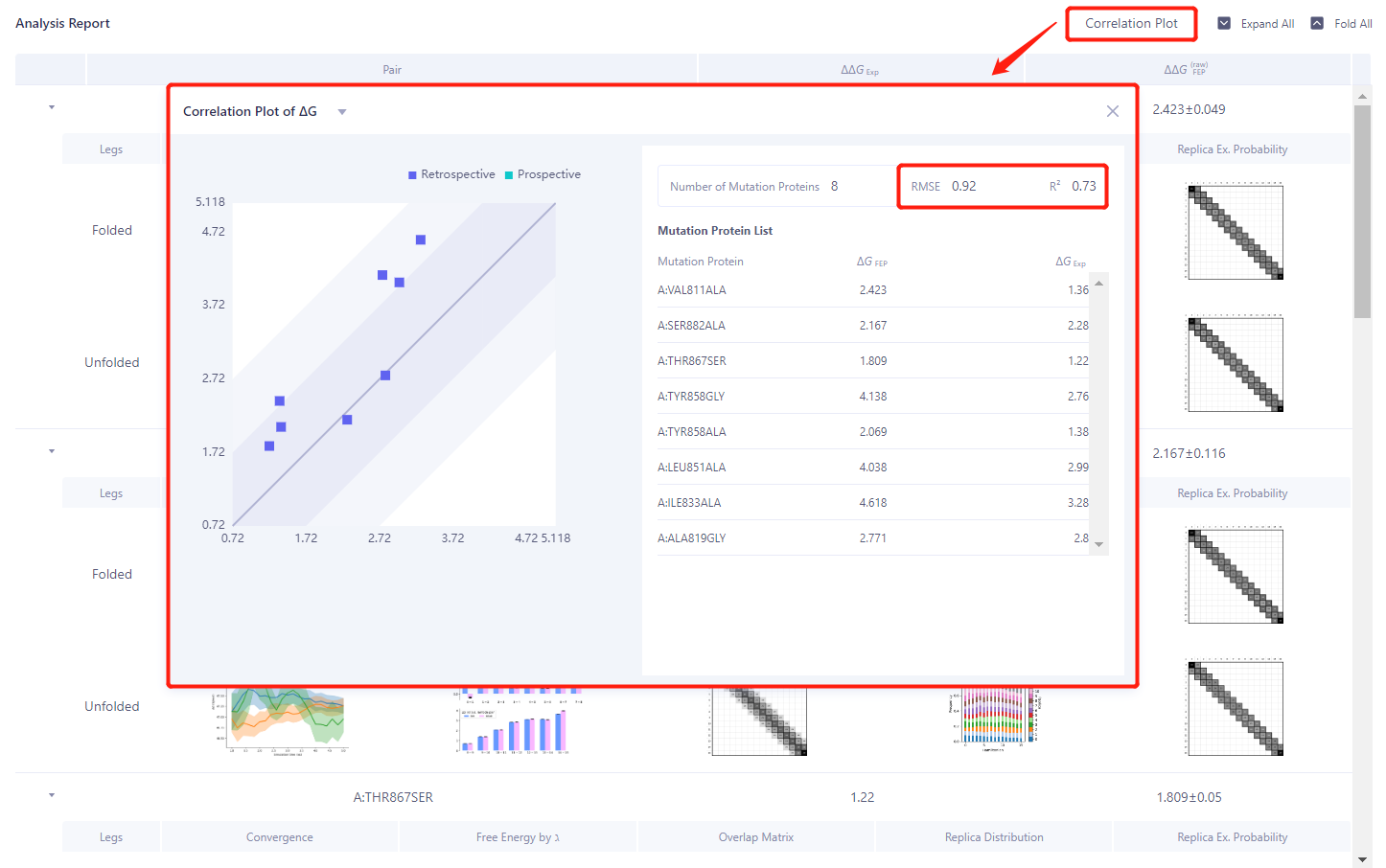
Green box: The reliability of each Pair calculation process can be analyzed.
- Red box: Click “Correlation Plot” and select “Correlation Plot of ΔG” to analyze the correlation between the experimental value ΔG and the calculated value ΔG. The RMSE and R ² are 0.92 kcal/mol and 0.73 respectively, showing a good correlation.
5. References
[1] Structure of a Fibronectin Type III Domain from Tenascin Phased by MAD Analysis of the Selenomethionyl Protein. SCIENCE. 6 Nov 1992. Vol 258, Issue 5084. pp. 987-991.
[2] The amino acid mutational thermostability experimental data used in this tutorial were derived from the database: Stourac, J., Dubrava, J., Musil, M., Horackova, J., Damborsky, J.; Mazurenko, S., Bednar, D., 2020: FireProt(DB): Database of Manually Curated Protein Stability Data. Nucleic Acids Research 49: D319-D324.