Loop Optimization
Introduction
Proteins are one of the most important molecules in life, and they play various roles in life, including catalyzing chemical reactions, transmitting signals, and supporting cell structures. The function of a protein is determined by its structure, and the loop region with strong flexibility often undertakes important processes such as protein recognition and functional regulation, and the structure of the loop region is difficult to accurately determine by structural biology through experimental means. It is very important to optimize the conformation of the protein loop to obtain a more reasonable loop structure. Hermite The Loop Optimization module of the platform provides the function of optimizing the conformation of the flexible loop structure of proteins. With the Hermite platform's friendly user interface, optimized algorithm and flexible Cloud Service resources, you can easily and quickly obtain a more reasonable protein loop conformation.
Transient Receptor Potential Channel 1 (TRPV1) is an ion channel protein primarily distributed in nerve endings and the Central Nervous System. It is involved in sensing and transmitting signals from heat pain, capsaicin (a spicy substance found in chili peppers) and other chemical stimuli. When the TRPV1 channel is activated, calcium ions flow into nerve cells, causing changes in signaling to sense heat pain and spiciness. TRPV1 plays an important role in pain and inflammatory responses and has therefore become a target for many research pain treatments. Its loop region is thought to play an important role in the activation and regulation of the channel.
In this tutorial, you will learn to use the Loop Optimization module to model and optimize the TRPV1 pore turret loop segment whose structure cannot be accurately determined, guiding the subsequent study of the mechanism of protein conformational delivery and the rational design of targeted drugs.
1. Create project and import structure
1.1 Login system
Login address https://hermite.dp.tech

1.2 Create a project
After entering the system, create a new project "7LQY Loop Optimization"
1.3 Imported protein structure
The left general menu bar Menu → File → Import Structure
Click Select Files, select the 7LQY.pdb file, and click Import to import the protein structure.
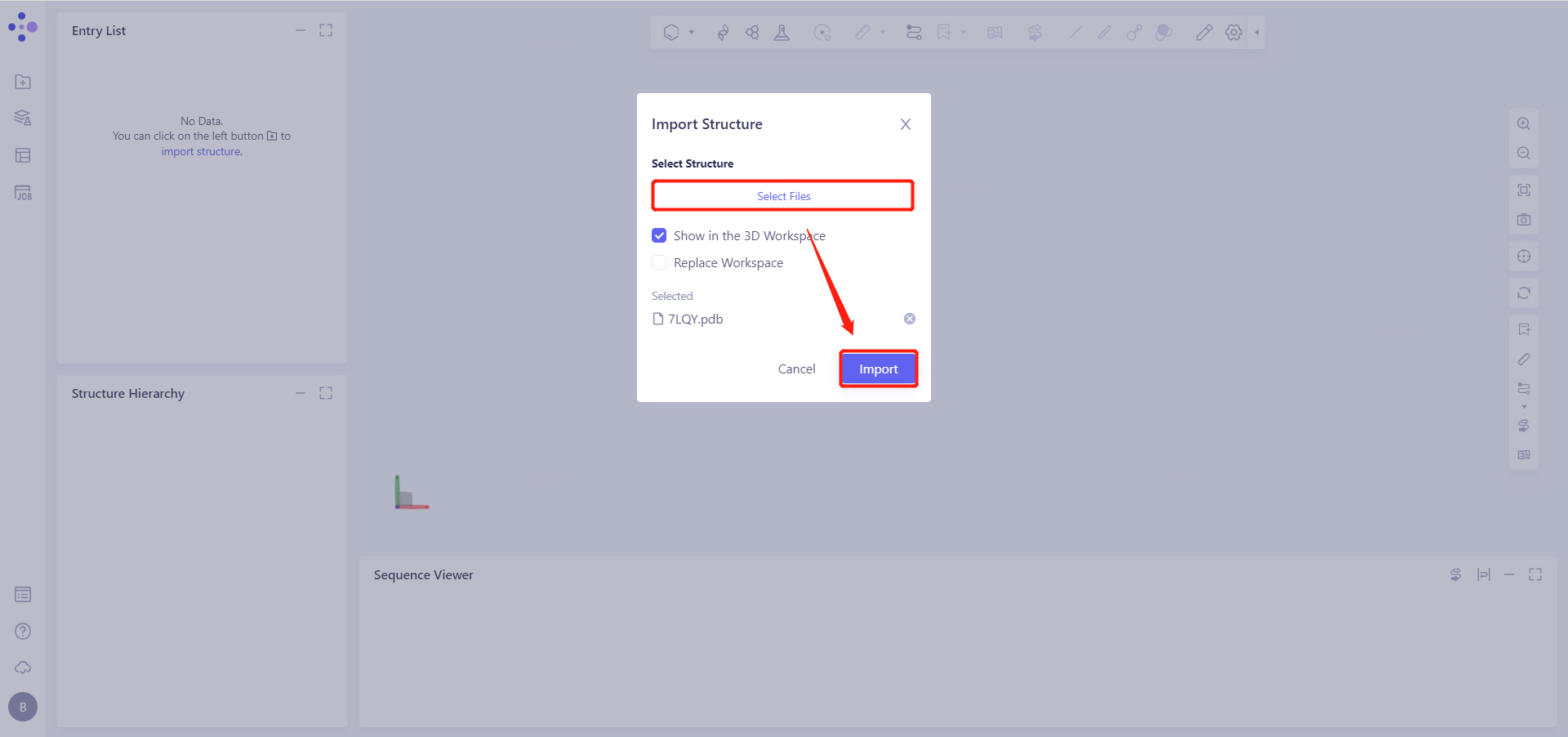
2. System preparation
2.1 Preparing the protein structure
2.2.1 Select Structure
Left General Menu Bar Function → Structure Modeling → Protein Preparation
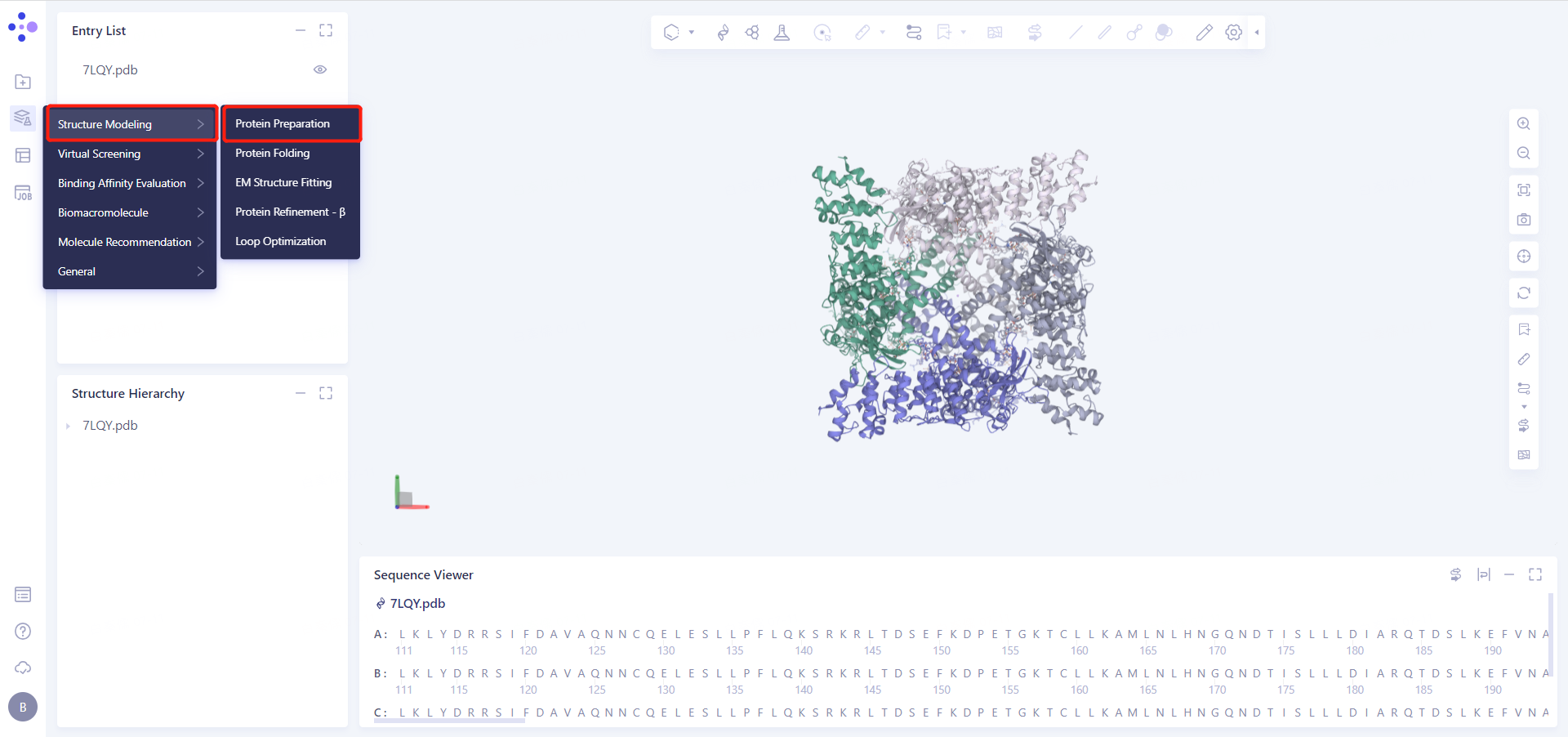
Select Structure from 3D Workspace: Select Protein in Structure Hierarchy and click OK.
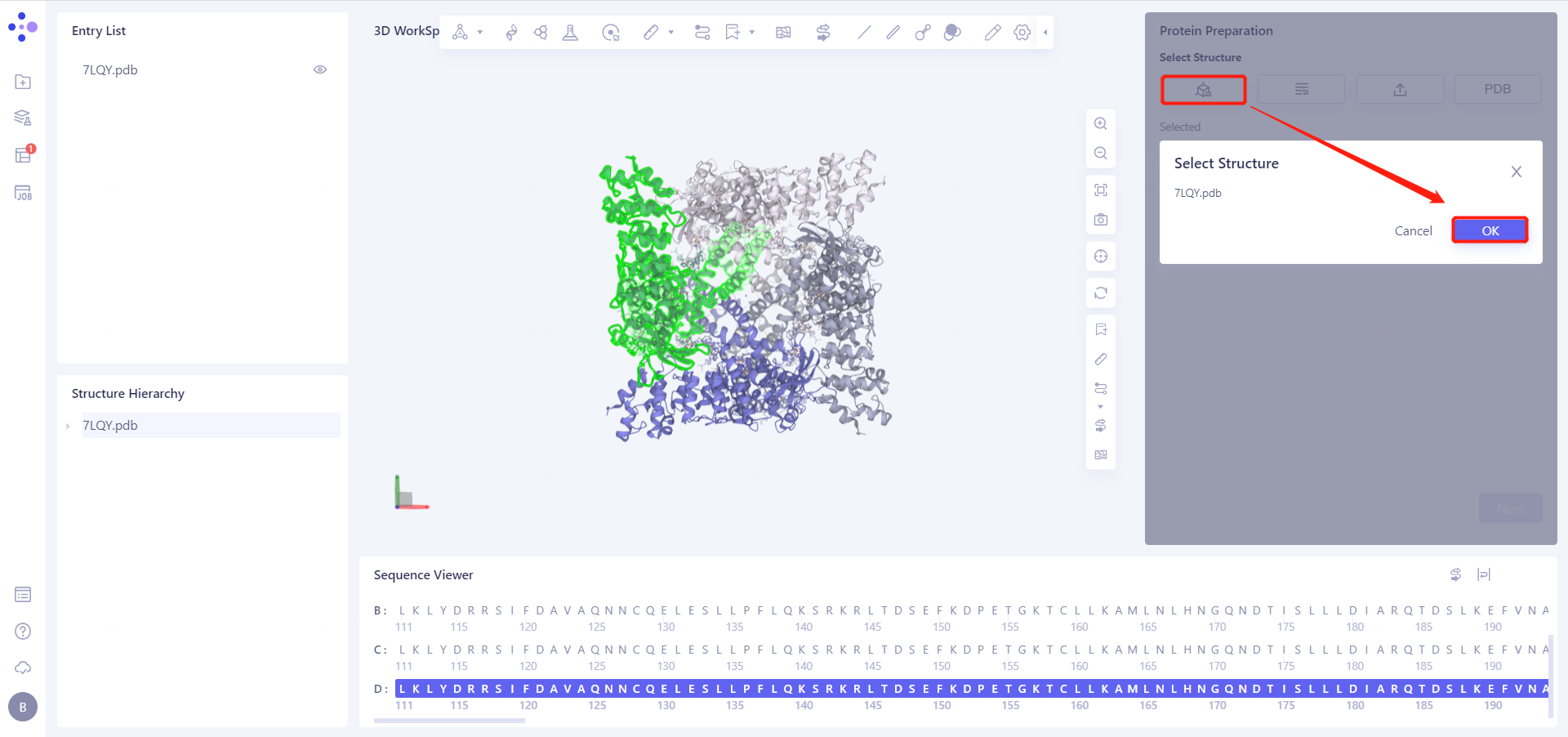
7LQY.pdb (ChainD) is loaded into the Protein Preparation parameter setting panel, click Next
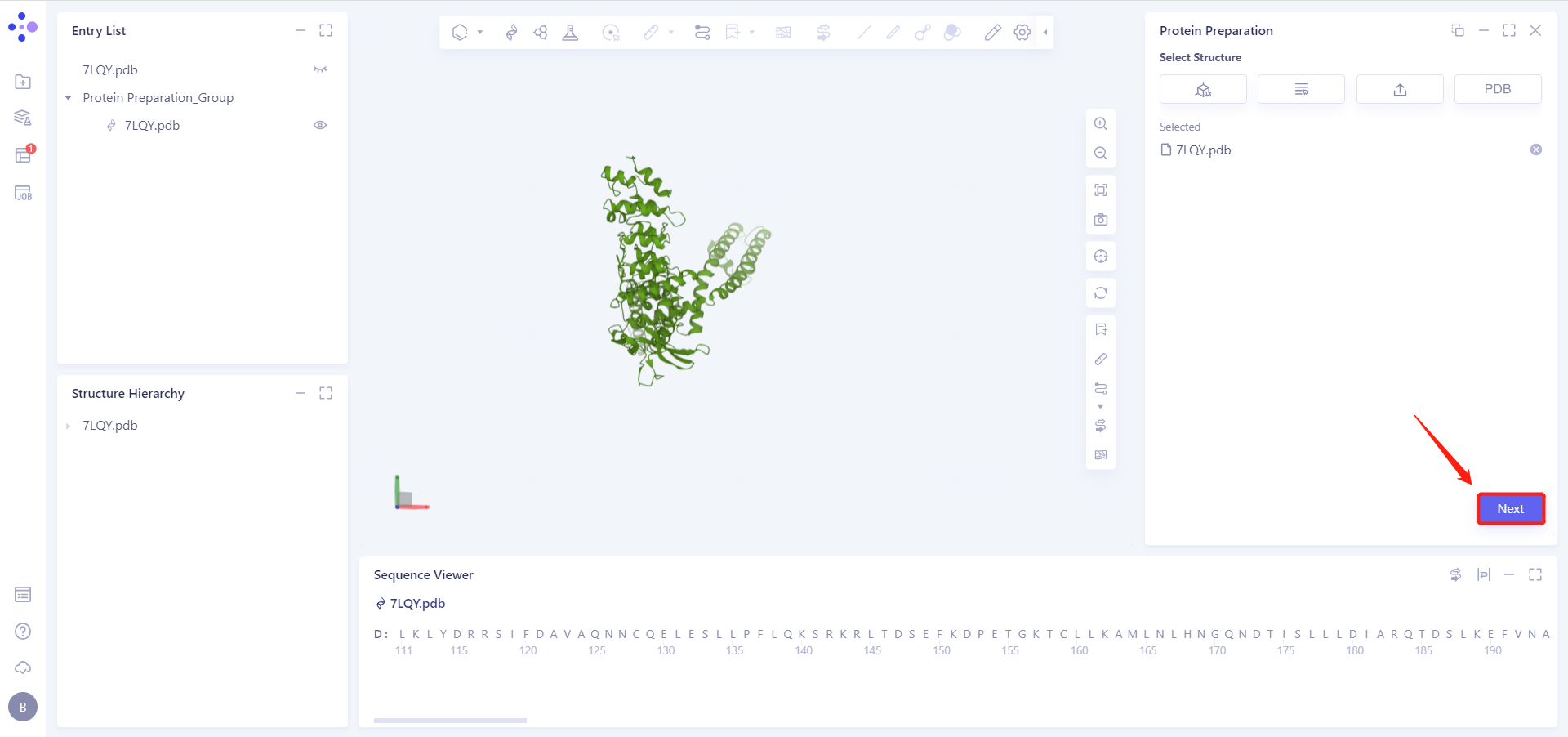
2.2.2 Select Polymer, Other Groups to Keep: Select D-Chain Protein and click Next
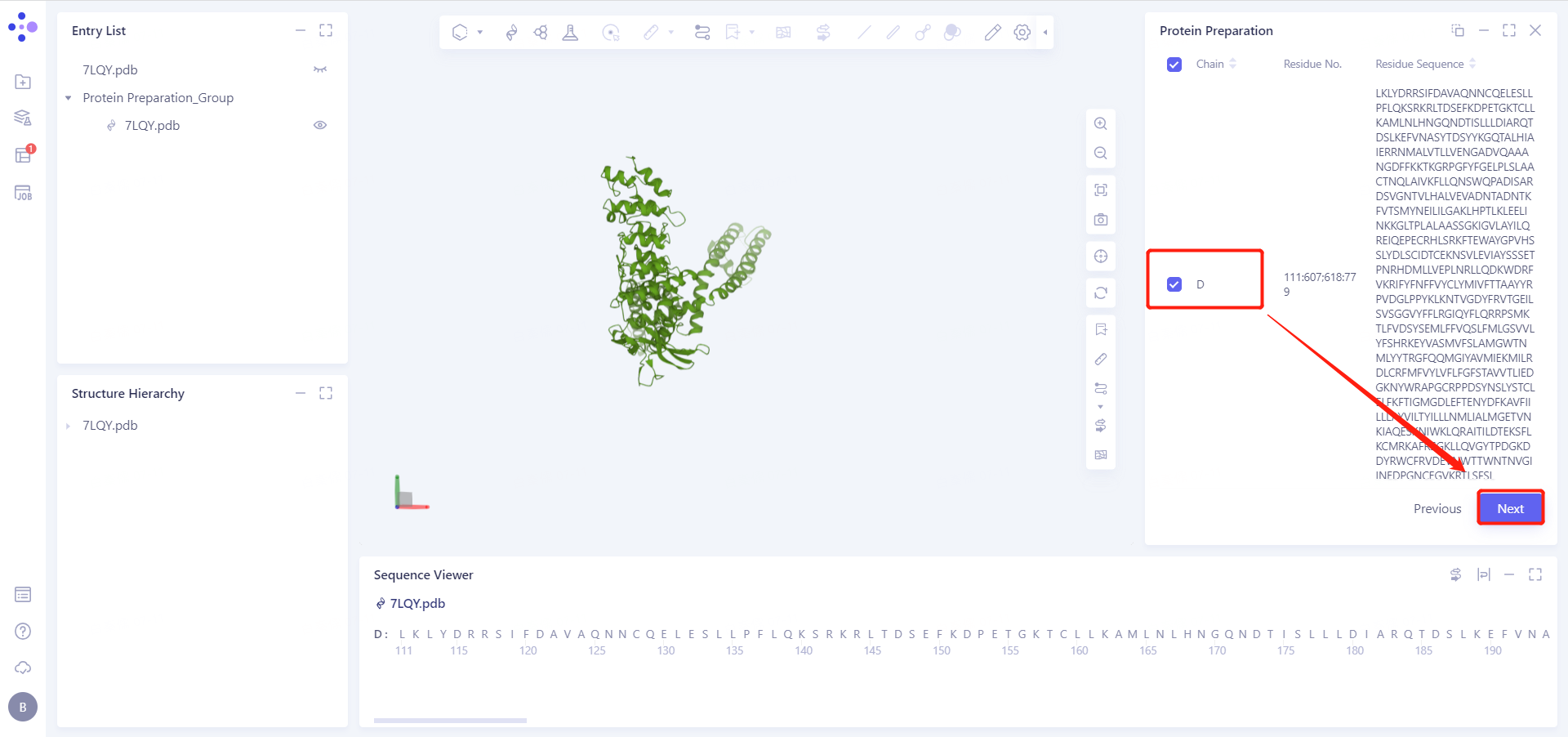
2.2.3 Select Missing Residues to Repair
In this case, we need to complete the missing residues 608-617 that cryo-EM failed to resolve accurately.
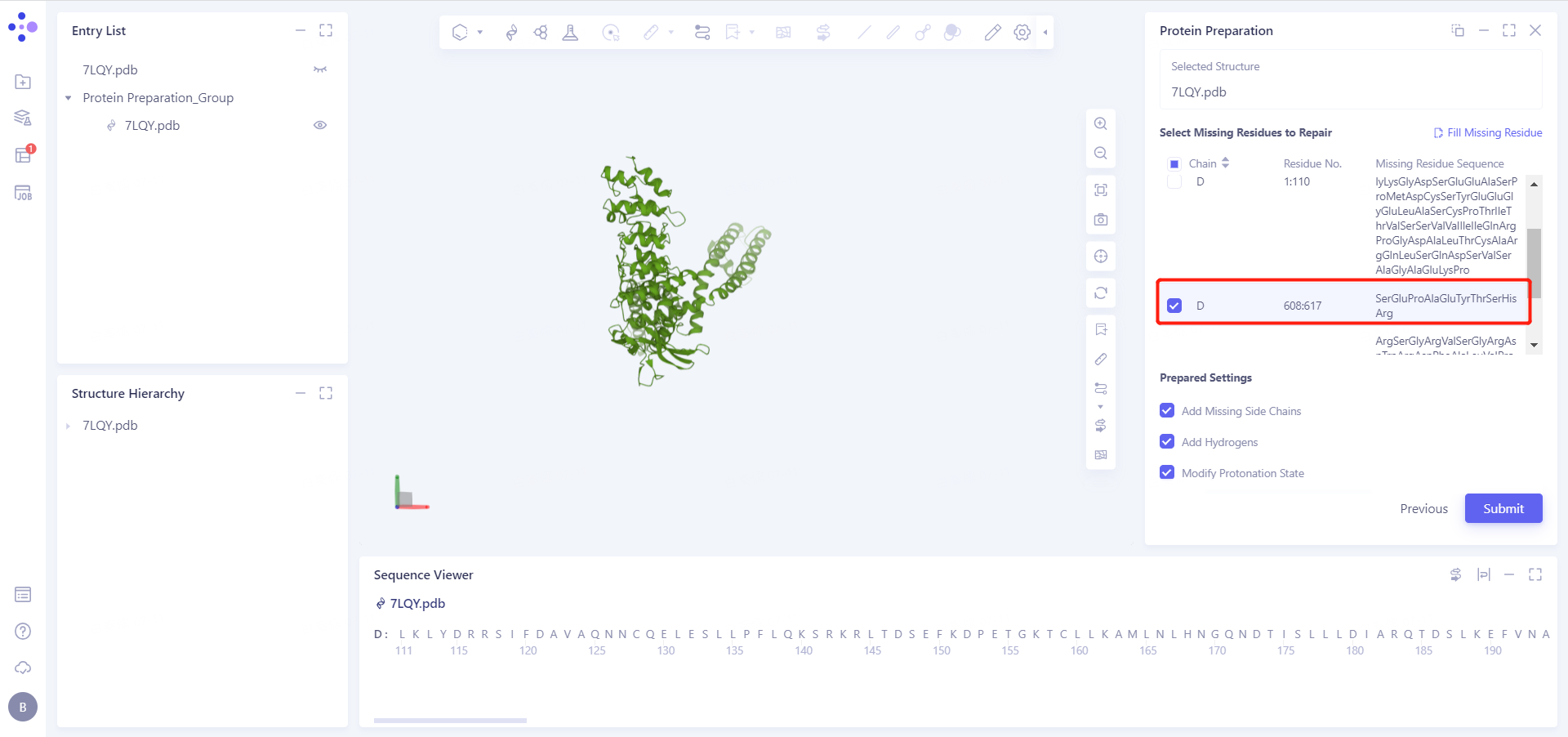
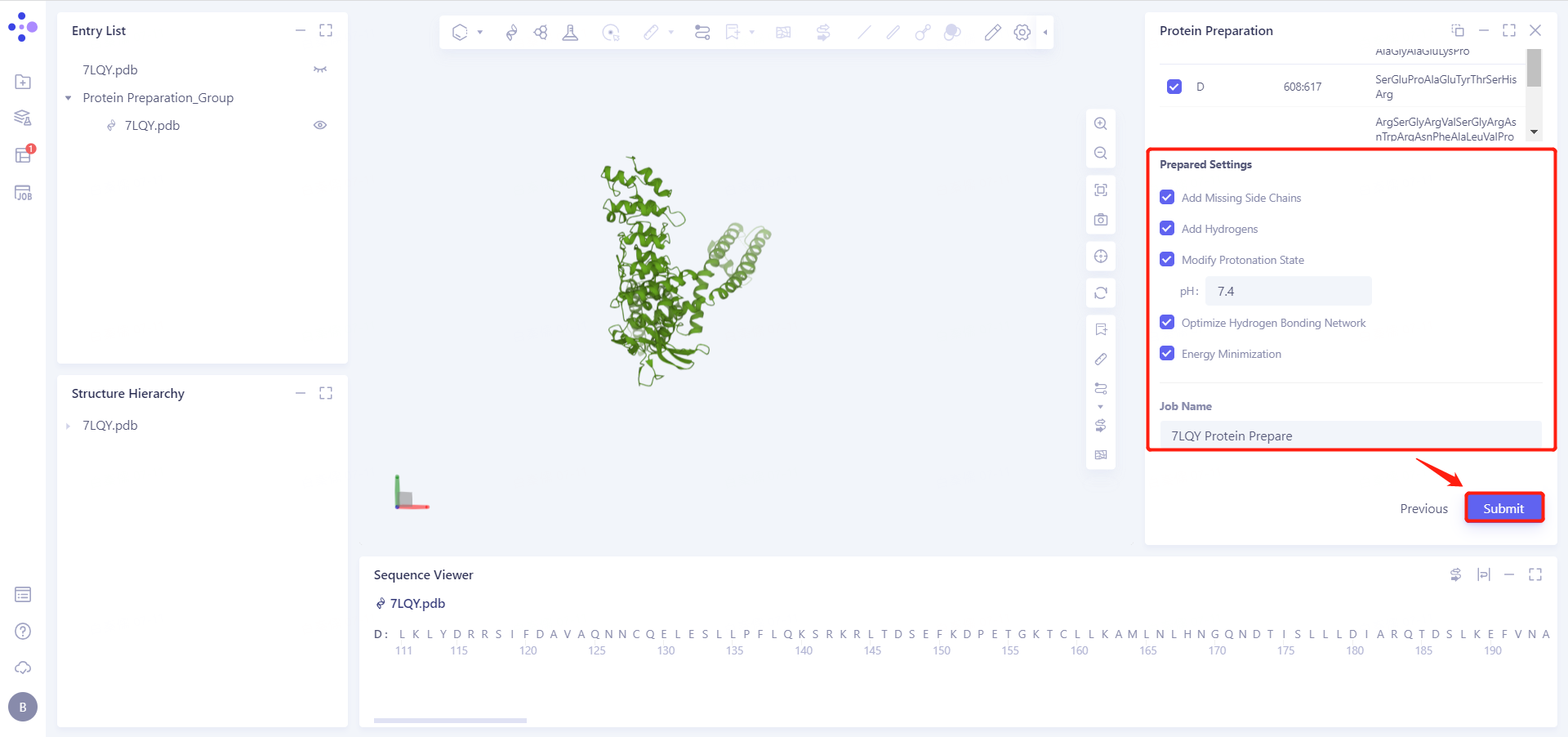
2.2.4 Prepared Settings
See the figure in the 2.2.5 for setting parameters
2.2.5 Name Job and Submit Task
Job Name is "7LQY Protein Prepare", click "Submmit" to submit the task
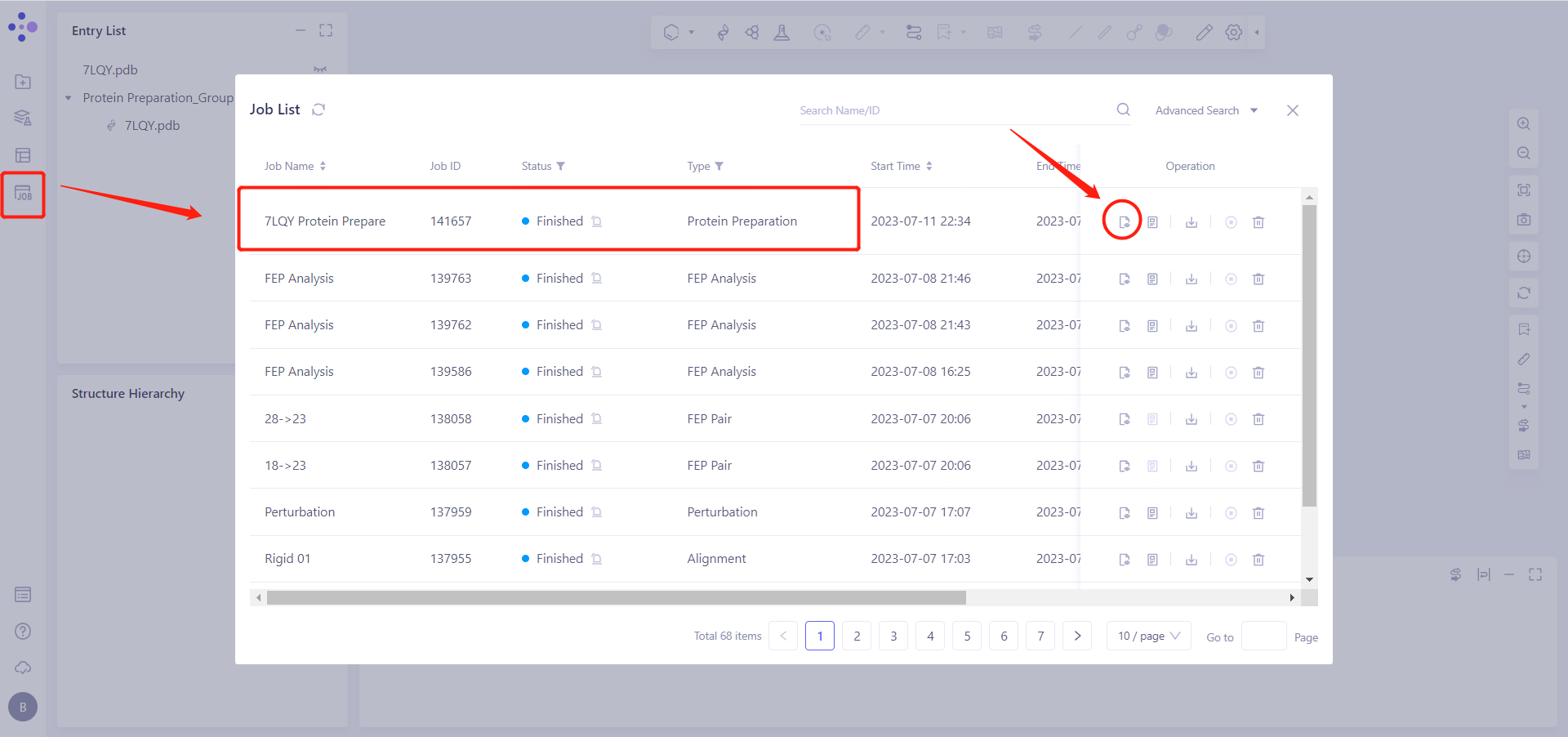
2.2.6 View Protein Preparation Results
Protein Preparation calculation tasks are generally completed within ten seconds to a few minutes. After the task is completed, click Jobs to view the corresponding task.
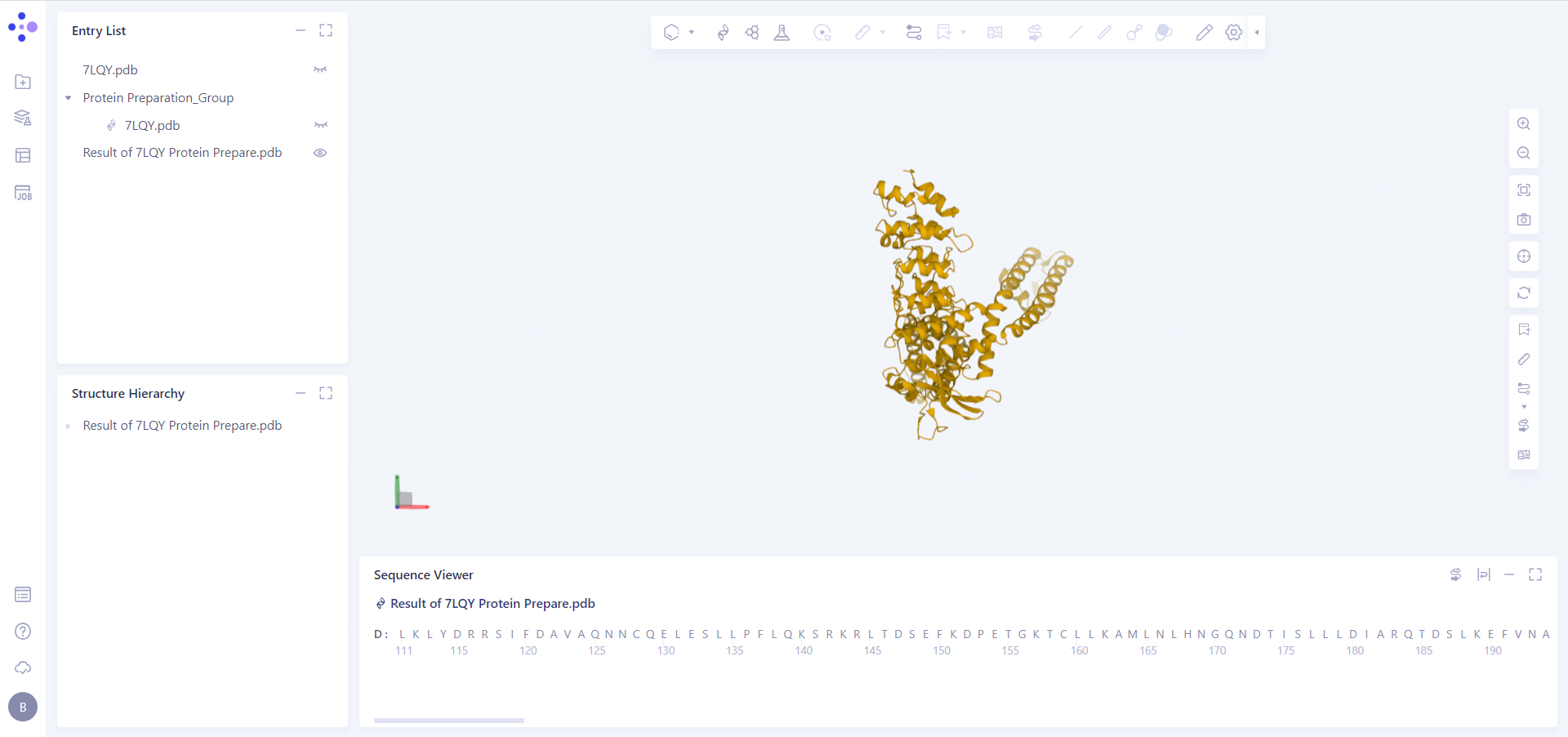
Click the "show" button to display the prepared protein structure in the 3D Workspace.
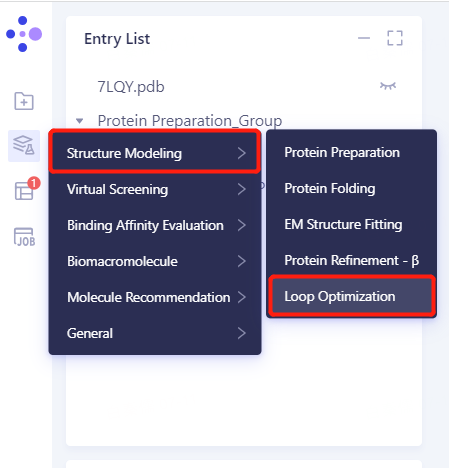
3. Create a Loop Optimization Task
3.1 Left General Menu Menu → Structure Modeling → Loop Optimization
3.2 Select Prepared Protein From Project
Select "7LQY_pdb_prepared" as the prepared protein structure and click OK.
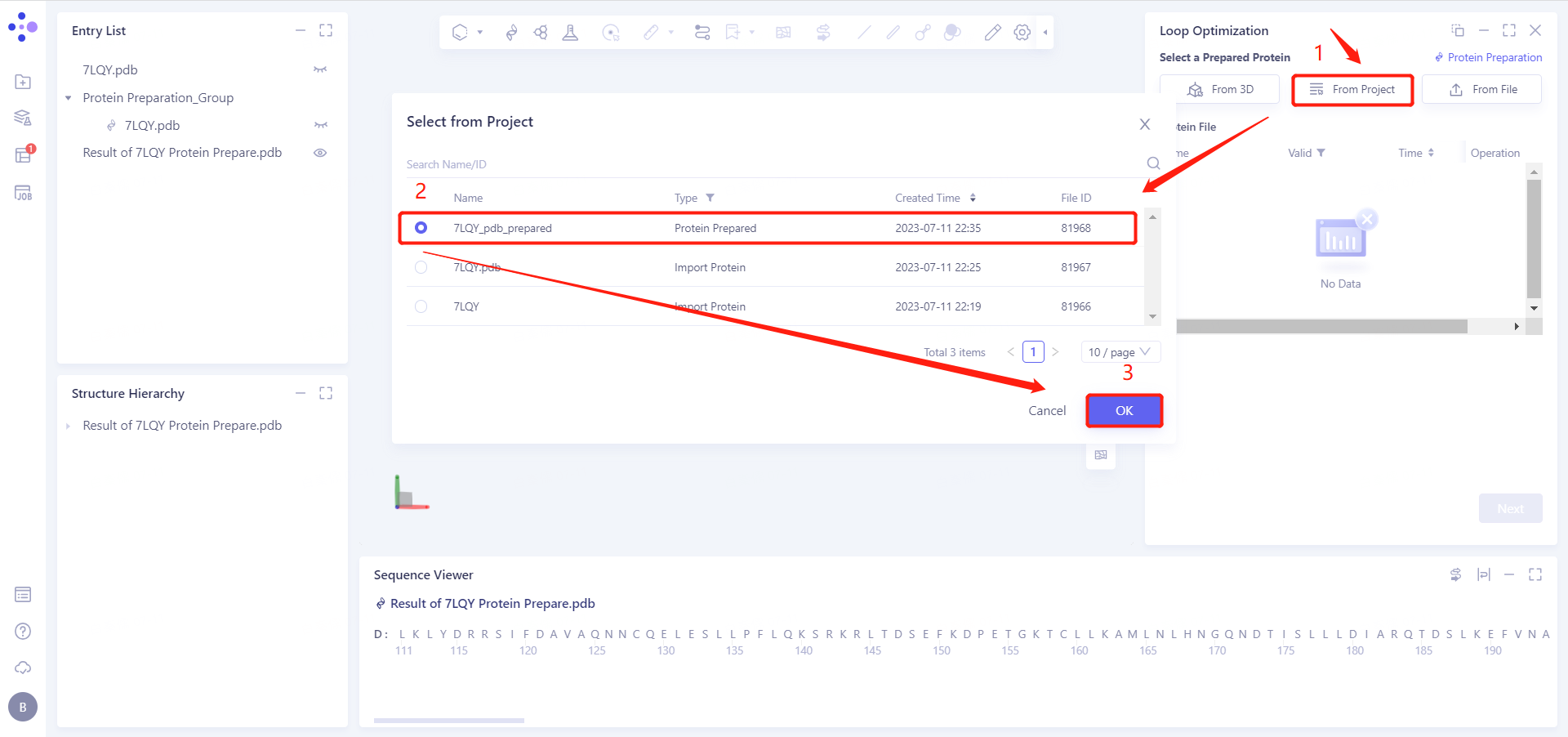
After clicking OK, the system will automatically check whether the input protein meets the calculation requirements, and the status is "Processing".
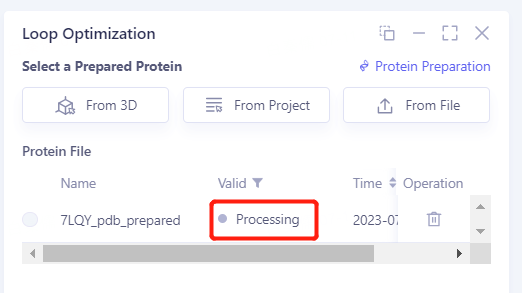
In less than 1 minute, the system will judge the prepared protein as "Valid" and click "Next".
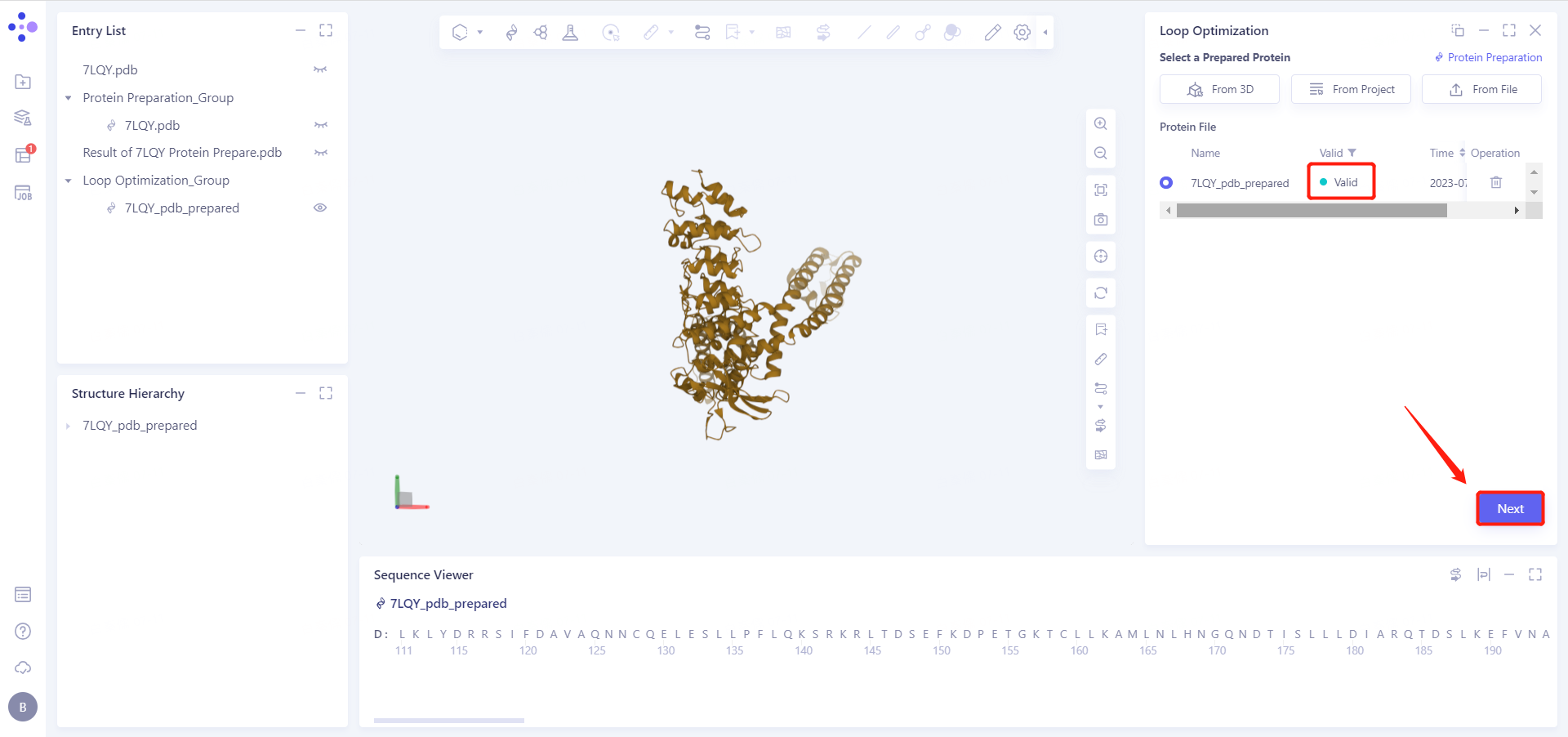
3.3 Select Optimization Site
Select the loop segment residues 602-631 to be optimized in the 3D Workspace;
Click "Add" to join "Select Residue "
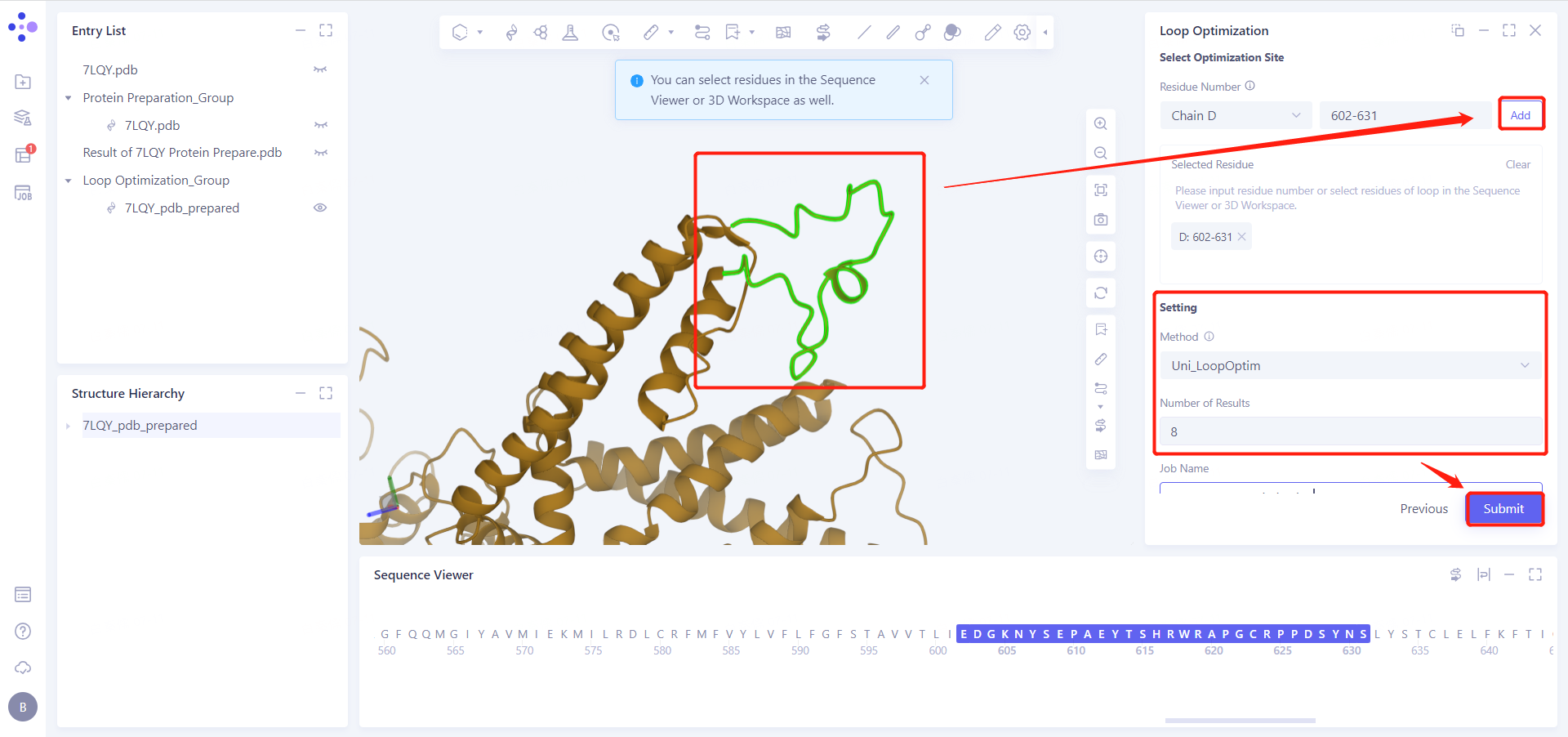
3.4 Setting and Submit
See Figure 3.3, "Method" select "Uni_LoopOptim";
Set "Number of Results" to 8;
Set "Job Name" to "7LQY_Loop_Optimization";
Click "Submit" to submit the task
4. Results analysis
4.1 Entrance
In the left general menu bar Job → Job List, find the "7LQY_Loop_Optimization" calculation task and click "Show".
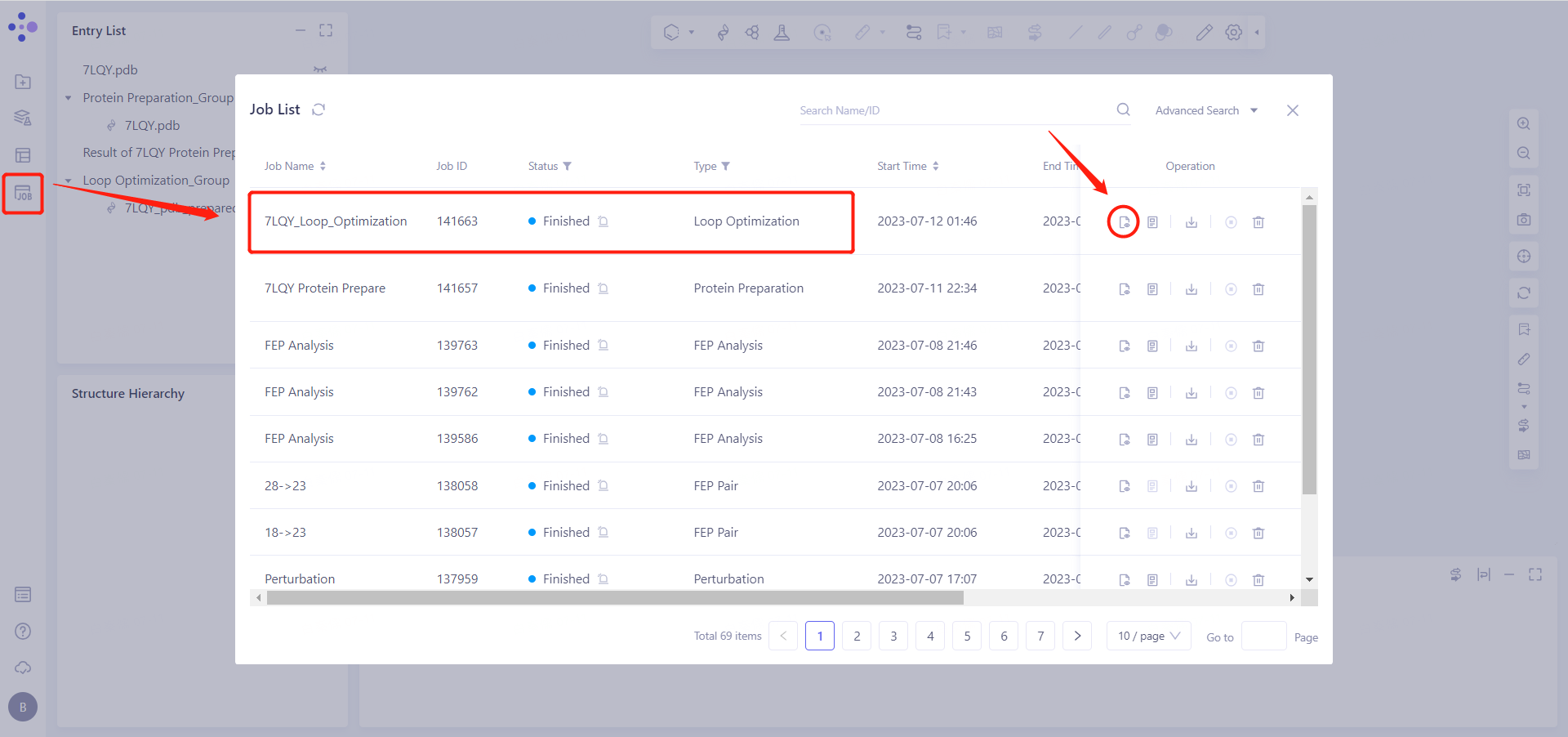
4.2 Display of results
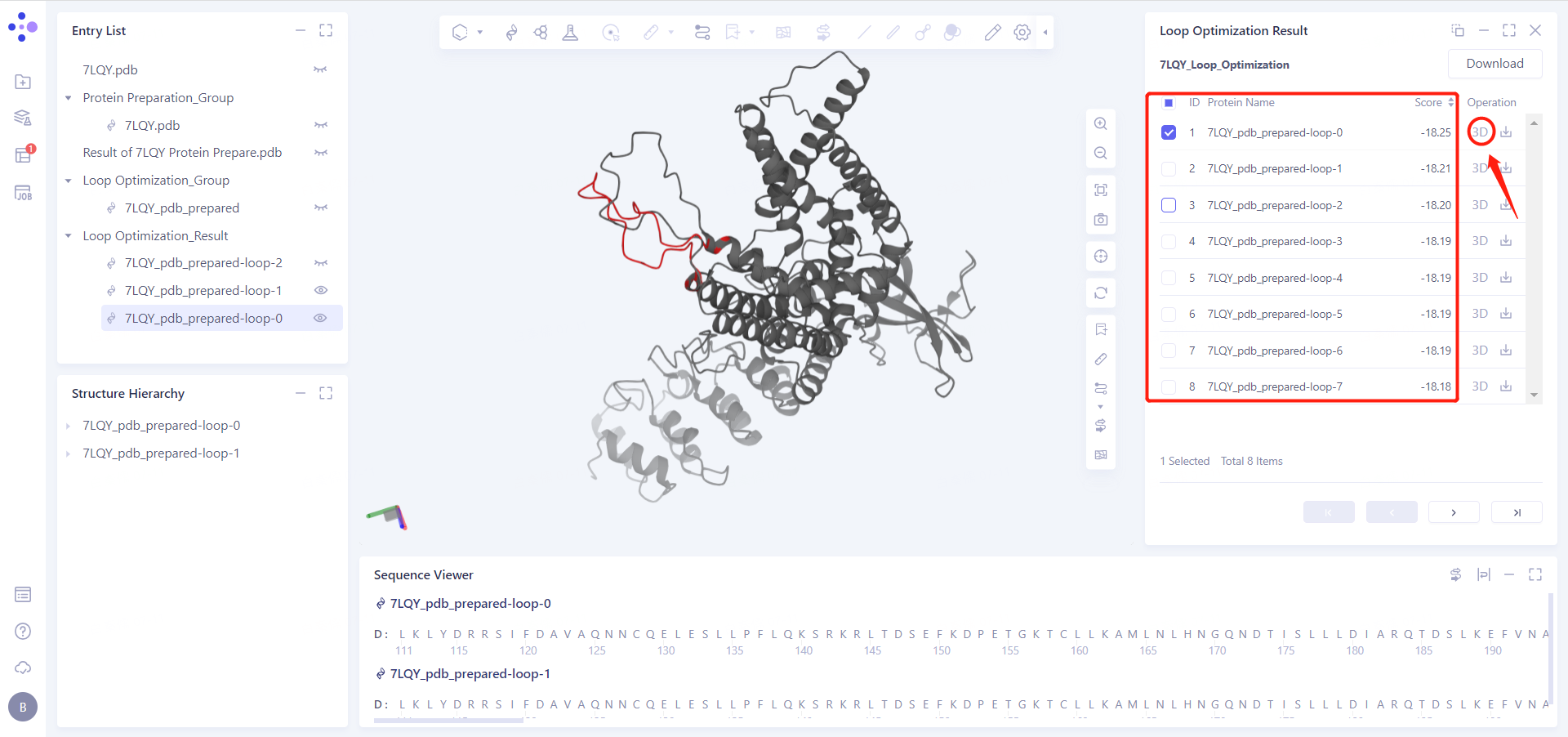
After selecting the generated model, click the "3D" button in the "Operation" column on the right side of the window to make the selected model appear in "3D Workspace";
The ranking of "Score" represents the judgment of the algorithm on the rationality of the model, and the lower the score value, the more reasonable the model is
5. References
[1] TRPV1 pore turret dictates distinct DkTx and capsaicin gating. Proc Natl Acad Sci U S A. (2018) 115 (50): E11837-E11846.