Induced Fit Docking
Introduction
The classical lock-key model clarifies the relationship between drugs and target proteins, that is, the spatial matching between drugs and target proteins in vivo will produce a recognition relationship similar to that between keys and locks. However, under actual physiological conditions, the target receptor and drug ligand are flexible, that is, in the binding process, the molecular conformation of the target protein and the substrate is changing, not only to meet the matching of space, but also to meet the matching of binding free energy. Therefore, the theory of “Induced Fit” was proposed D. E. Koshland in 1958. The core content is that the active site of protein changes through the interaction with ligands. Therefore, “Induced Fit” is more accurate than the prediction of “Lock and Key Model”.
Thrombin is a multifunctional serine protease, which catalyzes the conversion of fibrinogen to fibrin, leading to blood clotting and hemostasis.Thrombin is involved in the last step of the activation of the coagulation cascade, and excessive stimulation of the coagulation cascade can cause thromboembolic symptoms in clinic.Therefore, thrombin is a key target for anticoagulation and thrombus prevention in modern drug research. Considerable efforts have been made to develop safe and effective inhibitors of thrombin. A class of non-peptide inhibitors has been designed to achieve activity in the low nanomolar range [1-2].
Uni-IFD can accurately predict the binding mode of drug and target by simulating the “induced fit” effect when drug molecules bind to the target. In this tutorial, you will learn to use the Uni-IFD module to accurately predict the induced fit patterns of two Thrombin inhibitor complex and guide the optimal design of drug molecule binding to the target. (Thrombin inhibitor complex, PDB ID: 1KTT and 1C5N. The ligand for 1C5N is ES1 and the ligand for 1KTT is C02)
Protein structure files used in this tutorial:
Ligand structure files used in this tutorial:
Pocket files used in this tutorial:
1. Create a project and import the structure
1.1 Log in to the system
- Login address: https://hermite.dp.tech

1.2 Create the project
- Once in the system, create a new project IFD test ”.
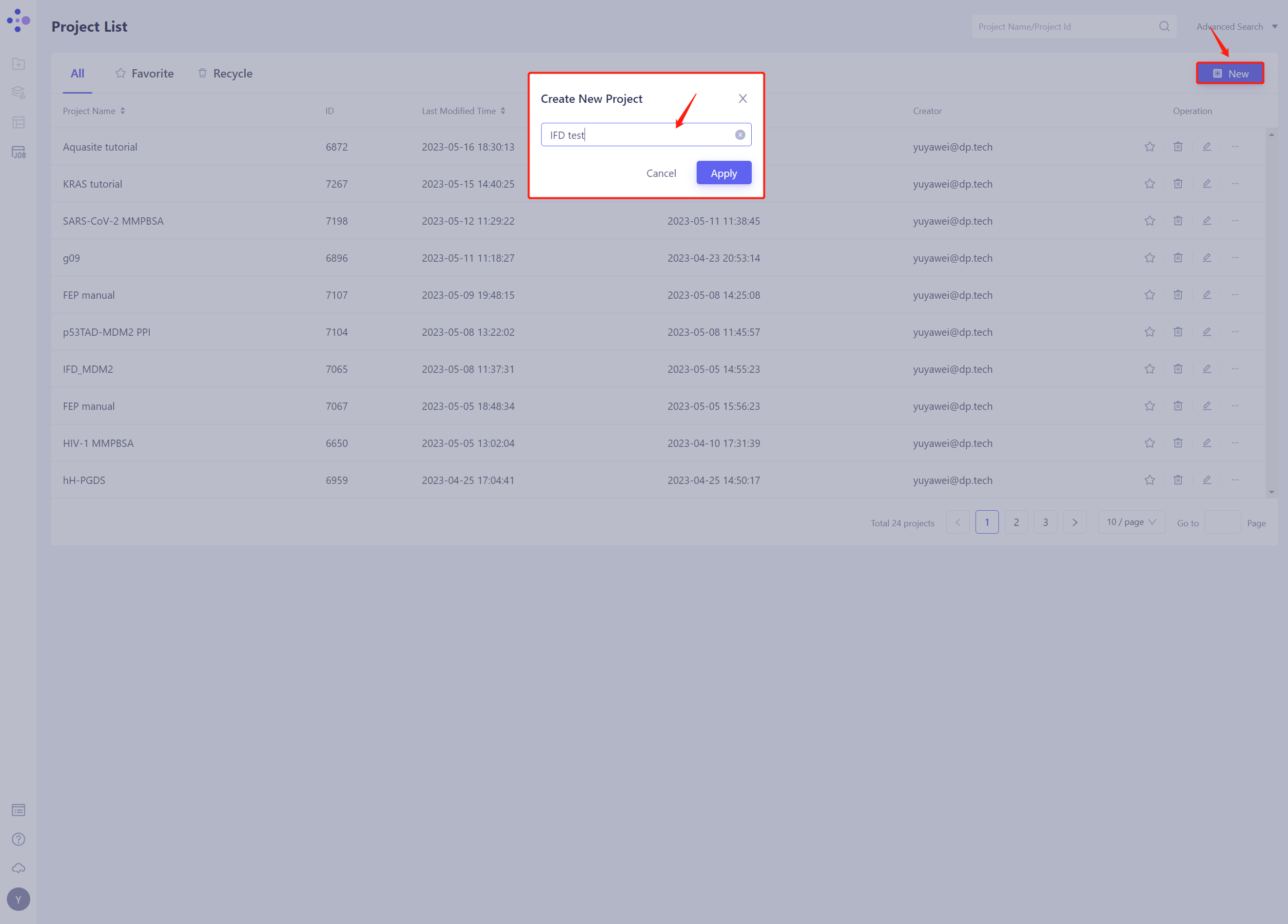
1.3 Import Protein Structure
- Left general menu bar Menu → File → Import Structure
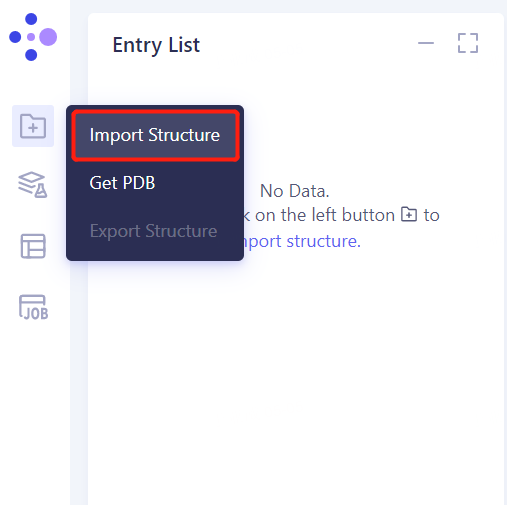
- Click Select Files, select 1C5N.pdb, 1C5N _ Ligand. SDF, 1KTT.pdb and 1KTT _ Ligand. SDF files, and click Import to import the protein structure
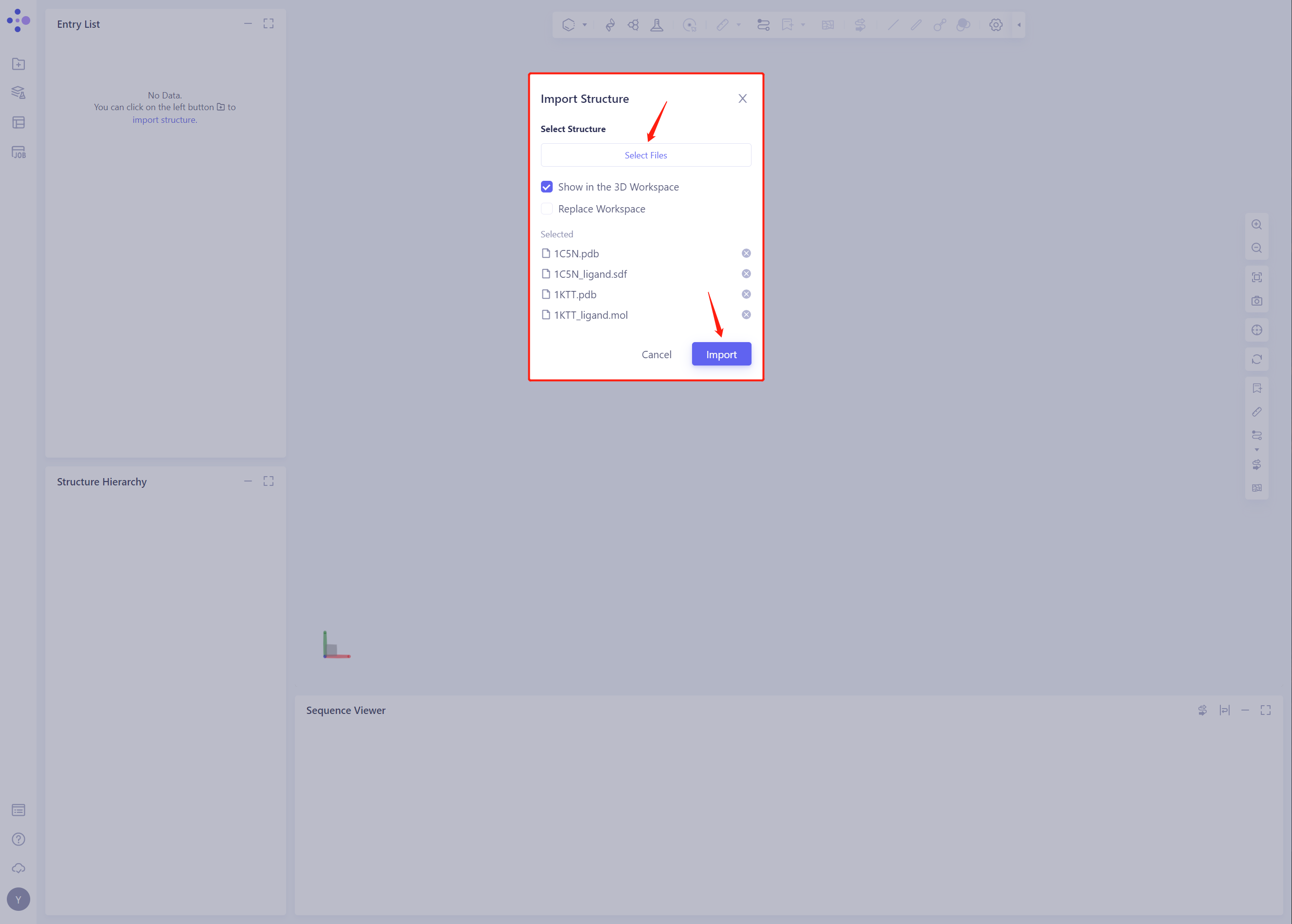
2. Create Induced Fit Docking Task
2.1 Click the general menu bar on the left: Menu → Function → Virtual Screening → Induced Fit Docking.
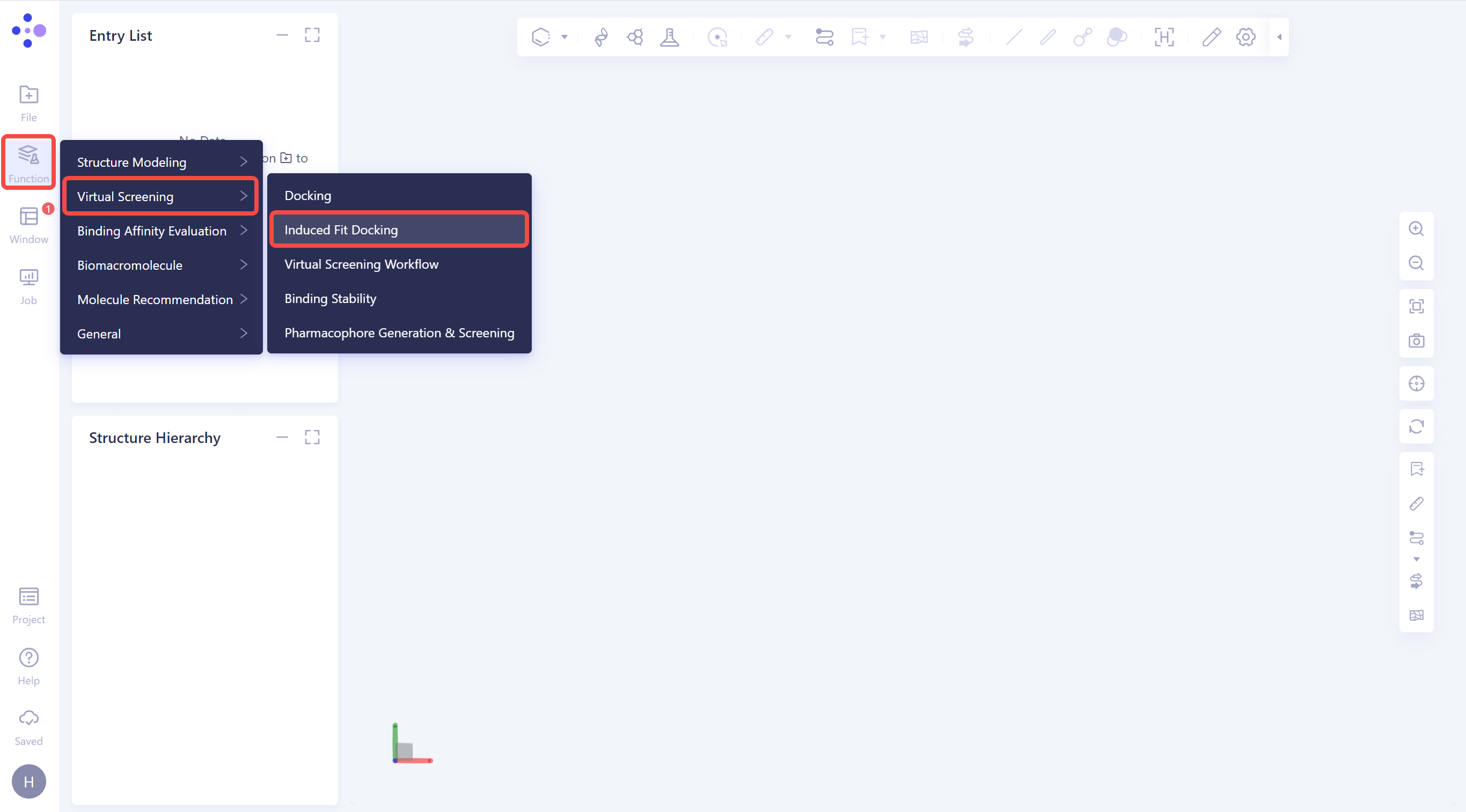
2.2 Select a Prepared Protein From 3D
- Select “1C5N.pdb” as the Receptor protein structure, and click OK
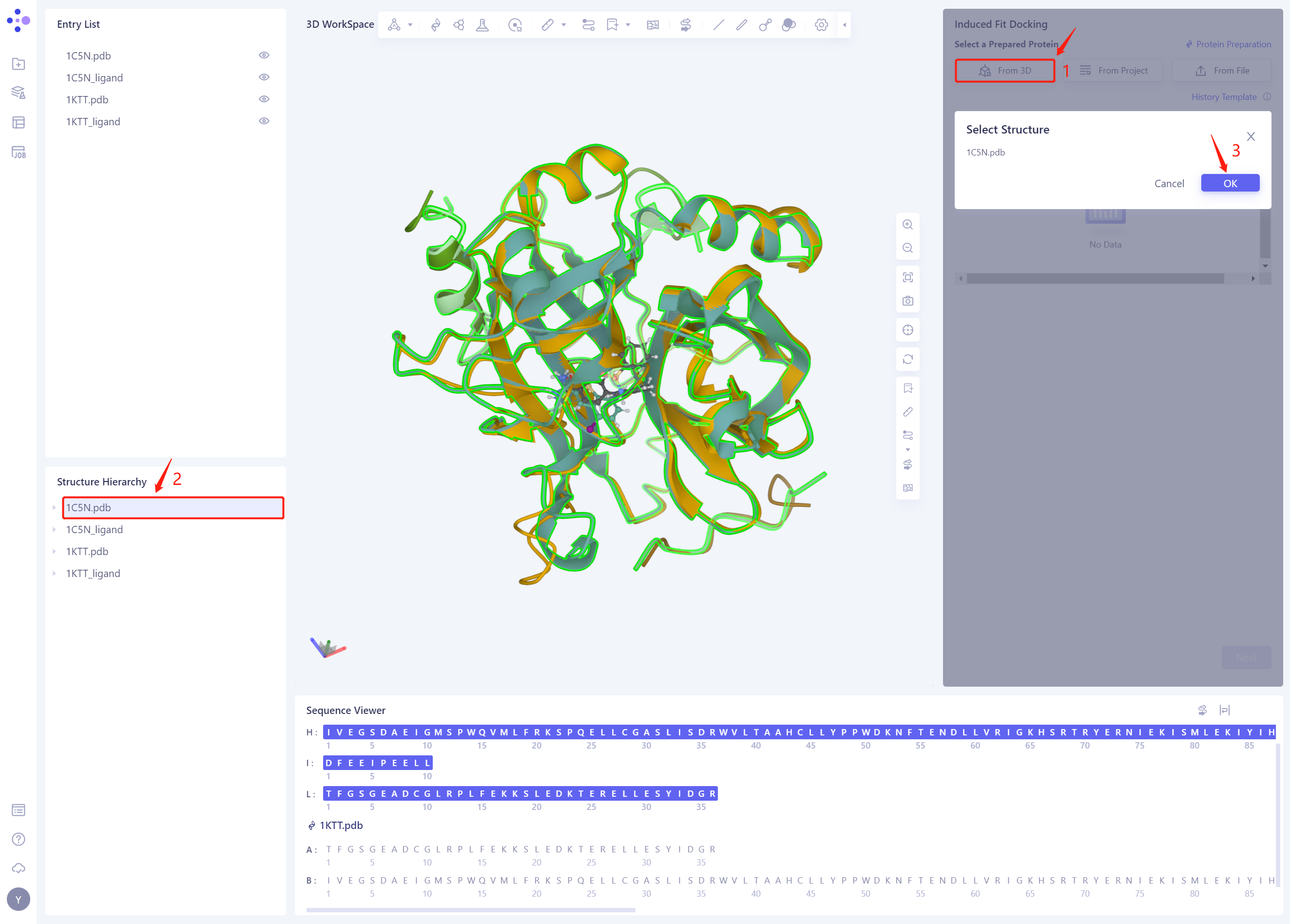
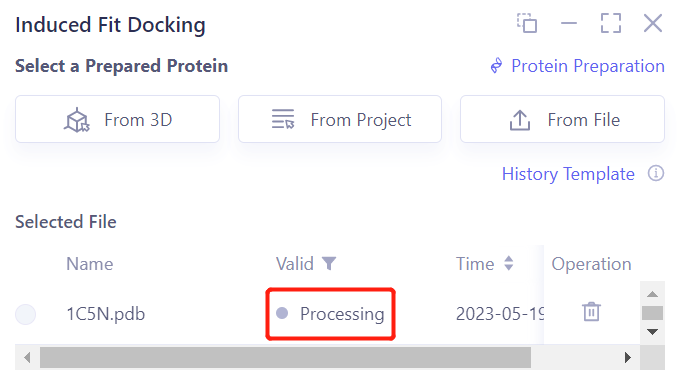
- After clicking OK, the system will automatically check whether the input protein meets the calculation requirements, and the status is “Processing”.
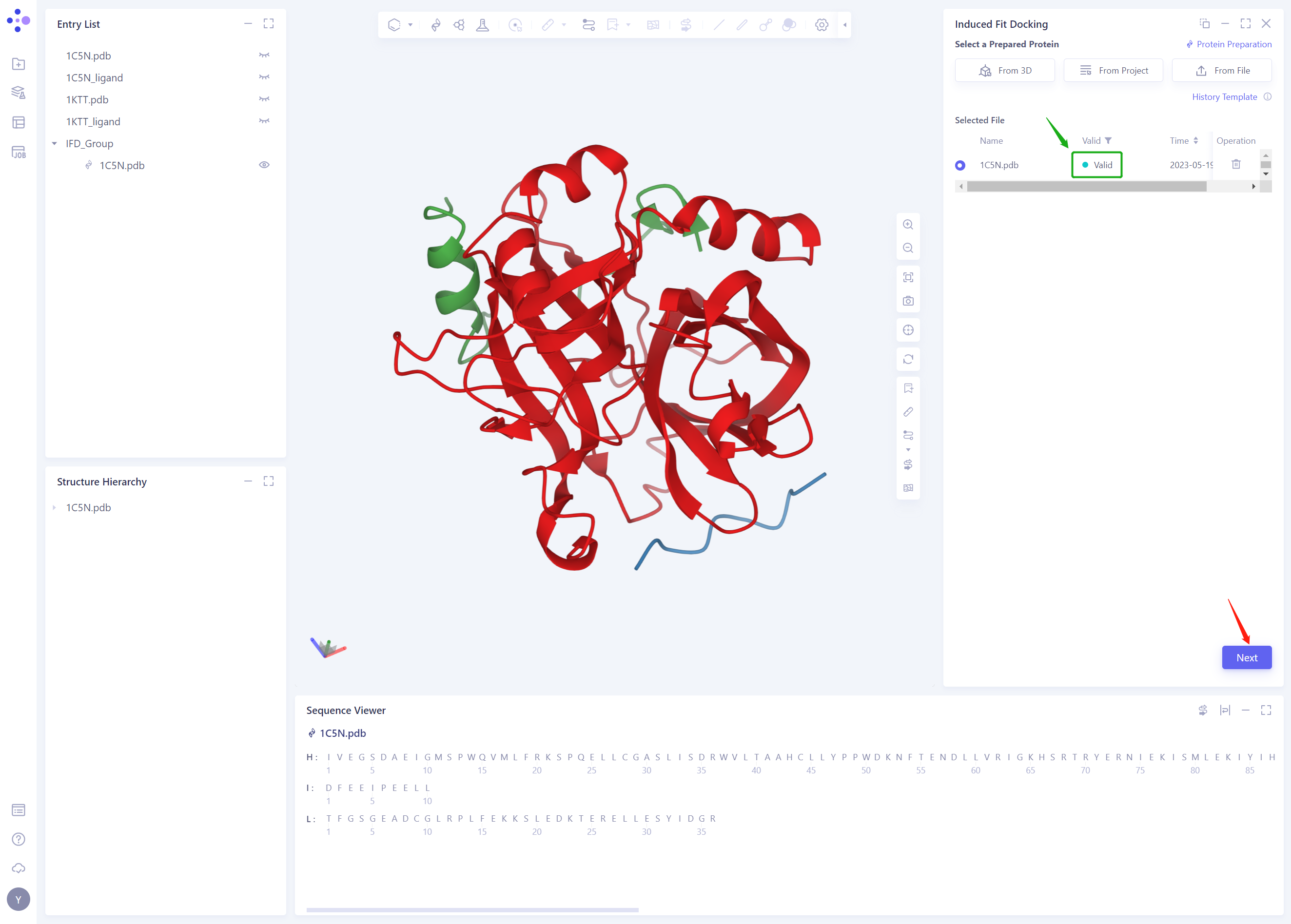
- In less than 1 minute, the system will judge that the protein is “Valid” and click “Next”.
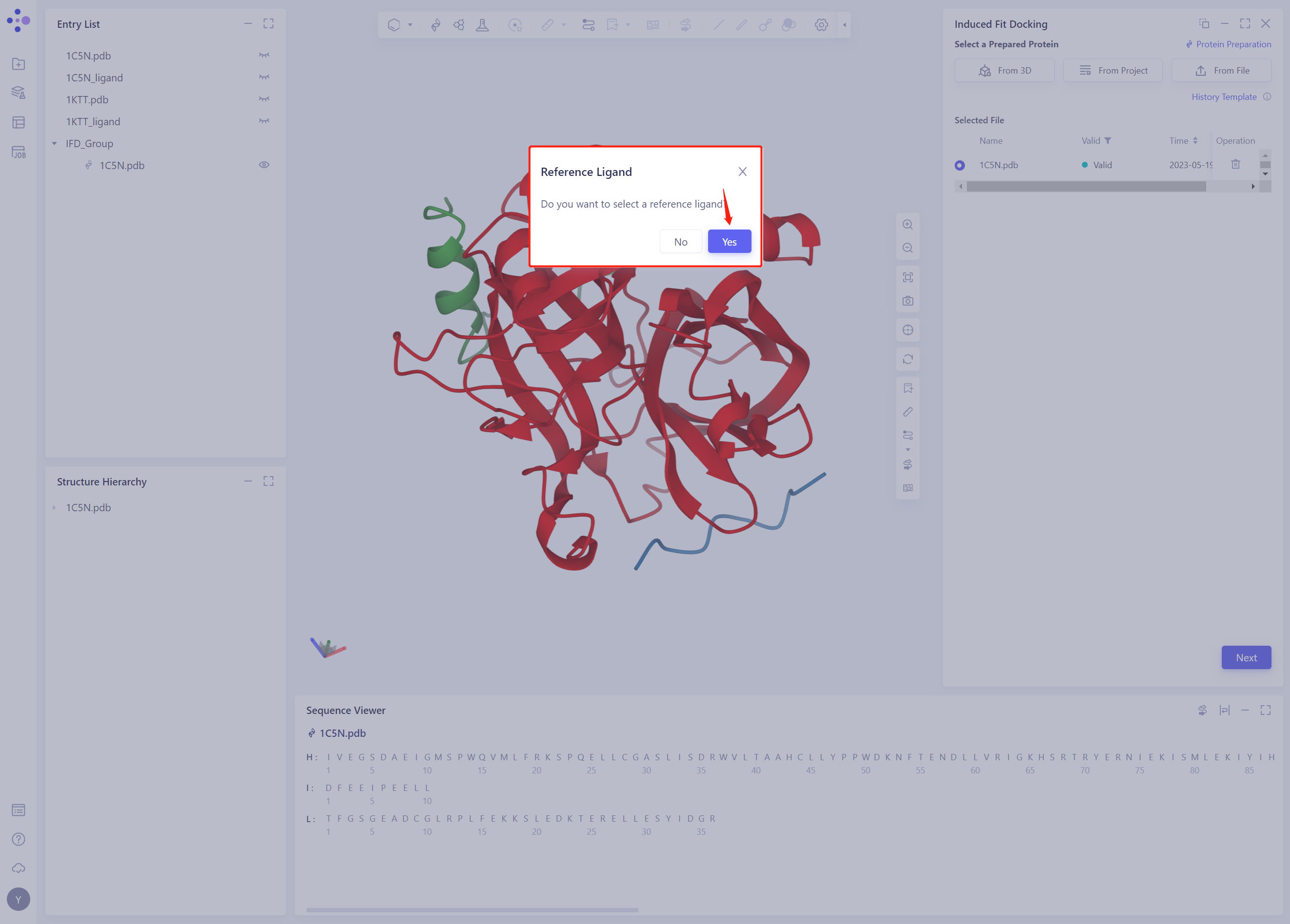
2.3 Select Prepared Reference Ligand 3D
- When prompted whether to select Reference Ligand, click “Yes”;
- Click “From 3D”, select the 1C5N _ Ligand. SDF ligand structure, and click Ok
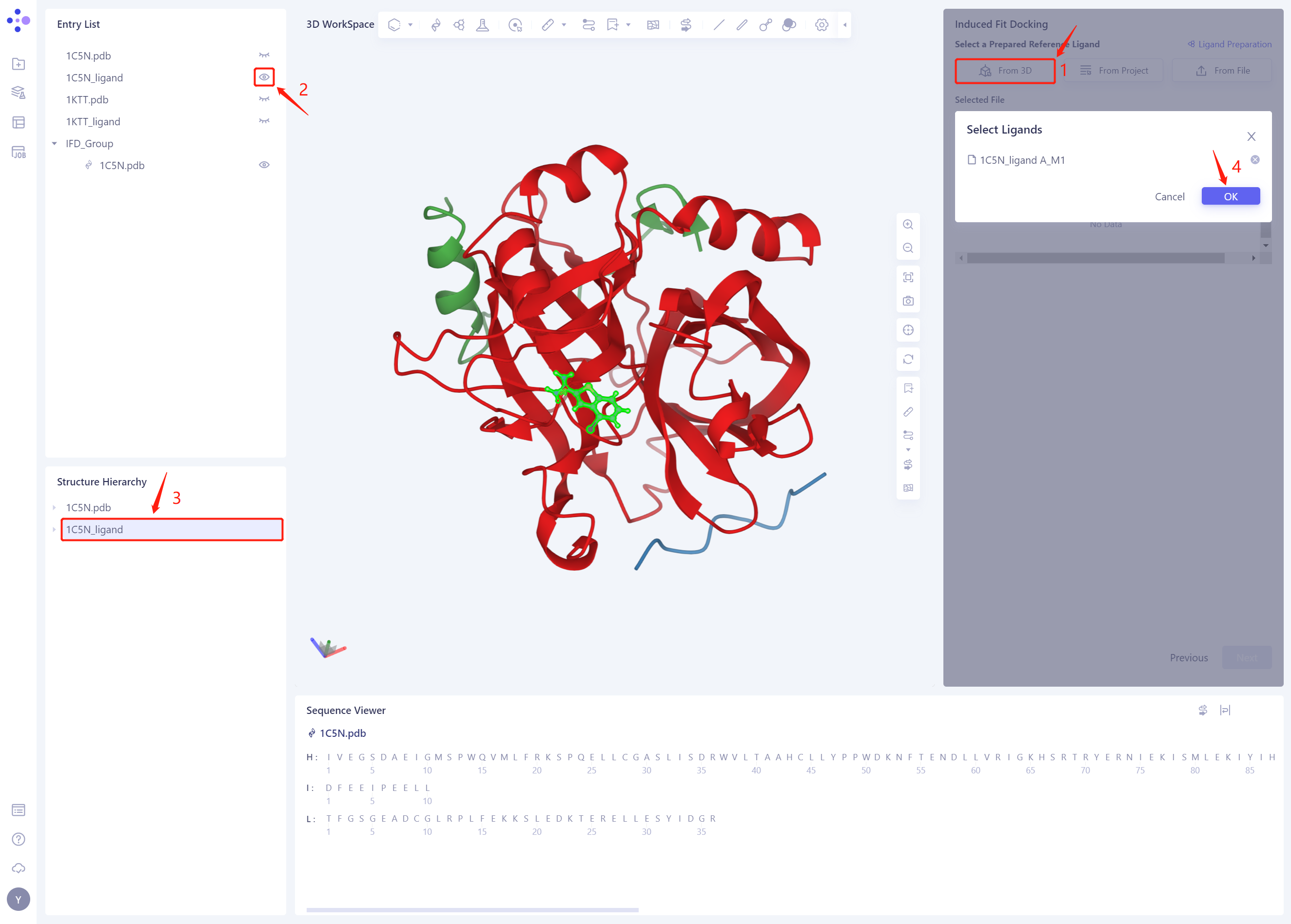
-
The ligand structure of the 1C5N _ Ligand. SDF will be included in the Selected File ”. The system automatically checks whether the Ligand entered meets the calculation requirements and the status is “Processing”, and then the system judges that the Ligand is “Valid”.
-
Click Next
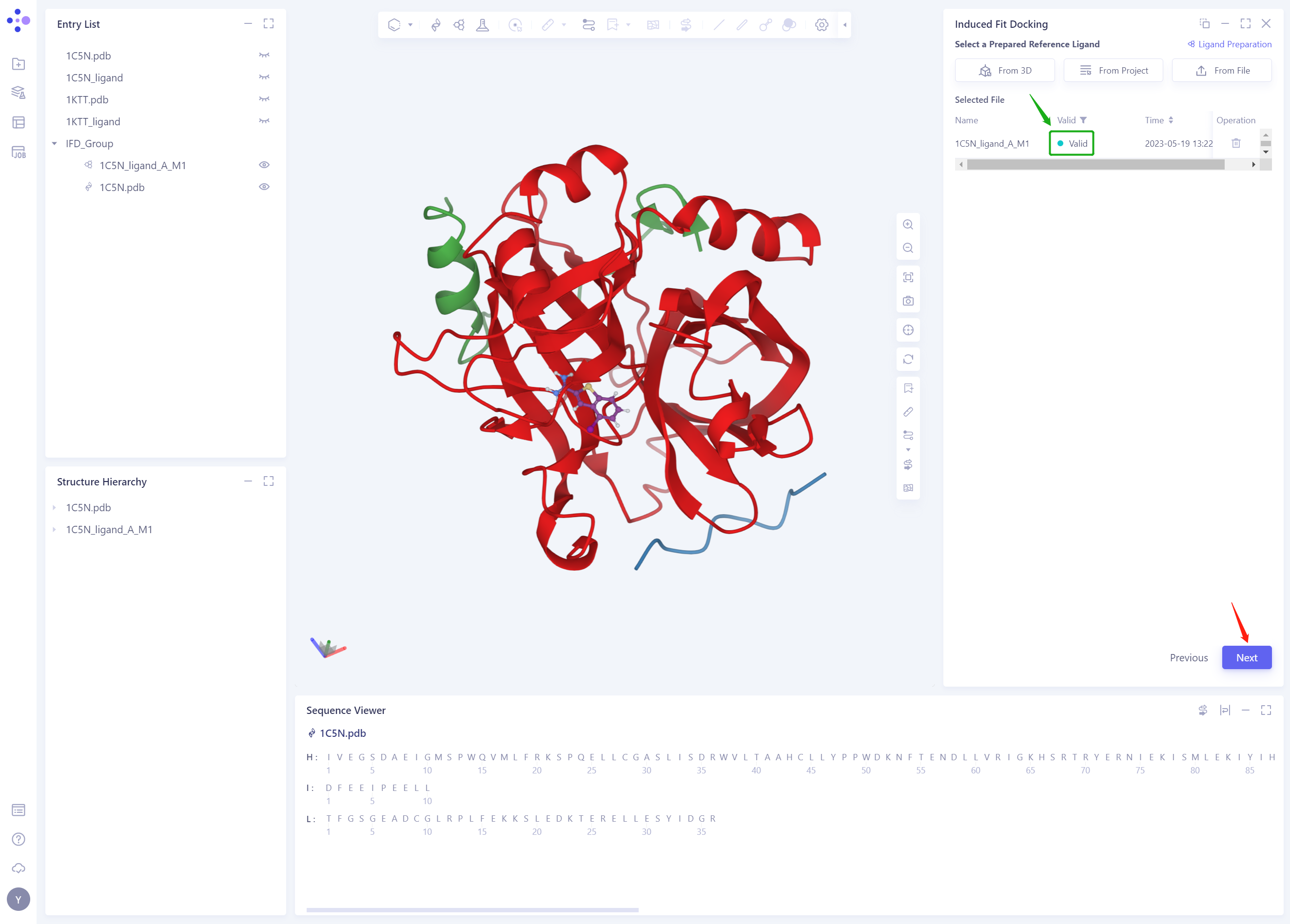
2.4 Select Ligands
- Select the ligand structure to be Docked: click “From 3D”, select the ligand structure of 1KTT _ Ligand. SDF, and click Ok
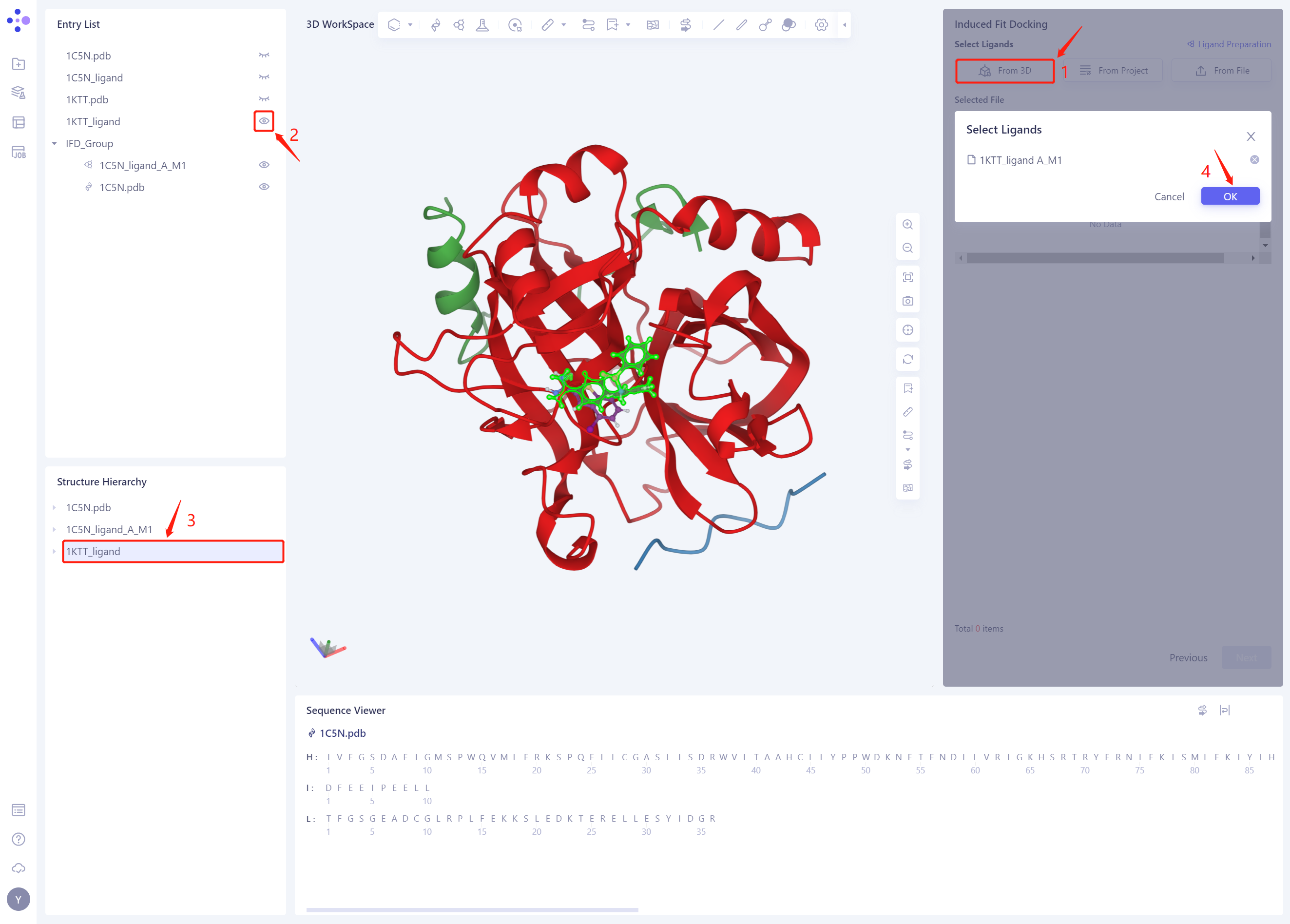
-
The 1KTT _ Ligand. SDF ligand structure will be included in the Selected File ”. The system automatically checks whether the Ligand entered meets the calculation requirements and the status is “Processing”, and then the system judges that the Ligand is “Valid”.
-
Click Next
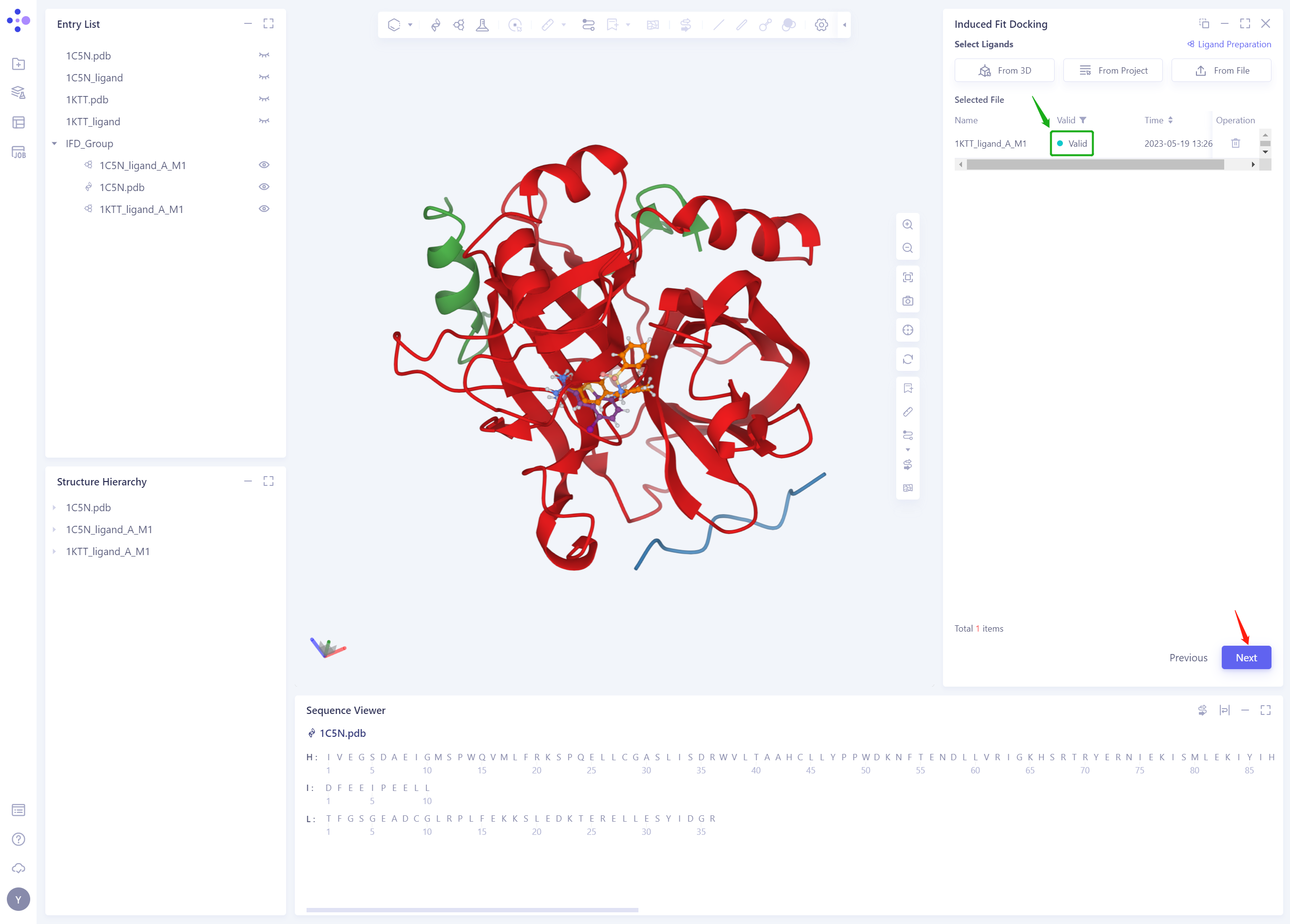
2.5 Pocket Setting
-
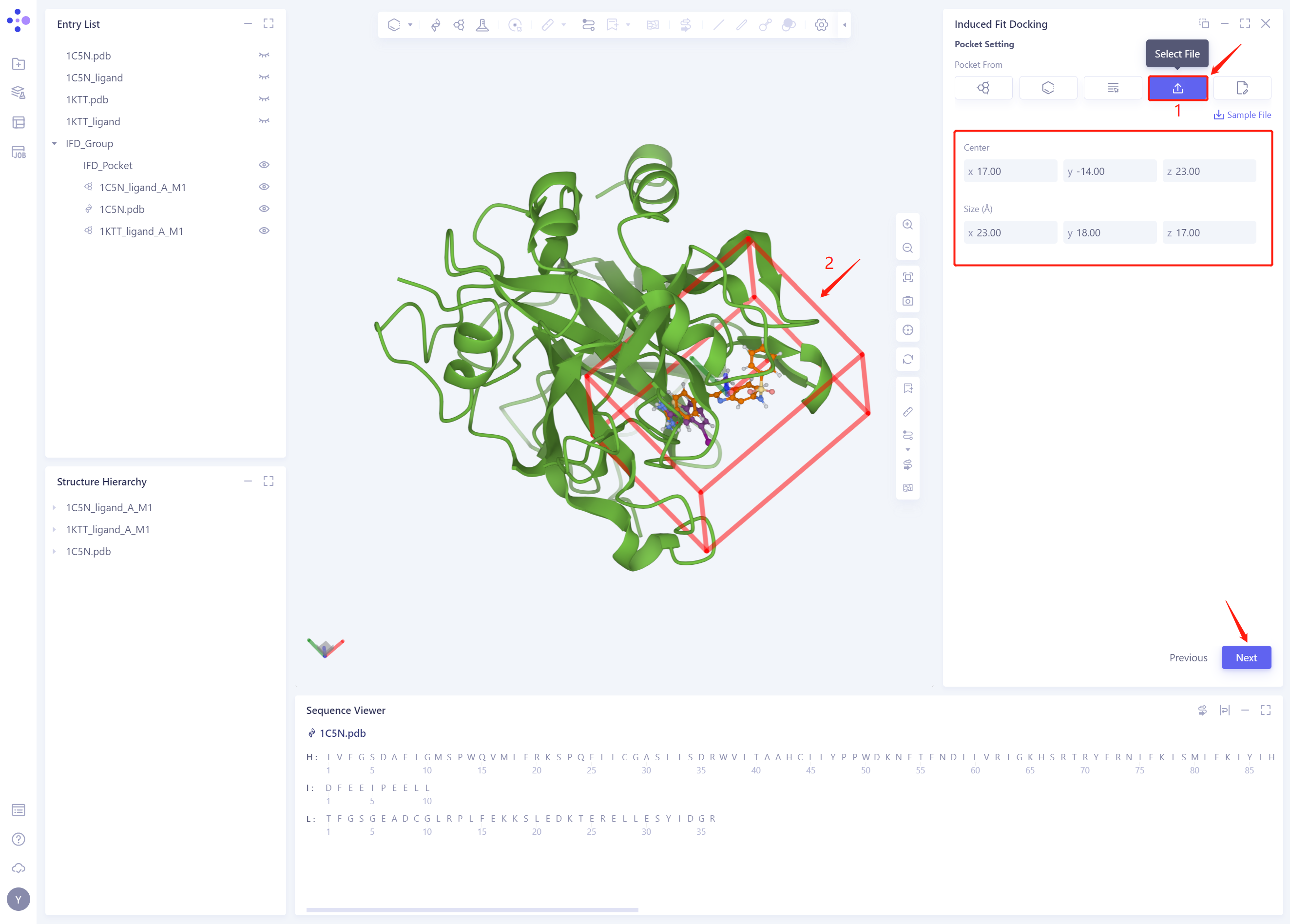
Click “Select File” to set up Pocket with the given Pocket file in this tutorial (you can also try setting up Pocket with Reference Ligand)
-
3D Works pace automatically loads the Pocket, and the system automatically identifies the Center and Size information of the Pocket
-
Click Next to proceed to the next step
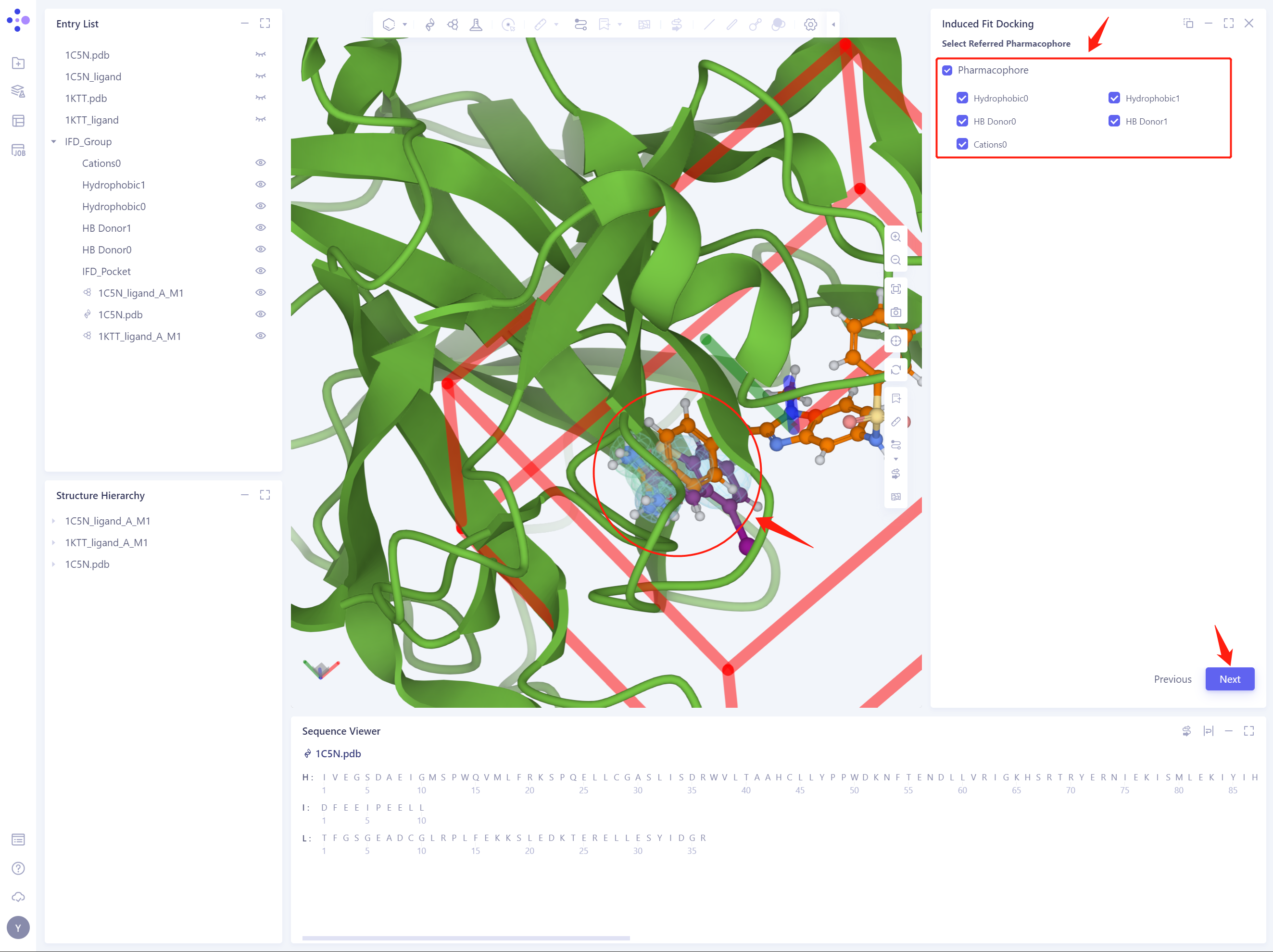
2.6 Select Referred Pharmacophore
- Select all reference pharmacophores and click Next to proceed to the next step
2.7 Confirm and Setting
- Check the selected Receptor protein structure and donor ligand structure, and check the selected reference pharmacophore by clicking Highlight All. The Job Name is named “Thrombin IFD” ”.
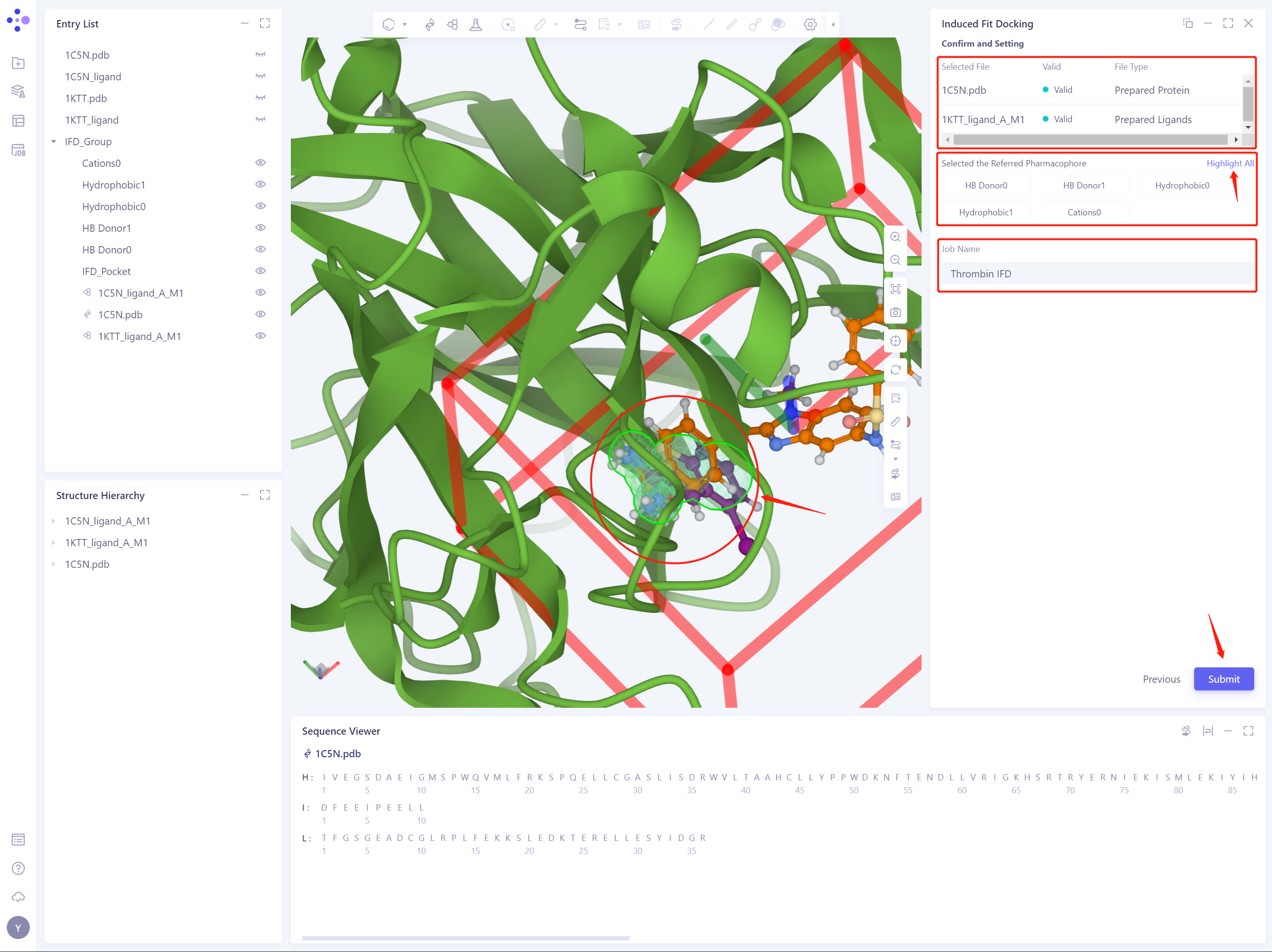
- Click Submit to submit the IFD task
3. Analysis of results
3.1 Entrance
- In the left general menu bar Job → Job List, find the “Thrombin IFD” Calculation Task “and click” Show ”.
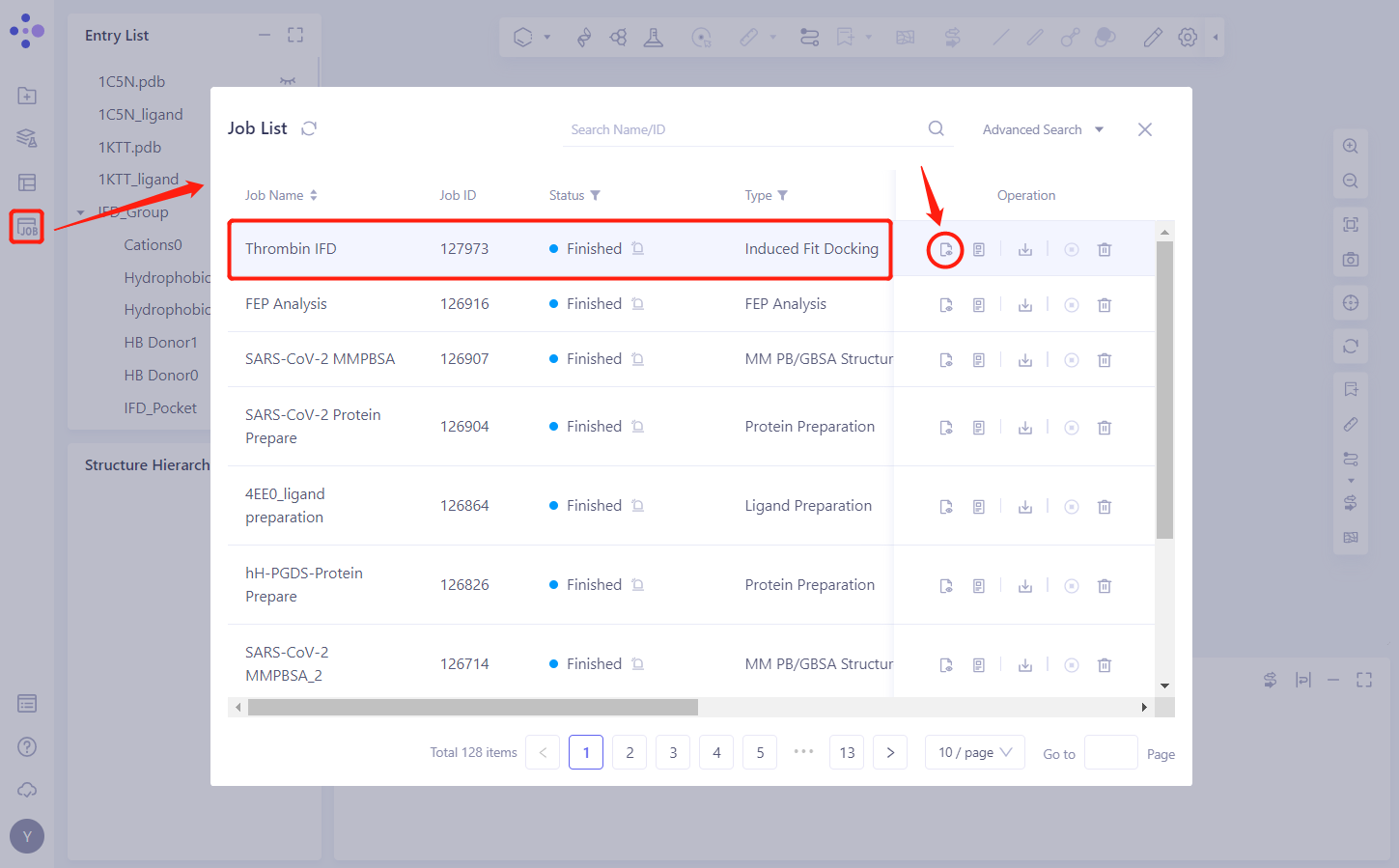
3.2 Results presentation
- Click the “3D” button of the 0th Pose to display the structure of the Induced Fit Docking in the 3D Workspace (here, the protein is shown in red, and the Docking small molecule is shown in green)
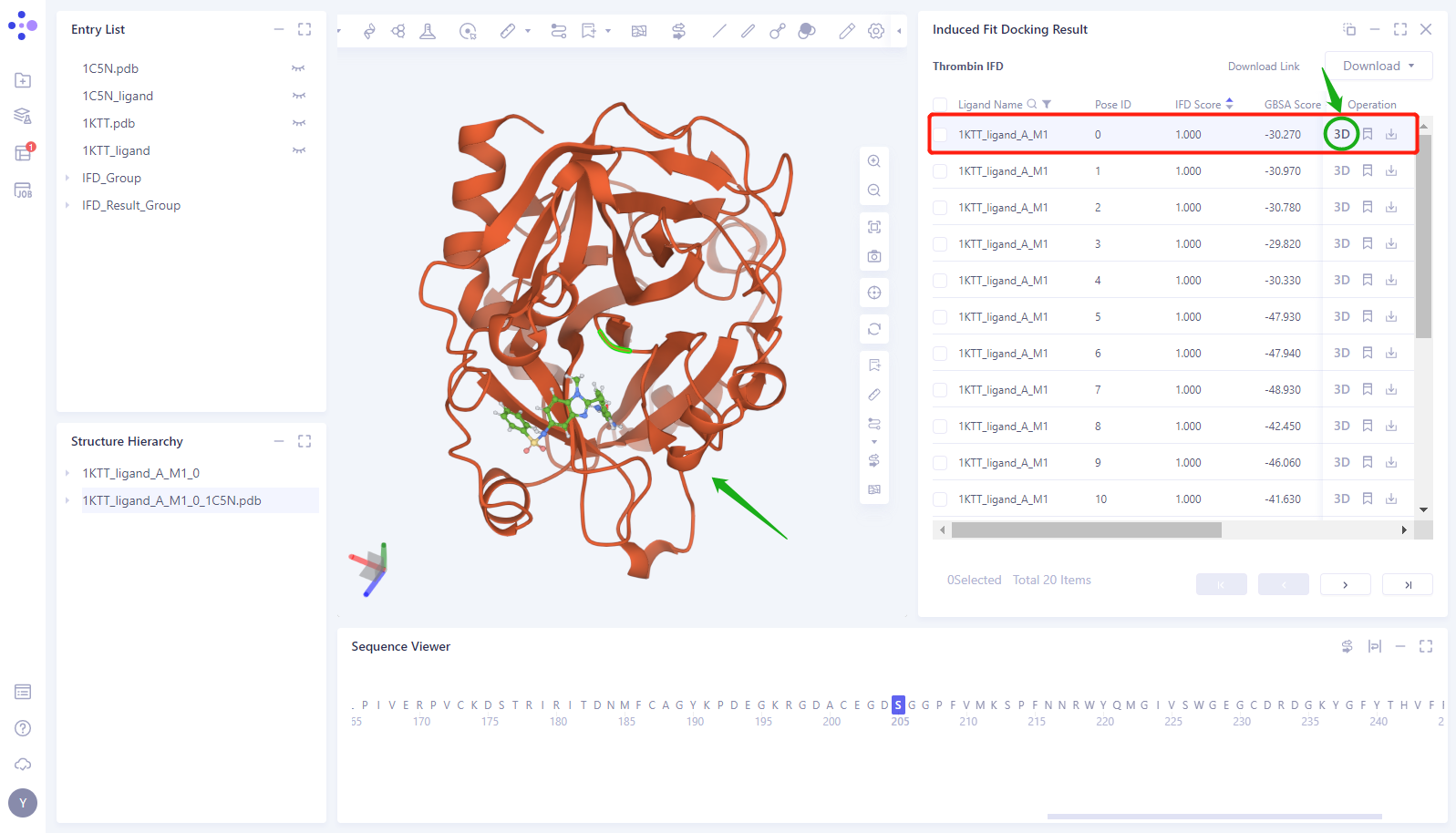
- Click the “Eye” button in the “Entry List” to load the 1KTT and 1KTT _ Ligand structures into the 3D Works pace (proteins are shown in gray, and small molecules in the crystal structure are shown in blue)
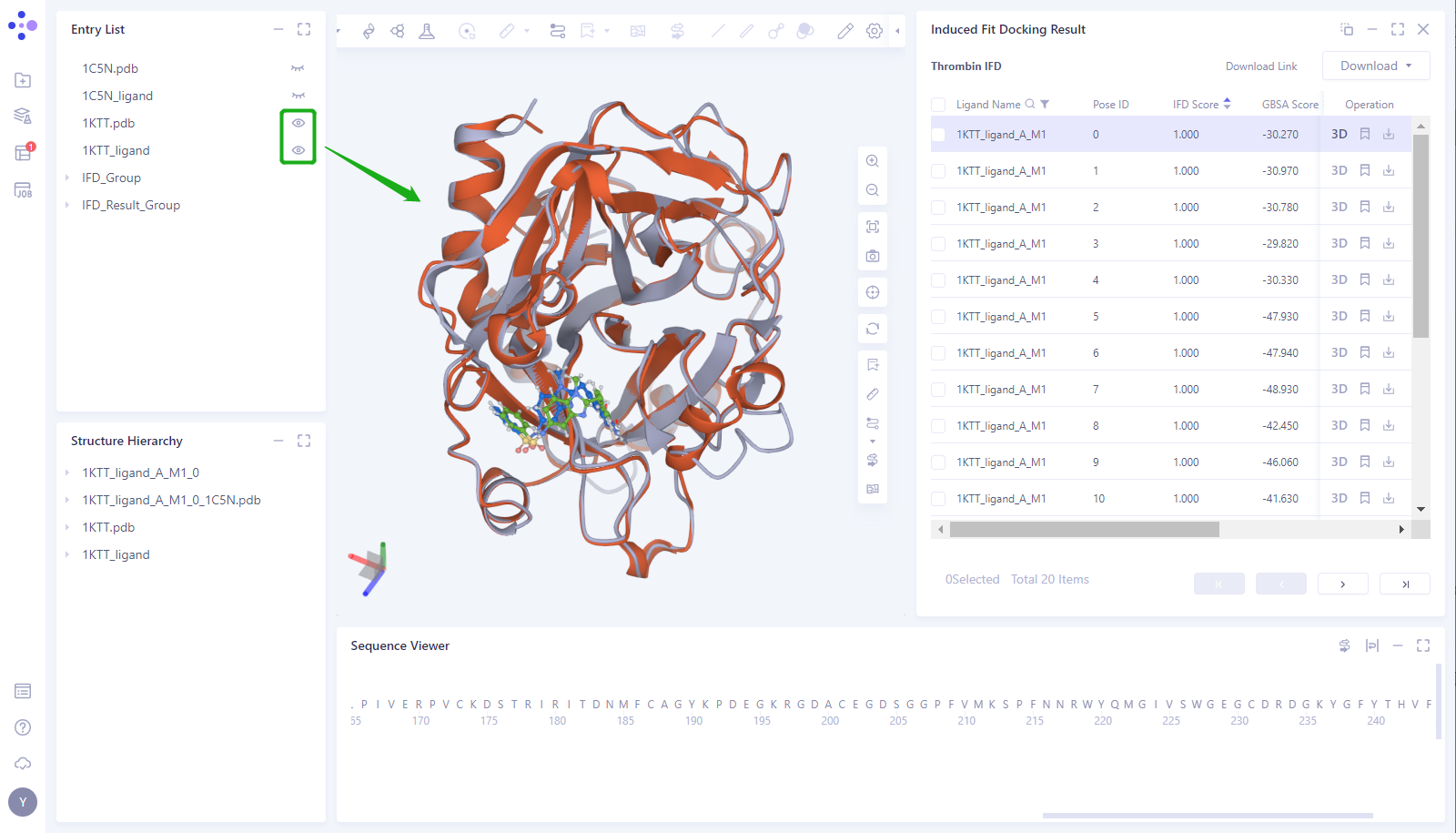
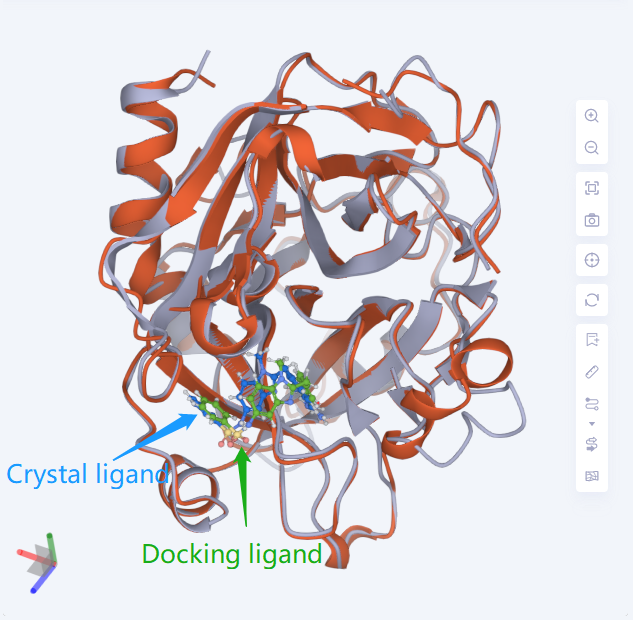
- It can be seen that the position of the small molecule predicted by the Induced Fit Docking is basically consistent with the position of the small molecule (1KTT _ Ligand) in the 1KTT eutectic structure.
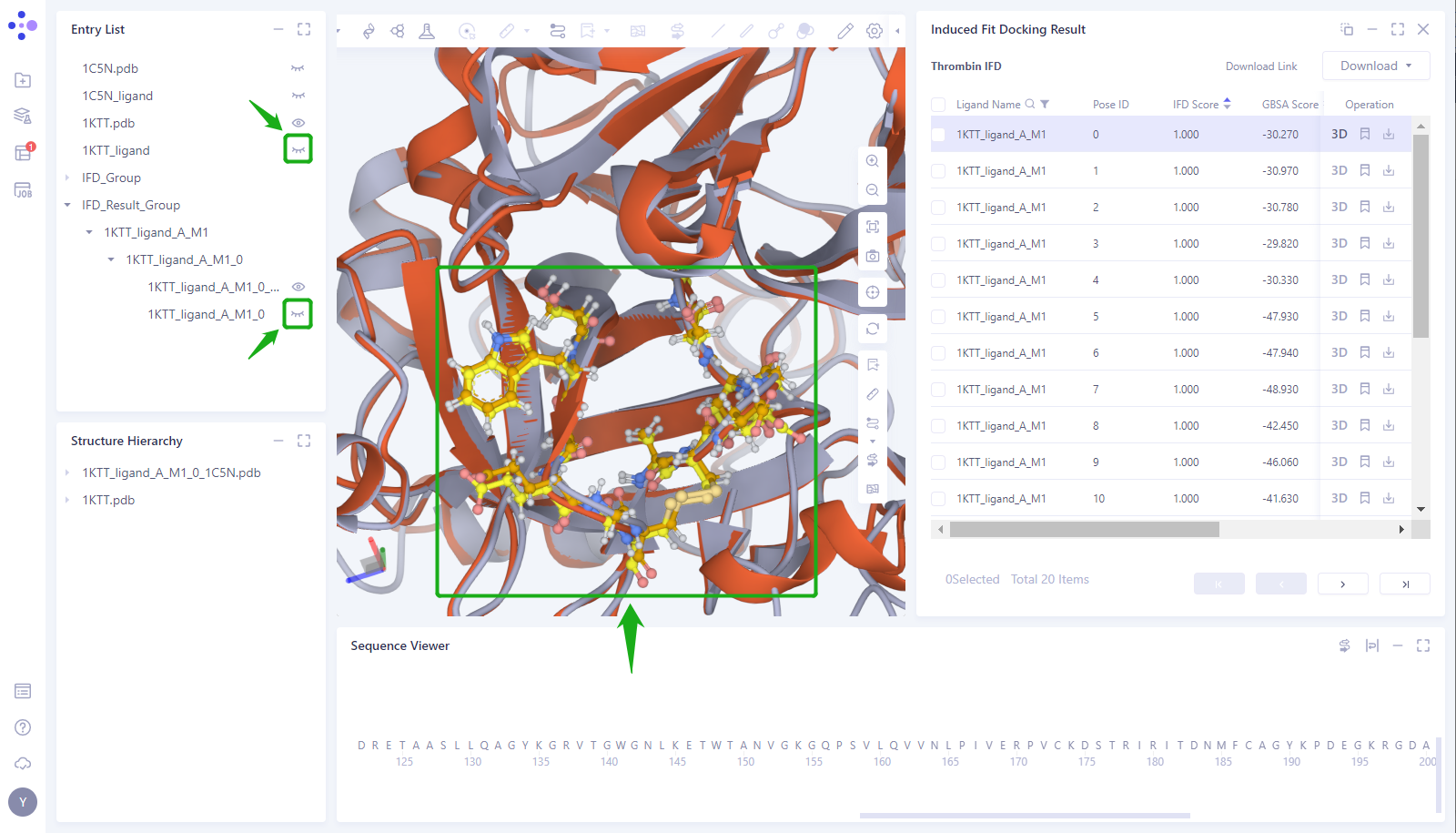
- Select the amino acid in the Pocket around the Ligand and display it as “Ball and Stick”. At the same time, the _ Ligand “of 1KTT and the _ ligand _ A _ M1 _ 0 of 1KTT were predicted by Docking in the Entry List”. It can be seen that in Pocket, the protein structure predicted by Induced Fit Docking is consistent with the structure of 1KTT crystal protein, and the position of amino acid side chain in Pocket is highly consistent.
4. Sum up
In this case, the thrombin inhibitor complexes 1 KTT and 1C5N were used as examples to study the “induced fit” mechanism of the co-crystal compound molecule of 1 KTT binding to the 1 C5N protein target by using the Uni-IFD module. The results of Induced Fit Docking show that Uni-IFD module can simulate the “induced fit” sample when drug molecules bind to the target, and accurately predict the binding mode between drug molecules and the target.
5. References
[1] Structure-Based Design of Novel Potent Nonpeptide Thrombin Inhibitors. J. Med. Chem. 2002, 45, 1757-1766
[2] Structural basis for selectivity of a small molecule, S1-binding, submicromolar inhibitor of urokinase-type plasminogen activator. Chemistry & Biology 2000, 7:299–312