Protein Selection, Ligand Selection and Pocket Setting
1. Protein Selection
Docking and MM PB/GBSA modules do not have "Get PDB".
| Protein import method | Icon | Operation | Interface display | Note |
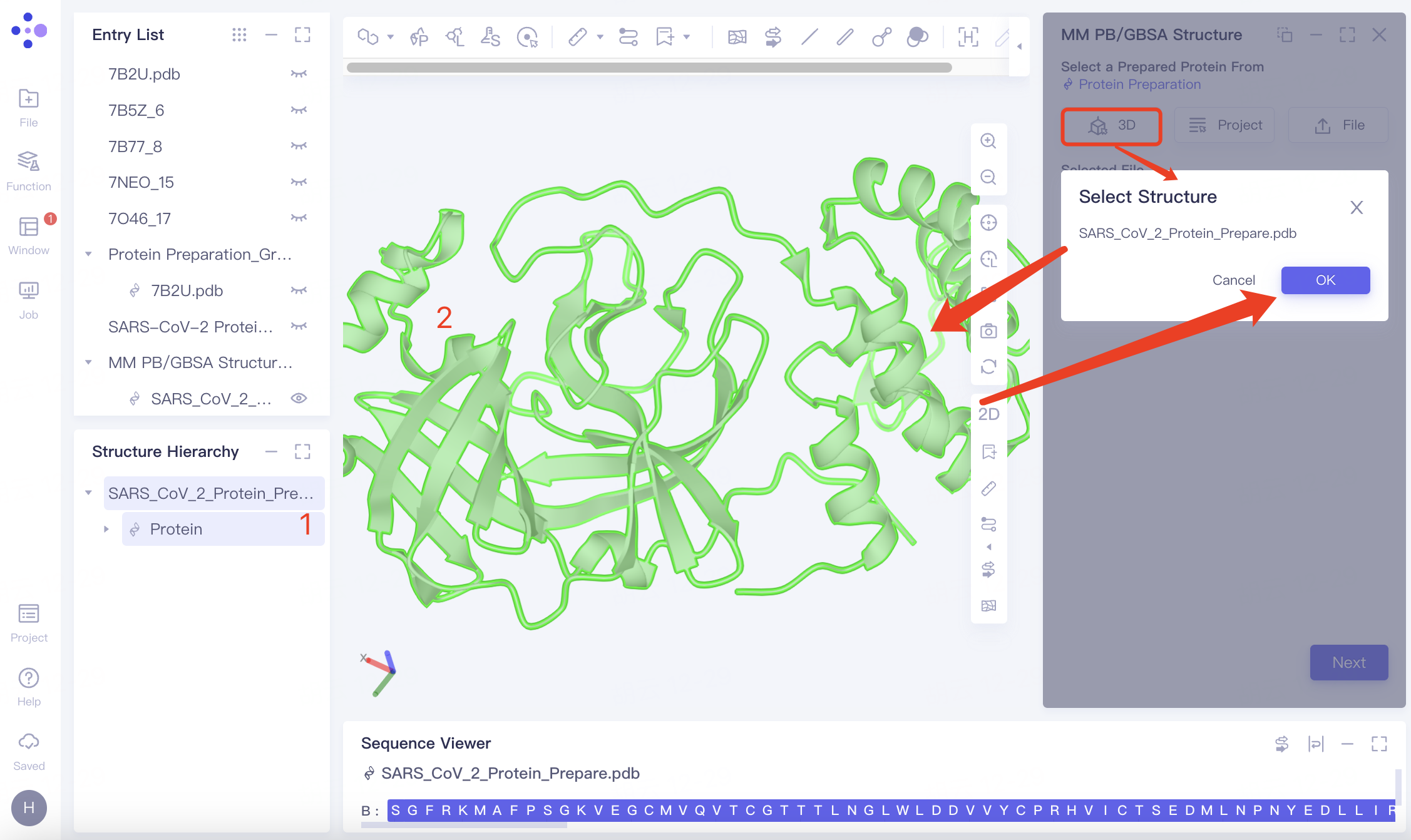
| Select from 3D Workspace | Click the "3D" button and select the protein from the 3D Workspace or the Structure Hierarchy |
| —— | |
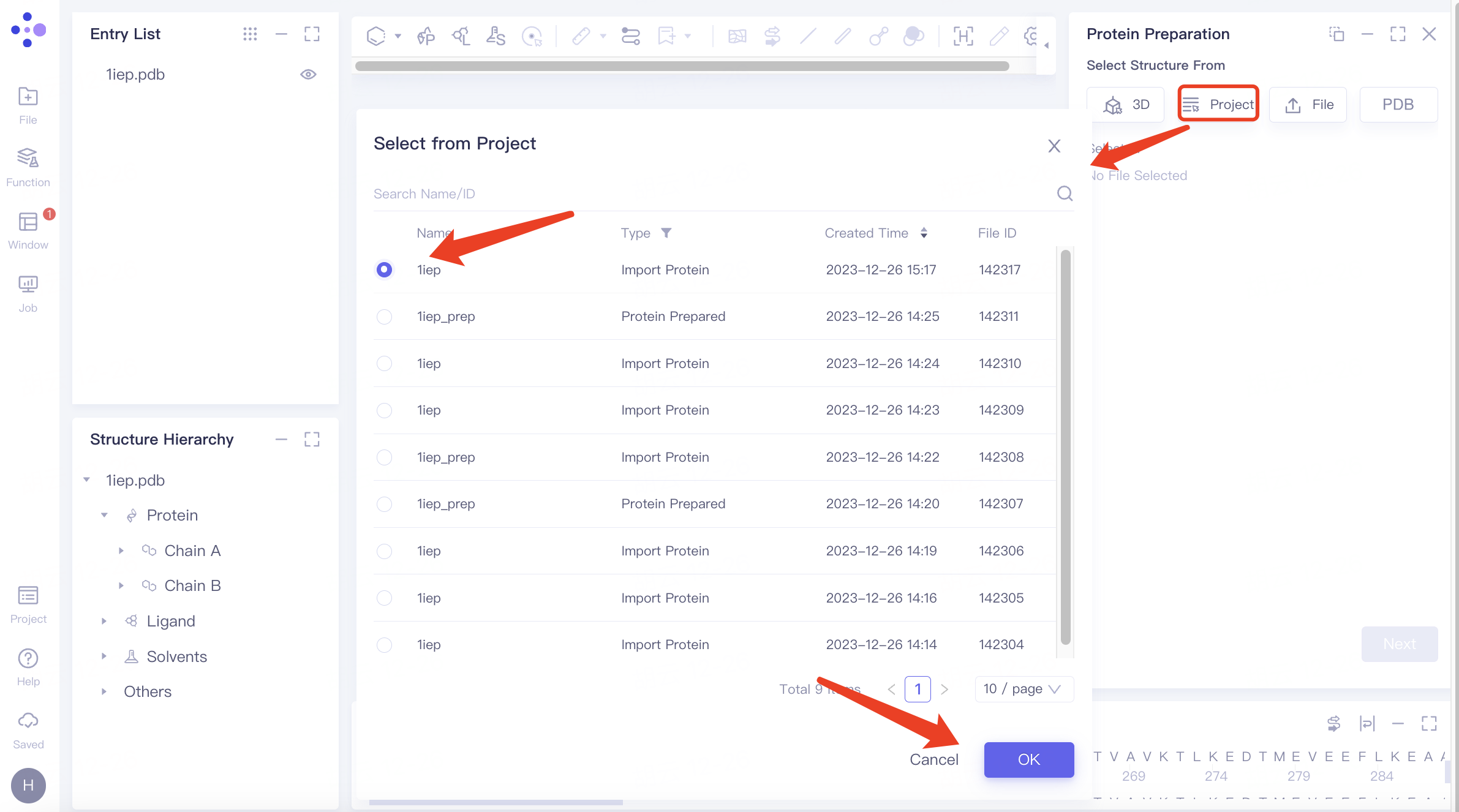
| Select from Project | Click the "From Project" button and select the protein from previous projects |
| You can locate files using the Search Name/ID feature by file name or ID, and protein files can be selected by type. | |
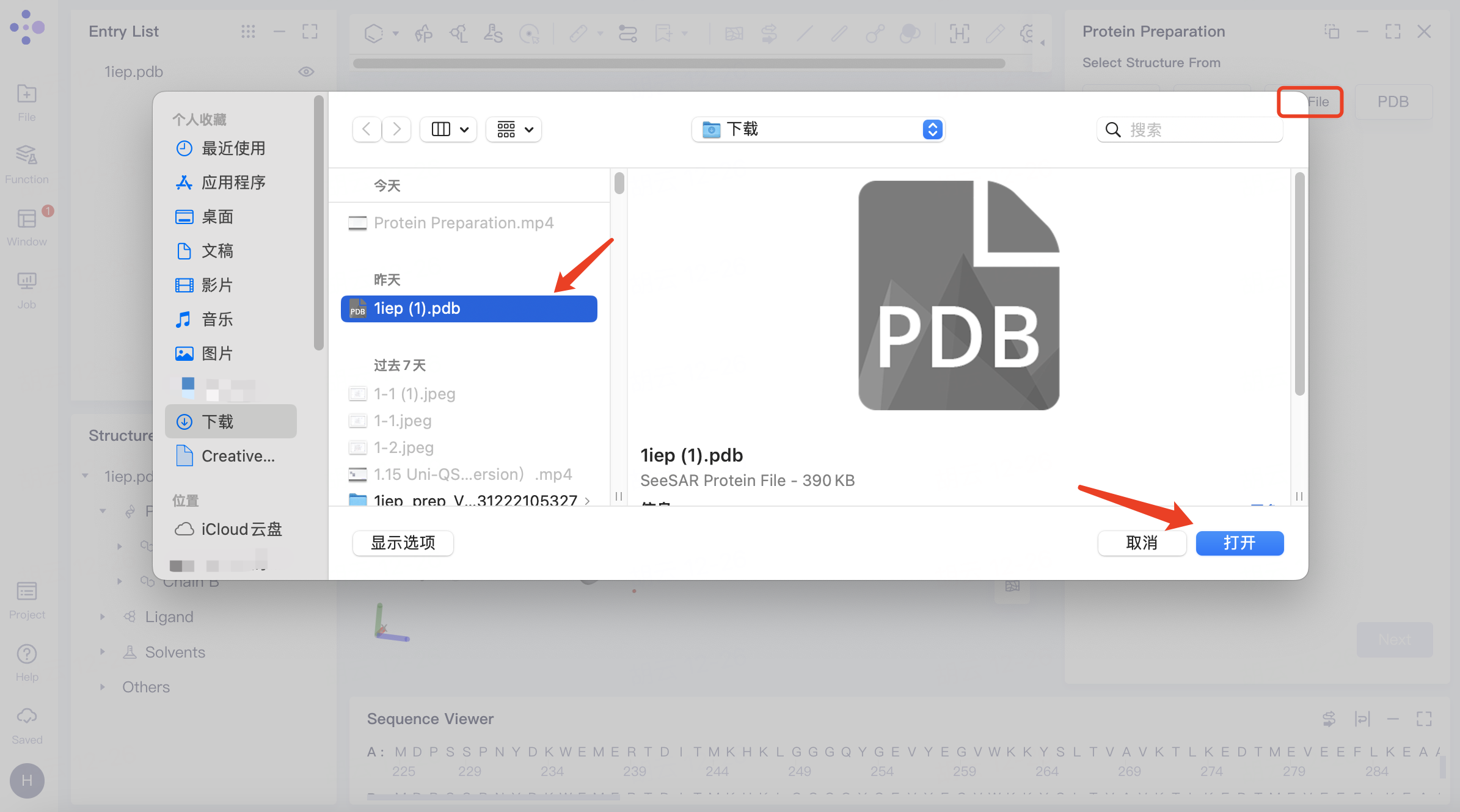
| Select from File | Click "From File" to upload a protein from a local file |
| The system supports .pdb file uploads. | |
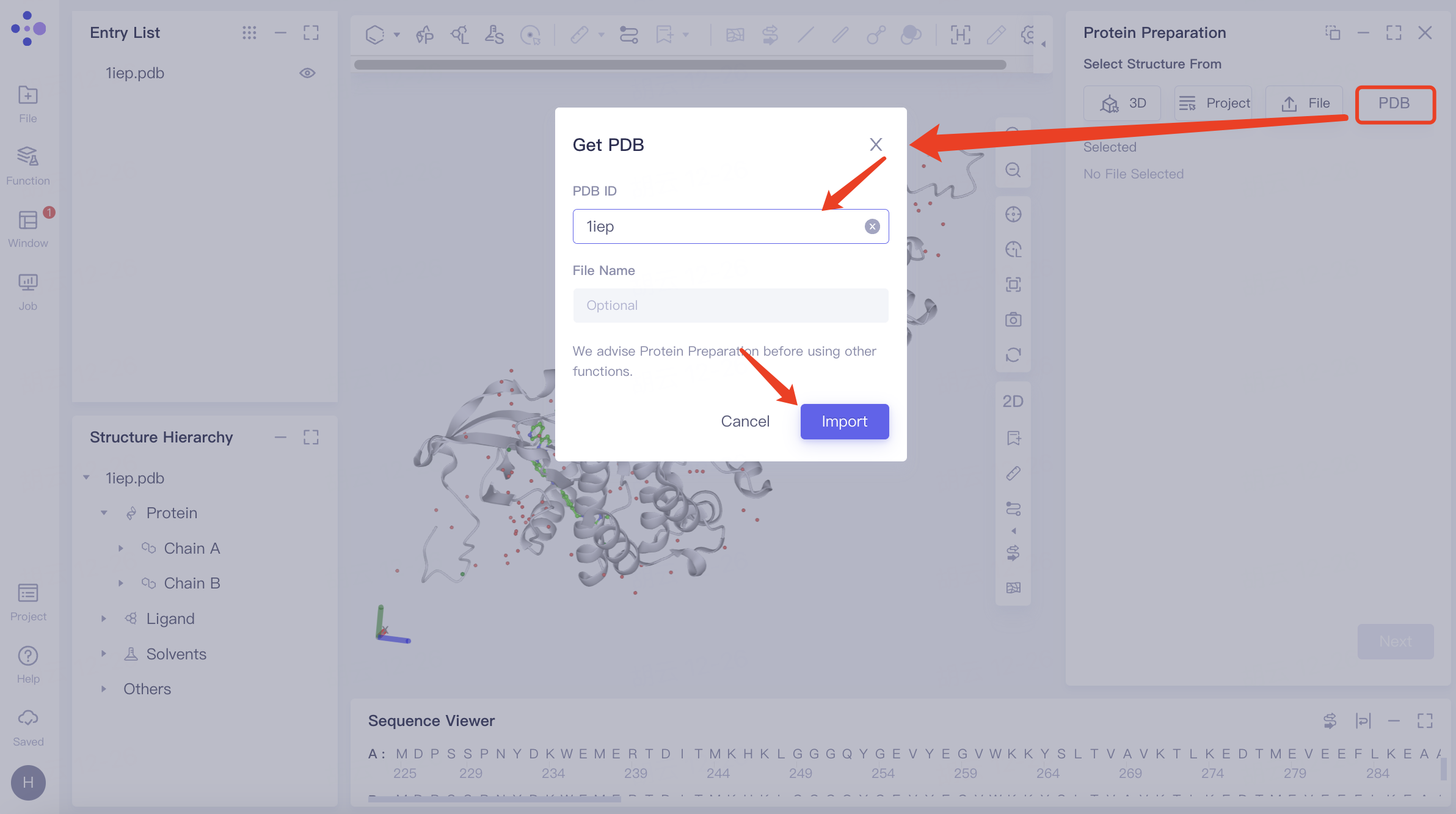
| Get PDB | Search protein by its PDB ID and then upload |
| —— |
2. Ligand Selection
Ligand preparation and MM PB/GBSA modules do not have "Select from Database", while Docking and MM PB/GBSA modules do not have "Select from SMILES".
| Protein selection method | Icon | Operation | Interface display | Note |
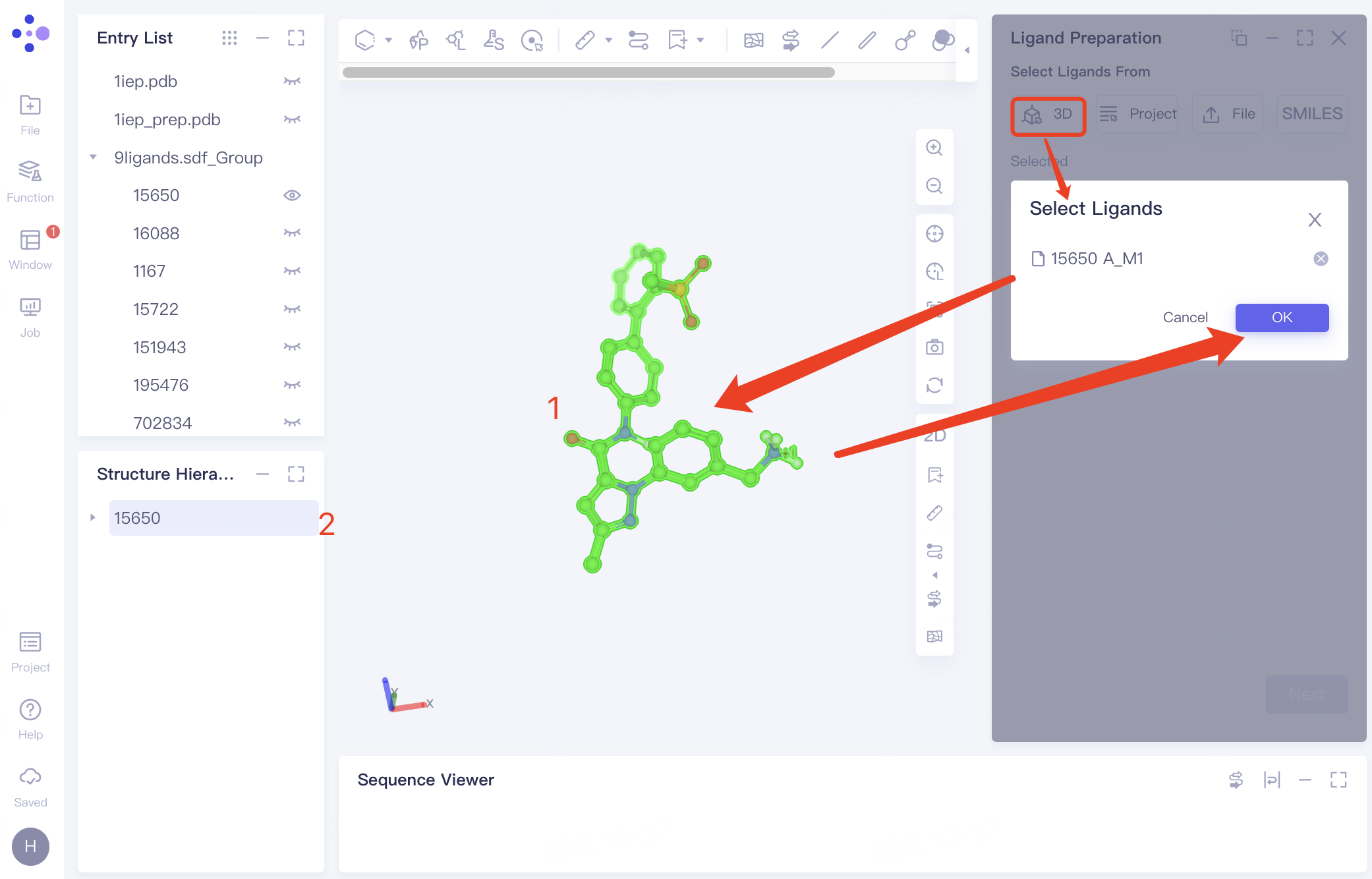
| Select from 3D Workspace | Click the "3D" button and select the ligand from either the 3D Workspace or the Structure Hierarchy |
| ||
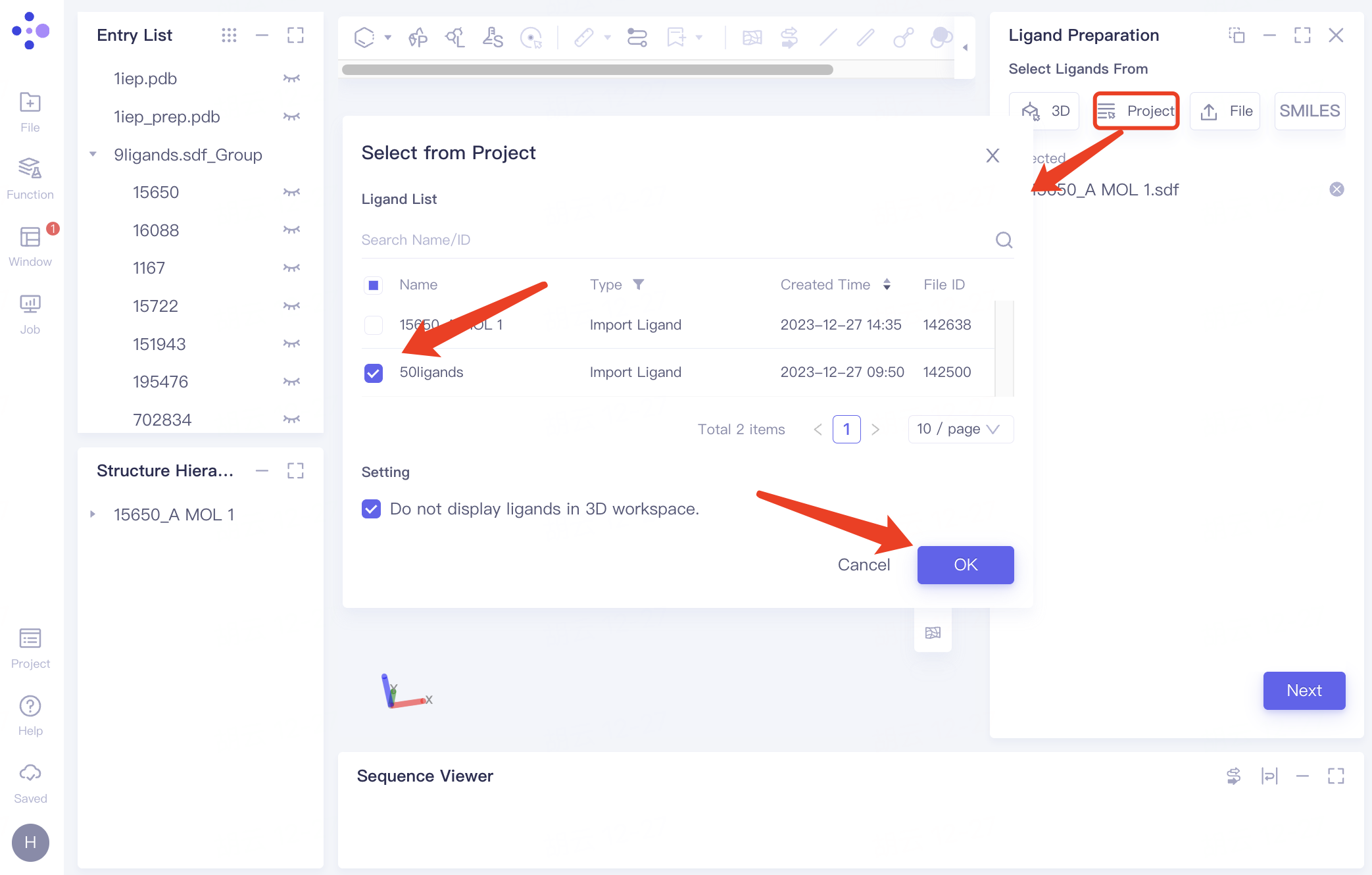
| Select from Project | Click the "Project" button to select ligands from previous projects |
| You can locate files using the Search Name/ID feature by file name or ID, and adjust the 'Type' setting to specifically select ligand files. | |
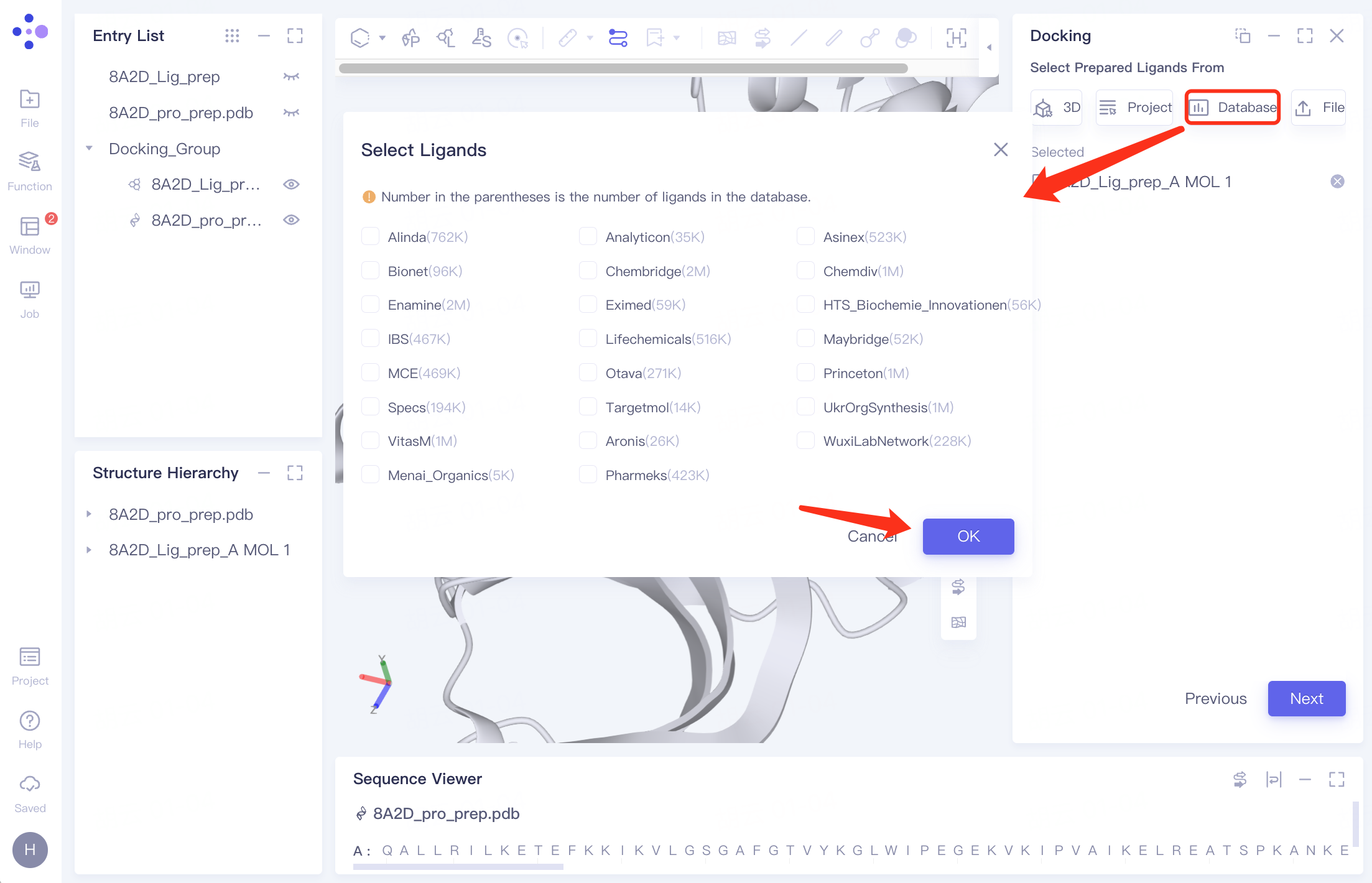
| Select from Database | Click the "Database" button and select the required molecular dataset from the "Select Ligands" interface that pops up |
| Choose from over 22 million molecules in our libraries for screening. Details about the database are provided in the table below. | |
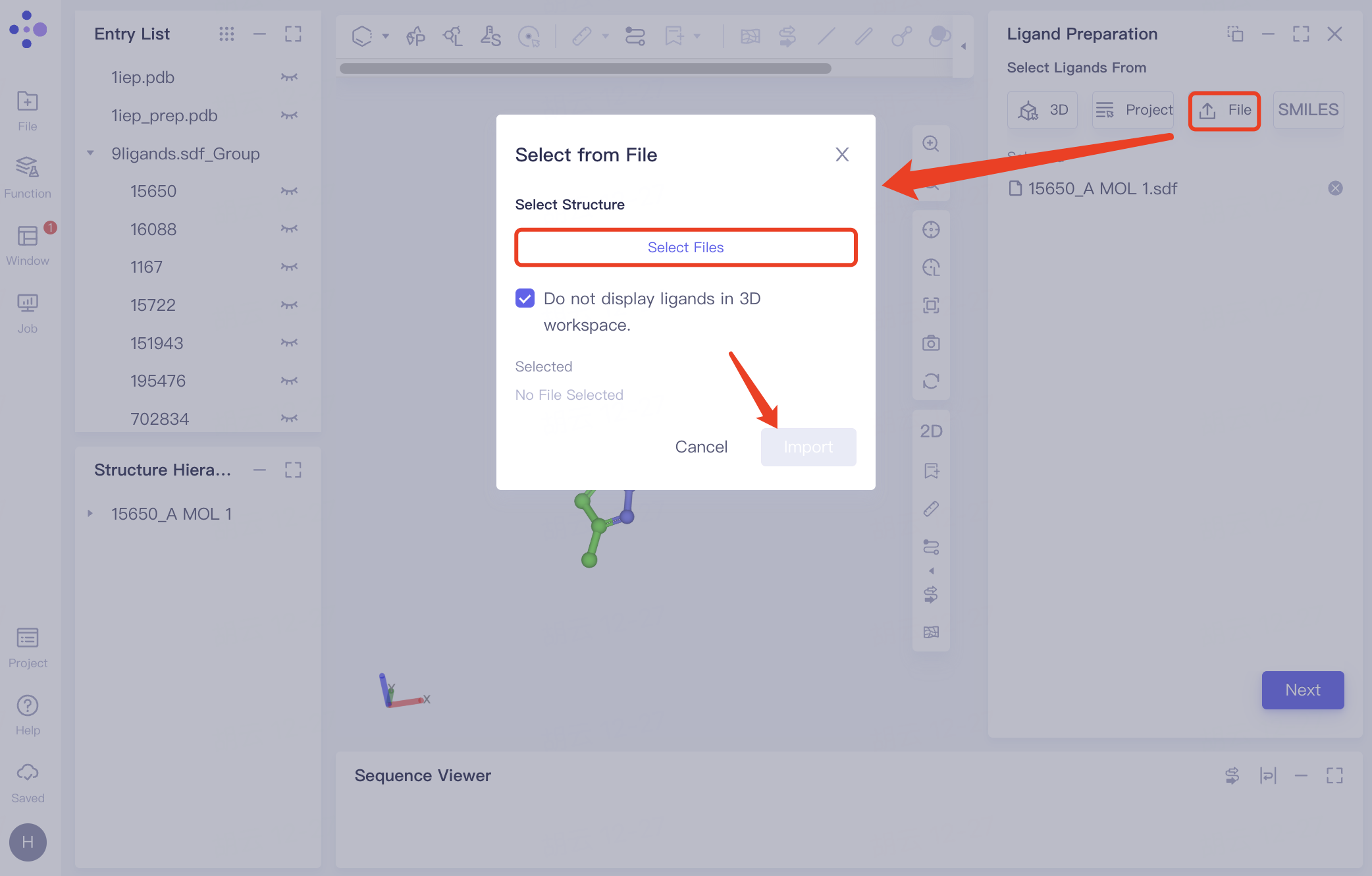
| Select from File | Click "File" to upload a ligand from a local file |
| The system supports .sdf, .mol, .smi files uploads. | |
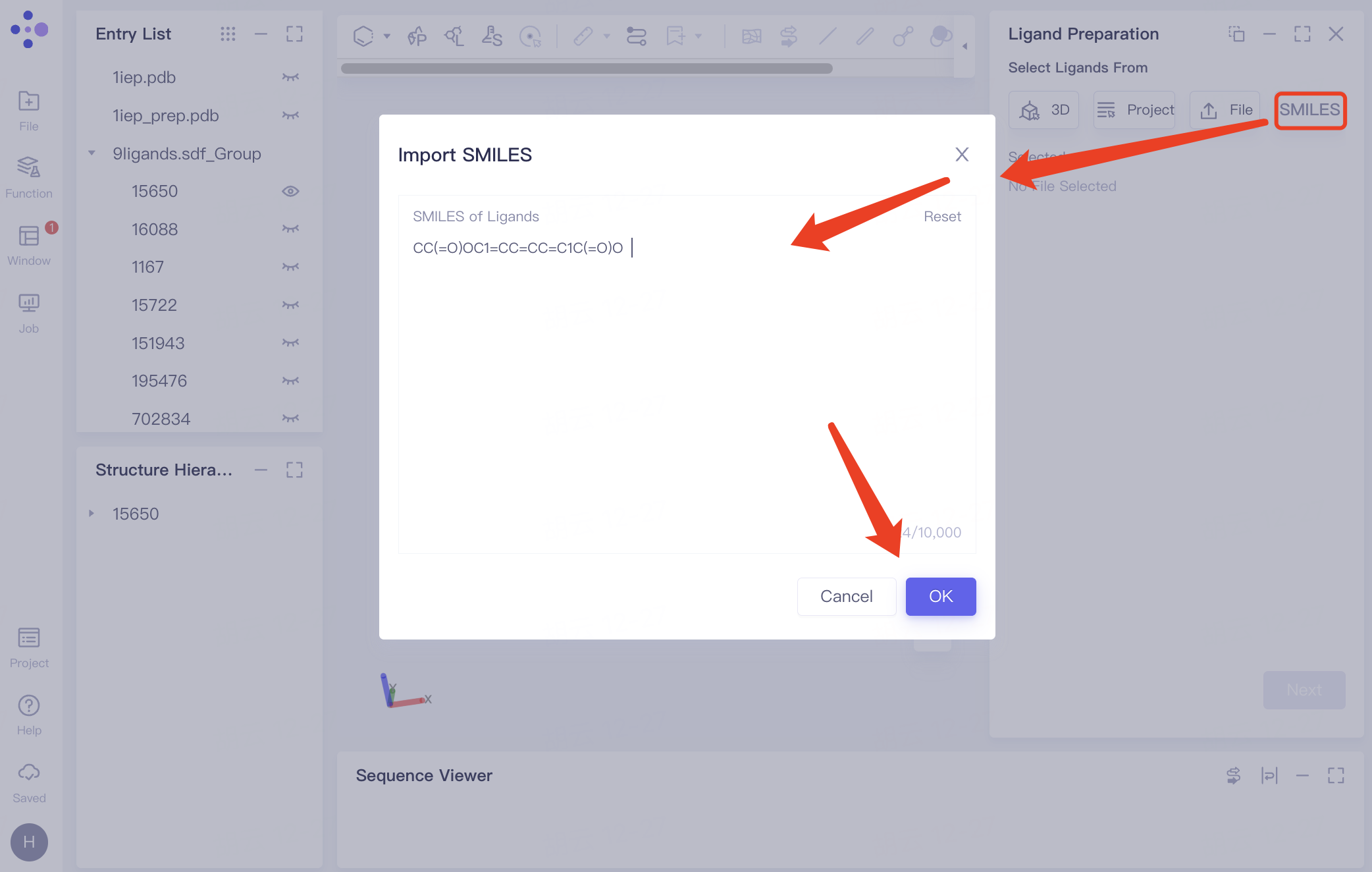
| Select from SMILES | Click "SMILES" and enter the SMILES molecular formula |
| Support up to 10,000 SMILES. |
| ID | Dataset | Description |
| 1 | Alinda(1M) | A compound brand originating from Russia, specializing in the production and sales of new screening compounds |
| 2 | Analyticon(62K) | A natural product brand originating from Germany, focusing on natural product extraction and analogue synthesis work |
| 3 | Aronis (34K) | A compound brand originating from Russia, with more than 20 years of experience in compound supply, sufficient inventory, and high cost performance |
| 4 | Asinex (987K) | A brand originating from the US, dedicated to the research and development of lead compounds and molecular building blocks for more than 20 years |
| 5 | Bionet (119K) | A brand originating from the United Kingdom with over 20 years of organic synthesis experience |
| 6 | Chembridge (3M) | A compound brand originating from the US, with a variety of popular compound libraries such as diversity libraries and large ring libraries |
| 7 | Chemdiv (2M) | One of the world's largest compound brands, with over 5,000 compound skeleton structures and over 100 compound libraries |
| 8 | Enamine (4M) | A compound brand originating from Ukraine, with strong compound research and development capabilities, and two types of products: high cost-effective compounds and high-value compounds |
| 9 | Eximed (81K) | A compound brand originating from Ukraine, dedicated to providing high-throughput screening compounds and related services for nearly 20 years |
| 10 | HTS_Biochemistry_Innovations (88K) | A compound brand originating from Germany, dedicated to developing unique compounds for pharmaceutical, agricultural, and biotechnology companies |
| 11 | IBS (705K) | InterBioScreen, a compound brand originating from Russia, has a variety of natural products and derivatives |
| 12 | Lifechemicals (796K) | A compound brand originating from Canada, with more than 2,900 compound skeleton structures and relatively complete compound specifications |
| 13 | Maybridge (68K) | A compound brand originating from the United Kingdom, under Thermofisher, with a small number of products and a large inventory of each product |
| 14 | MCE (469K) | MedChemExpress |
| 15 | Otava (409K) | A Canadian compound brand specializing in the development and production of specialty compounds, biochemicals, and bioanalytical reagents |
| 16 | Princeton (2M) | A compound brand originating from the US, designing unique small molecule compounds for drug research and development for more than 20 years |
| 17 | Specs (284K) | A compound brand originating from the Netherlands with obvious price advantages |
| 18 | Targetmol (20K) | Advantageous products are known active compounds and natural products, with high cost performance |
| 19 | UkrOrgSynthesis (2M) | A young and rapidly growing compound brand originating from Ukraine, mainly used for high-throughput screening and drug discovery |
| 20 | VitasM (2M) | A compound brand originating from the US, with a shipping center in Hong Kong and moderate prices |
| 21 | WuxiLabNetwork (565K) | A CRO company originating from China with rich experience in compound synthesis |
| 22 | Menai_Organics (7K) | A compound brand originating from the United Kingdom with over 30 years of experience in compound supply and service |
| 23 | Pharmeks (666K) | A compound brand originating from Russia, specializing in the synthesis and supply of high-quality new compounds for virtual screening for more than 20 years |
3. Pocket Setting
Docking and Virtual Screening Workflow:
| Pocket setting method | Icon | Mode of operation | Interface display | Note |
| Select Ligand in the Structure as Center | Click the "Select Ligand in the structure as center" button. Define the docking box based on the ligand's region displayed in the 3D Workspace window. The "Cutoff" option sets the box size. | 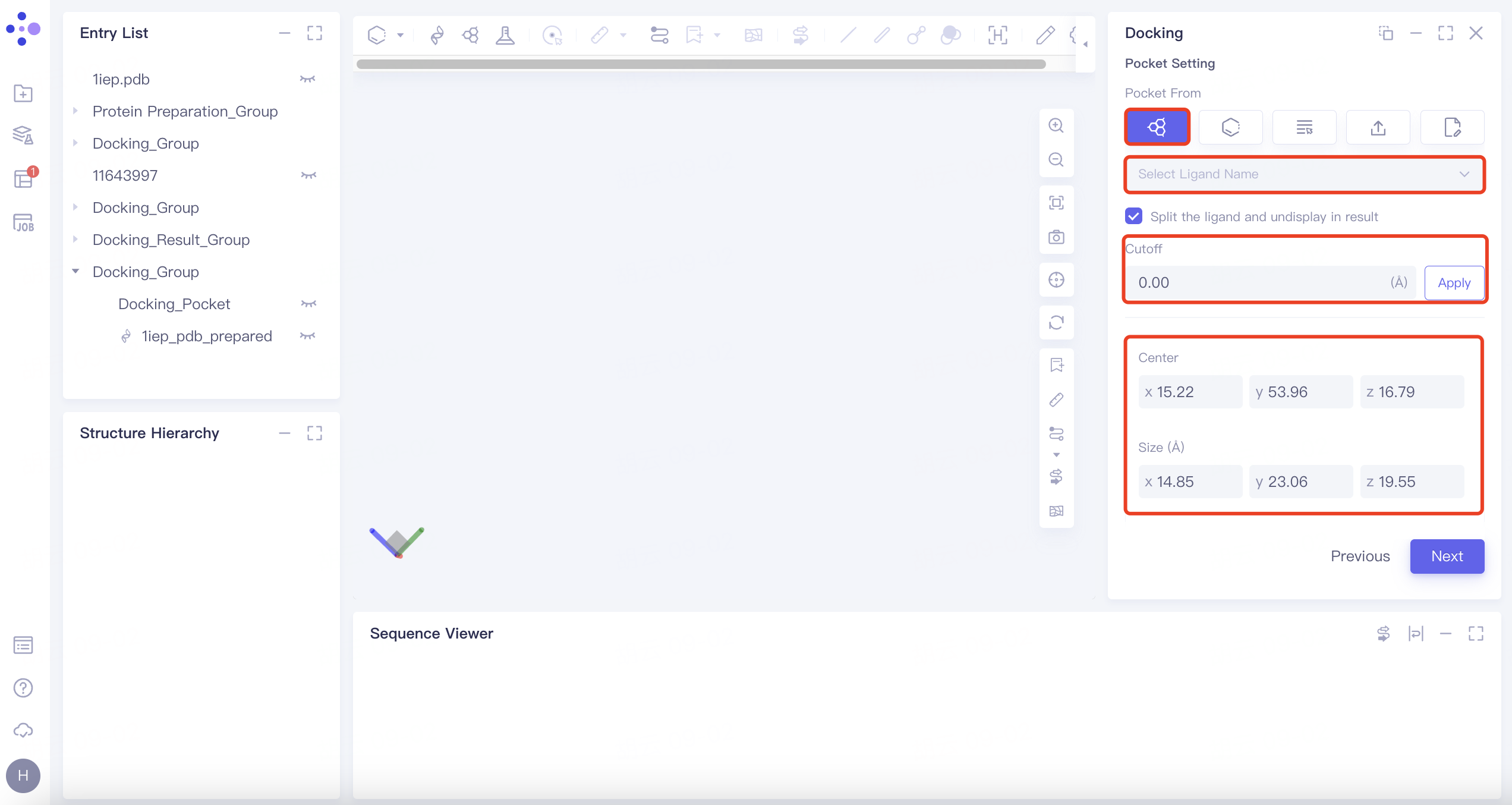 | 1) The box size is adjustable via the cutoff value; a larger value results in a larger box. 2) The option "Split the ligand and undisplay in result" ensures the selected ligand does not enter the docking task and is not displayed in the result. | |
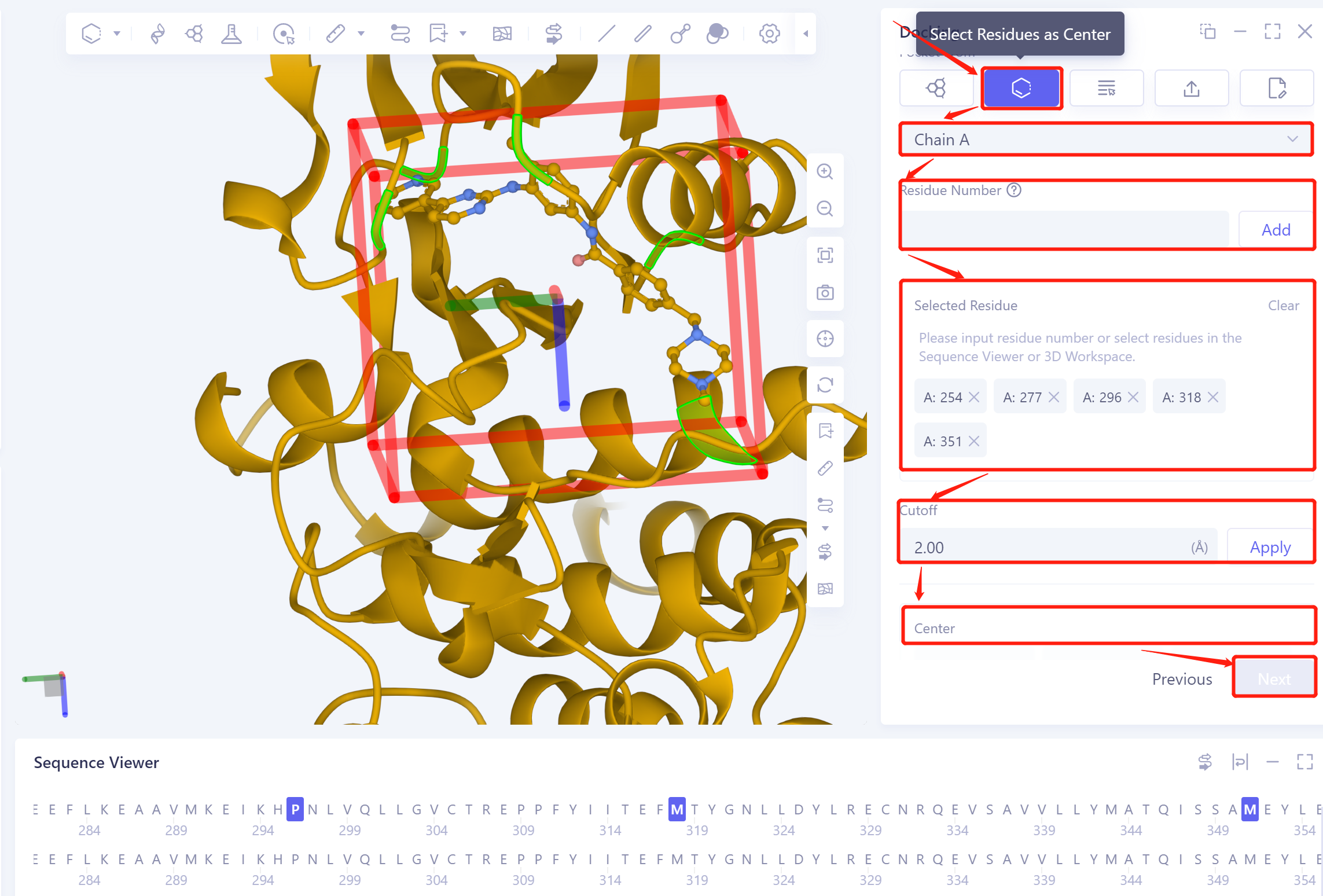
| Select Residues as Center | Click the "Select Residues as Center" button. Choose the protein chain name in the "Select Residue Chain ID" checkbox. Select the amino acid residue using the "Residue Number", 3D Workspace window, or Sequence Viewer window. Set the docking box size with "Cutoff". |
| The residue on the edge of the docking box is used to define the box | |
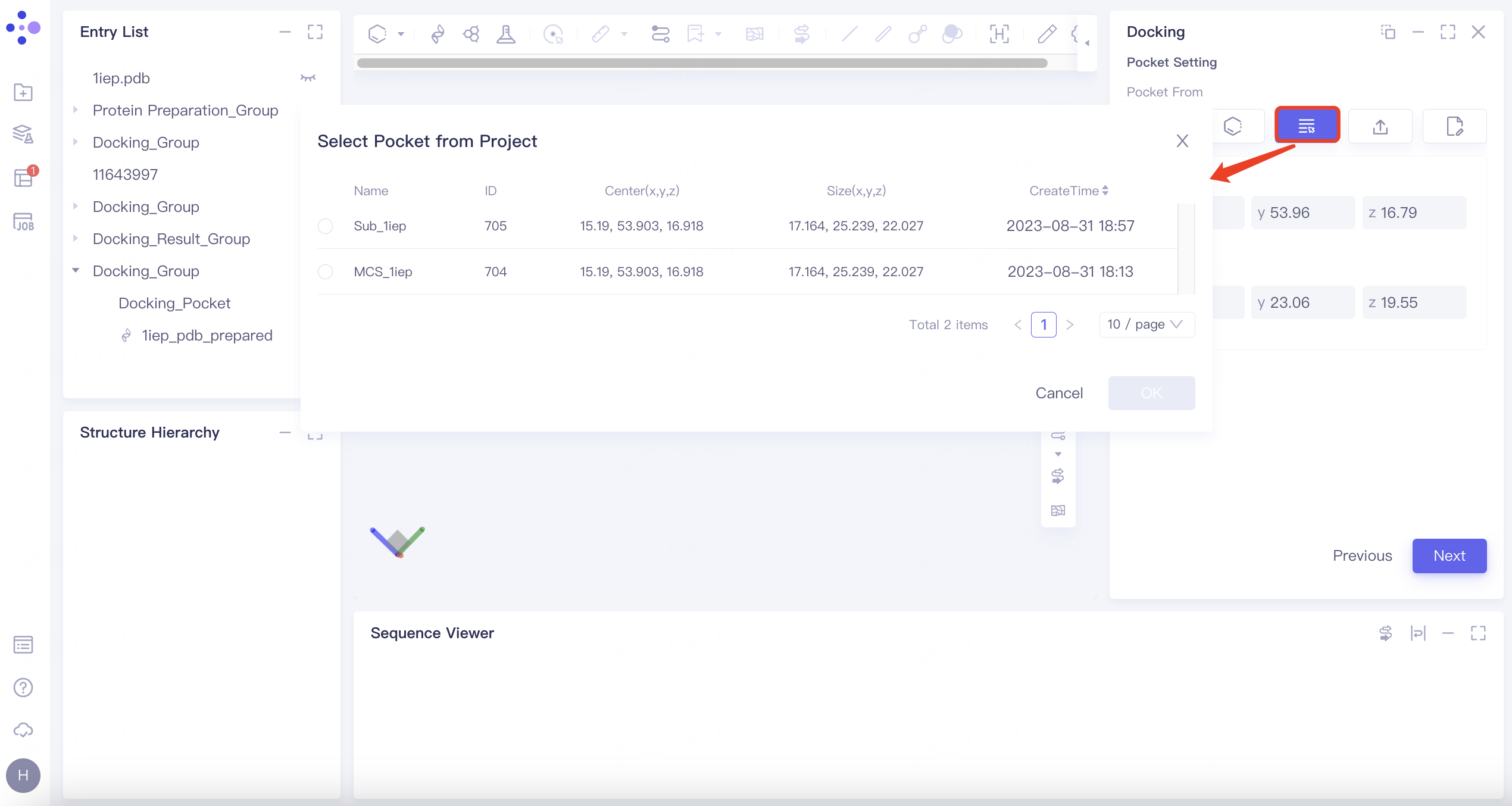
| Select Pocket from Project | Click the "Select Pocket from Project" button. Choose the pocket from a previous task in the "Select Pocket from Project" |
| This method utilizes previously defined pockets from historical tasks | |
| Select from File | Click the "Select File" button. Choose the pocket file from the local file folder |
| Support uploading .txt files | |
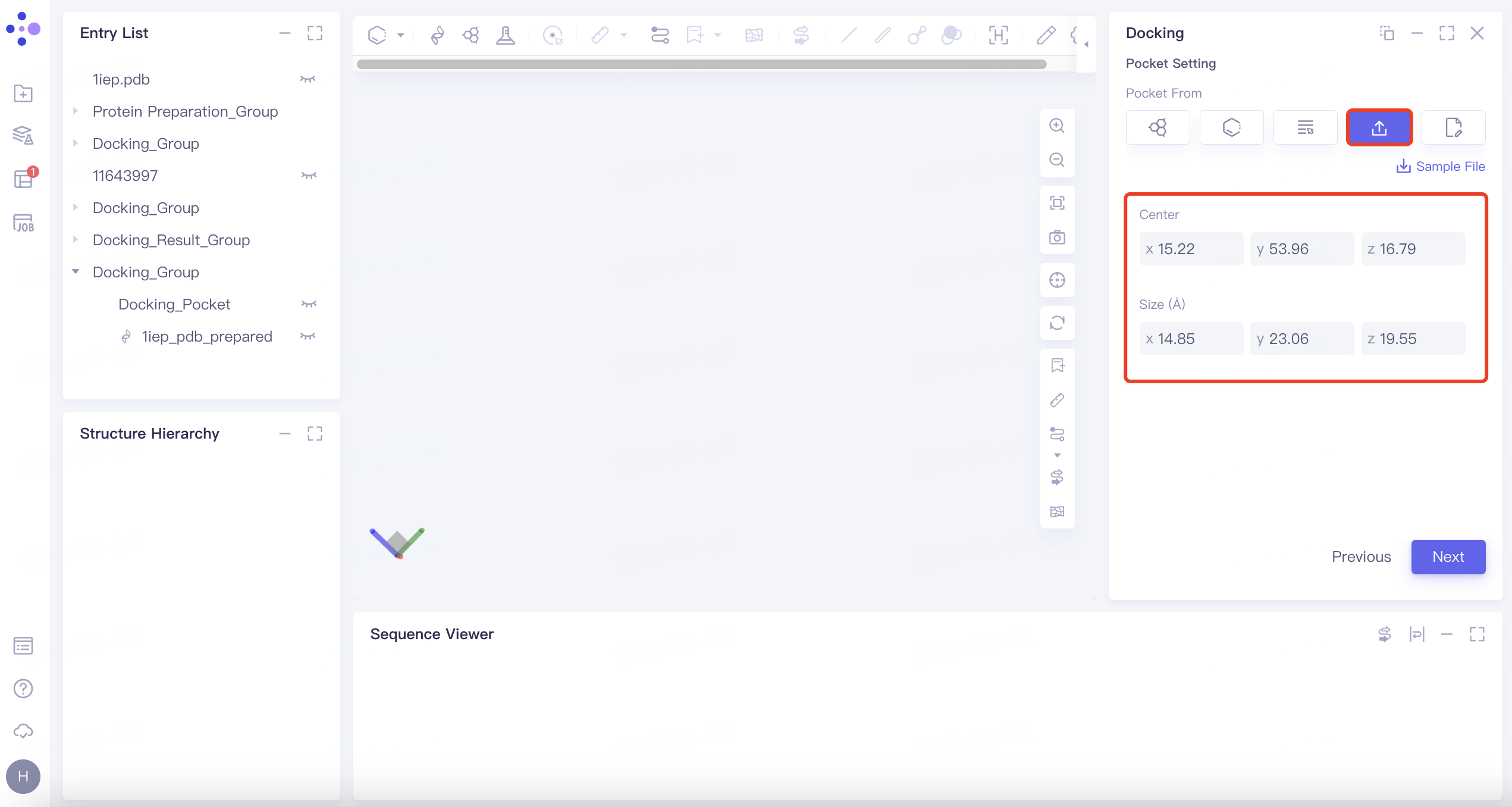
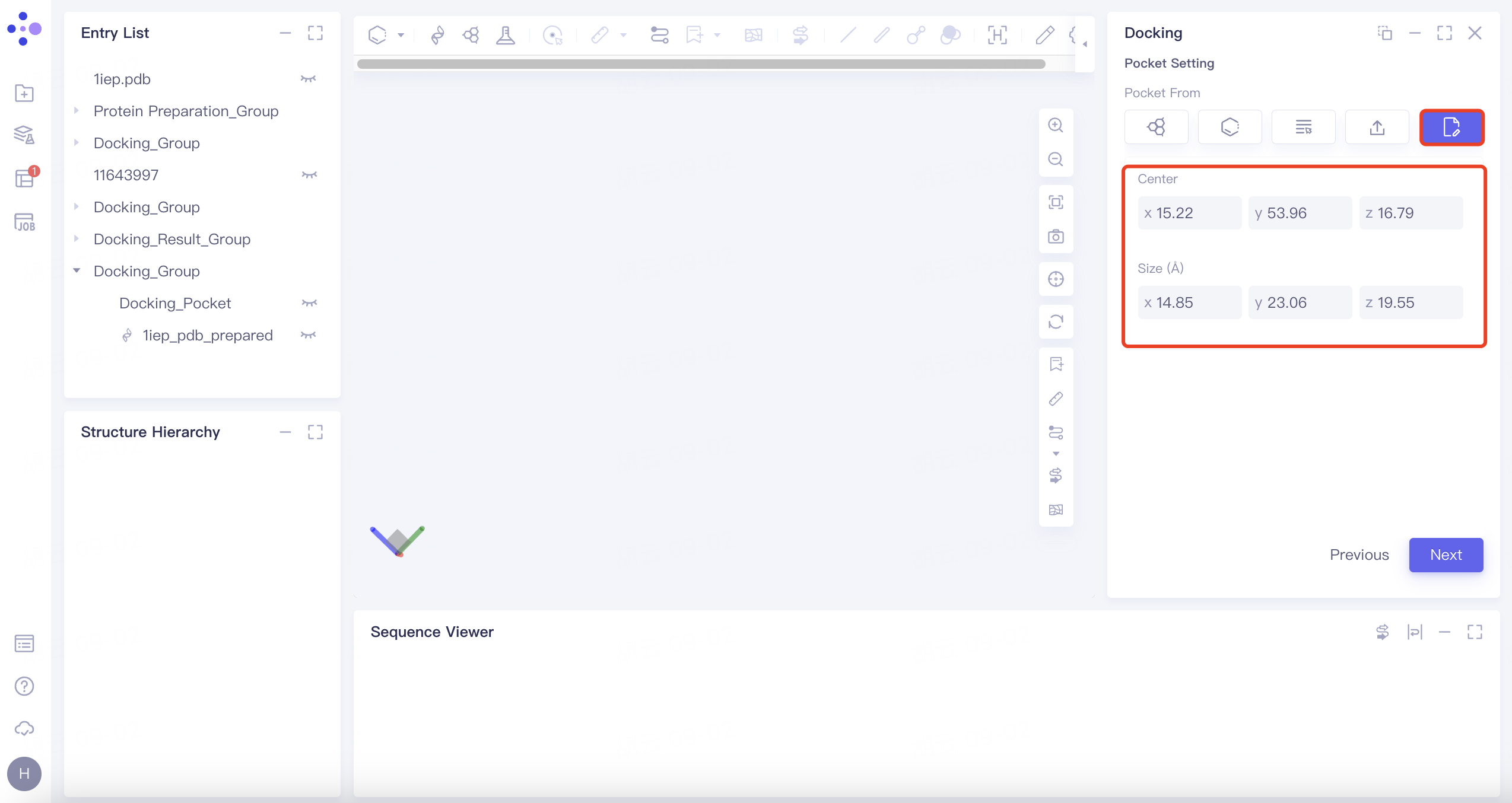
| Input | Click the "Input" button. Manually edit box parameters. |
| This method allows for manual input and customization of the docking box parameters. |