General Operation
1. Register and log in
To use the Hermite ® platform for the first time, you need to register an account. Enter https://hermite.dp.tech in the browser address bar to open the Hermite login page:
Click " Create New Accout " to create a new account. After entering the registered email address, click the " Get Verification Code " button to get the registration verification code. At this time, the system will automatically send the verification code to your registered email.
Note: If you have not received the verification code email within 5 minutes, please check whether the email is recognized as Email Spam, Junk Email by your email system first. If you are sure that you have not received the verification code email, you can contact the official operation staff of Hermite for help.
After entering the registered mobile phone number, click the " Get Verification Code " button to get the registration verification code, and the system will automatically send the verification code to your mobile phone SMS.
Note: If you do not receive the SMS verification code within 5 minutes, you can contact the official operator of Hermite for help.
After entering the verification code and new account password, click the Register button to complete the registration of the new user.
2. Universal menu bar
When you first enter the Herimite ® platform, you will automatically enter the main interface. At the same time, the Hermite ® platform uses the Project method to manage your research work and computing tasks. The main interface includes four graphical windows of the Hermite ® platform and a universal menu bar on the left. The functions of the left menu bar include:
2.1 File
Import Structure
See the "Graphical Functions" section for details.
Get PDB
See the "Graphical Functions" section for details.
2.2 Function
| Structure Modeling | Protein Folding | Predicting protein structure from protein sequence based on Uni-Fold |
| EM Structure Fitting | Flexible construction tool for all-atomic structure model based on cryo-EM density map | |
| Loop Optimization | Conformational optimization of protein flexible loop structures | |
| Virtual Screening | Docking | Molecular docking program based on GPU acceleration, Uni-Mol optimization combined with pose |
| Induced Fit Docking | Flexible docking of proteins and ligands based on induced fit theory | |
| Virtual Screening Workflow | Integrate Protein Preparation, Ligand Preparation, Docking and MM PB/GBSA to complete ultra-high throughput virtual screening in one stop | |
| Binding Stability | 100 ns full atomic simulation of protein-small molecule complexes to evaluate binding stability | |
| Pharmacophore Generation & Screening | Virtual screening tool based on pharmacophore | |
| Binding Affinity Evaluation | MM PB/GBSA Structure | Measuring ligand and target binding affinity |
| FEP Calculation | A tool for evaluating the affinity differences between a series of structurally similar candidate drug molecules (Ligands) and target proteins (Proteins) based on free energy perturbation theory, molecular dynamics simulation, and high-performance computing | |
| FEP Analysis | Results Analysis Tool for FEP Calculation or FEP Recalculation | |
| FEP Protein Mutation | In the process of FEP calculation, the amino acids in the supporting protein undergo Alchemical Transform changes, and the effect of point mutation on protein properties is accurately calculated (currently only the function of predicting the difference in protein thermal stability before and after mutation is available). | |
| Aquasite | Computational analysis of the distribution and stability of water molecules at the target pocket | |
| Biomacromolecule | Antibody Folding | Prediction of antibody structure based on light and heavy chain sequences |
| Sequence Alignment | Protein sequence alignment tool based on multiple sequence alignment algorithm (MSA) | |
| Antibody Numbering | Tool for labeling CDR regions of antibody sequences based on antibody numbering system (supported: IMGT, Kabat, Chothia) | |
| Antibody Humanization | A tool for humanizing antibodies based on databases and statistical models. Supports CDR labeling and transplantation, human antibodies, and provides graphical functions for humanizing antibodies | |
| Antibody Properties | A tool for evaluating the developability of antibodies (including viscosity, aggregation, clearance, isoelectric point, extinction coefficient, etc.) based on databases and statistical models, while supporting structural modeling of antibody variable regions (Fv) | |
| Nanobody Properties | Tools for evaluating the developability of nanobodies (including clustering, clearance, isoelectric points, etc.) based on databases and statistical models | |
| PTM Prediction | A tool based on artificial intelligence algorithms to predict protein post-translational modification ( PTM ) check points (e.g. phosphorylation, glycosylation, ubiquitination, nitrosation, methylation, acetylation, etc.) | |
| Protein-Protein Docking | Protein-protein rigid docking tool based on fast Fourier transform ( FFT ) algorithm | |
| Protein-Protein Interaction | Interactions between proteins | |
| AI Protein Mutation | Based on the artificial intelligence algorithm , the MultiModal Machine Learning features such as protein structure and dynamics information are fused to quickly predict the difference in thermal stability before and after protein mutation | |
| Molecule Recommendation | VD-Gen | Based on Virtual Dynamics artificial intelligence algorithm , molecular de novo design function relying on protein pocket |
| Similar Molecules Search | Generate molecular fingerprints of query molecules based on various means, matching their similarity to target molecules | |
| Substructure Search | SMARTS based on substructure matches the same part of the target molecule | |
| Maximum Common Substrucrture | Find the largest common substructure in a batch of molecules | |
| Compound Clustering | Clustering molecules with similar properties into a class based on molecular fingerprints | |
| Uni-QSAR | A quantitative structure-activity relationship ( QSAR ) modeling and prediction tool based on multiple models of artificial intelligence (Deep learning, Machine Learning ), and a new 3D molecular descriptor based on molecular 3D structure pre-training model (Uni-Mol) is added on the basis of molecular 1D and 2D descriptors, greatly improving the accuracy of QSAR models | |
| Uni-QSAR Analysis | Uni-QSAR model performance comparison and analysis tool | |
| General | Protein Preparation | The protein is processed for subsequent tasks |
| Ligand Preparation | Processing the ligand for subsequent tasks | |
| Protein Alignment | A tool for 3D superposition of protein structures (according to: backbone, Ca, sidechain), and supports the superposition operation of multiple protein structures |
2.3 Windows
Evokes the Hermite platform resident window (any of the four graphical components).
Arouse the function you have already activated (i.e., the function you minimized during use is restarted from here).
Restore the default layout of the page.
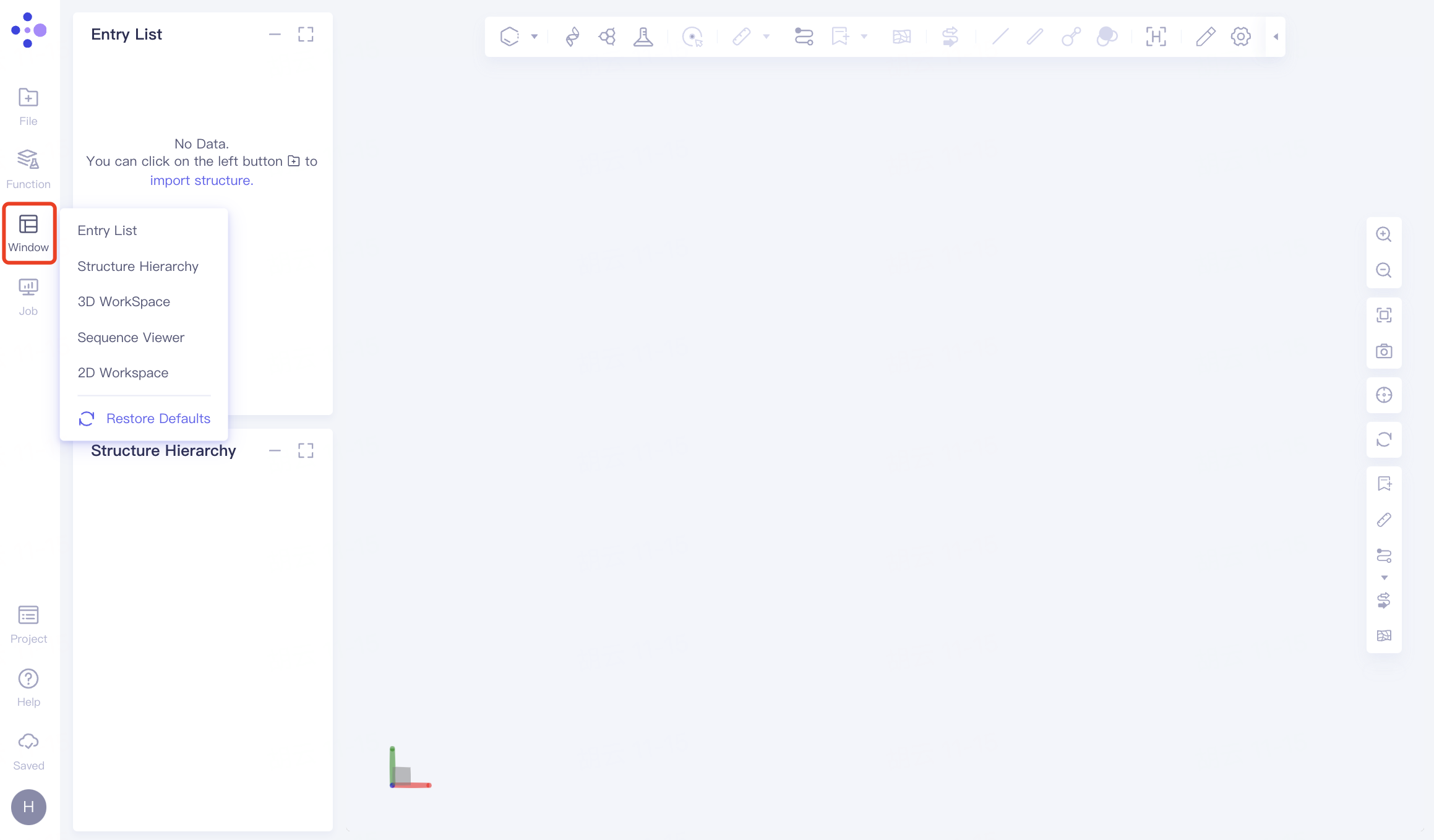
2.4 Job List
2.4.1 entrance
When you are not currently in a project, click on the left common menu bar "Job" to expand the "Job List" interface, where you can view all historical jobs.
When you are currently in a project, click on the left common menu bar "Job" to expand the "Job List" interface, you can view all the jobs under that project.
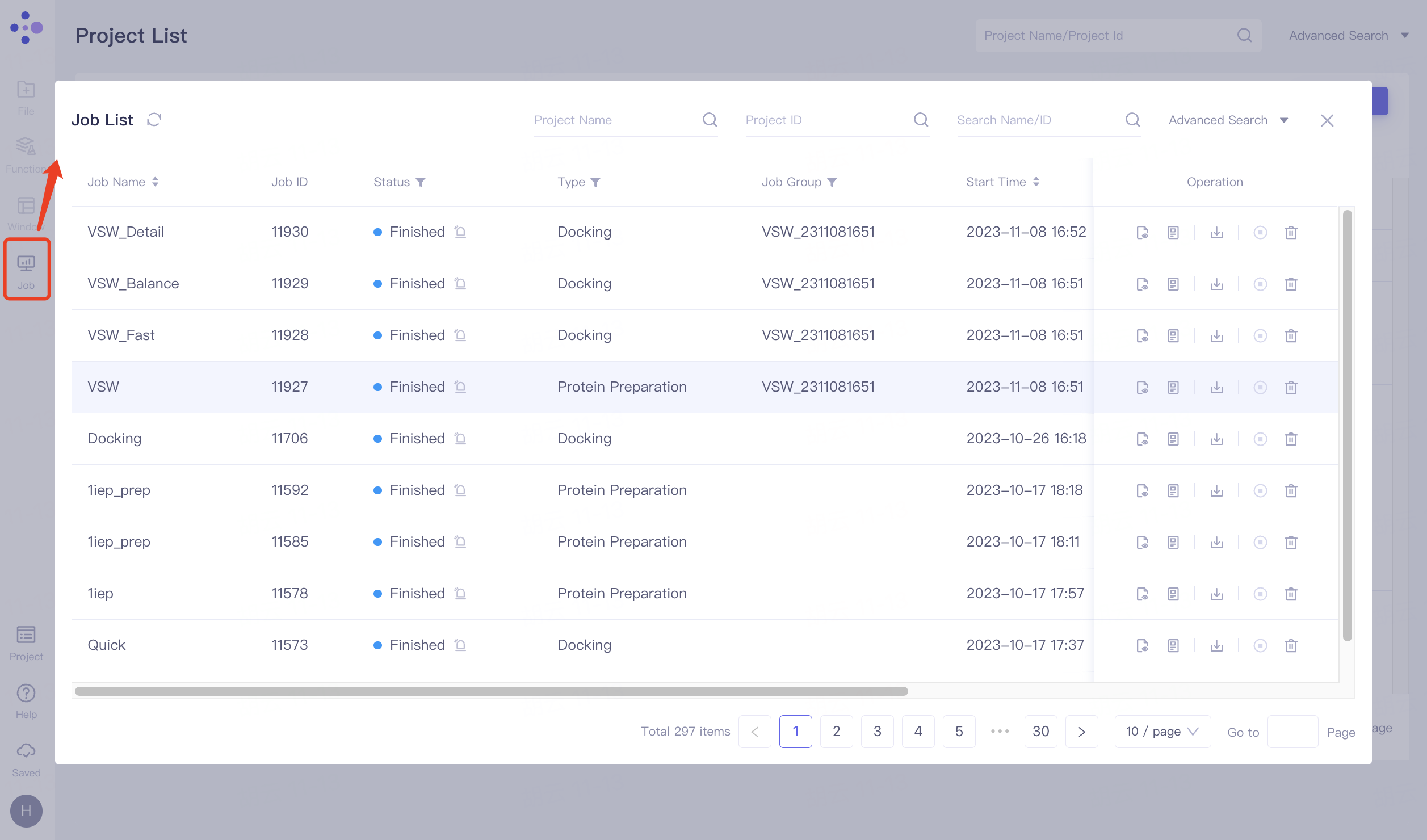
2.4.2 operation
The default interface supports searching by Project Name, Project ID, and Name/ID conditions.
Click "Advanced Search" to expand advanced search. Support searching by Job Name, Job ID, Job Status, Operator, Project Name, Project ID, Start Time and End Time. Click "Search" to search, and click "Reset" to reset the search term.
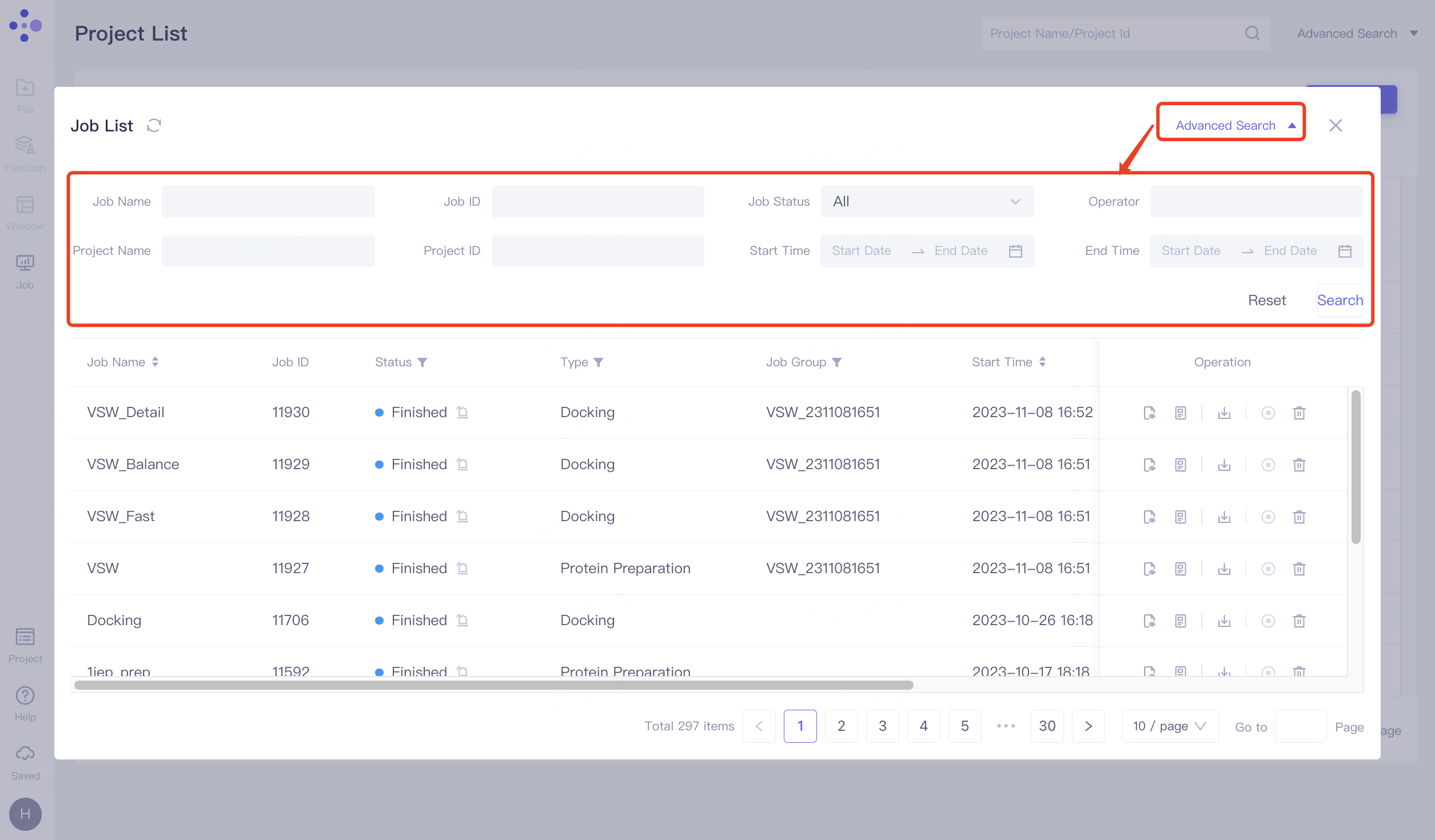
Support sorting by Job Name, Job Status, Start Time, and End Time.
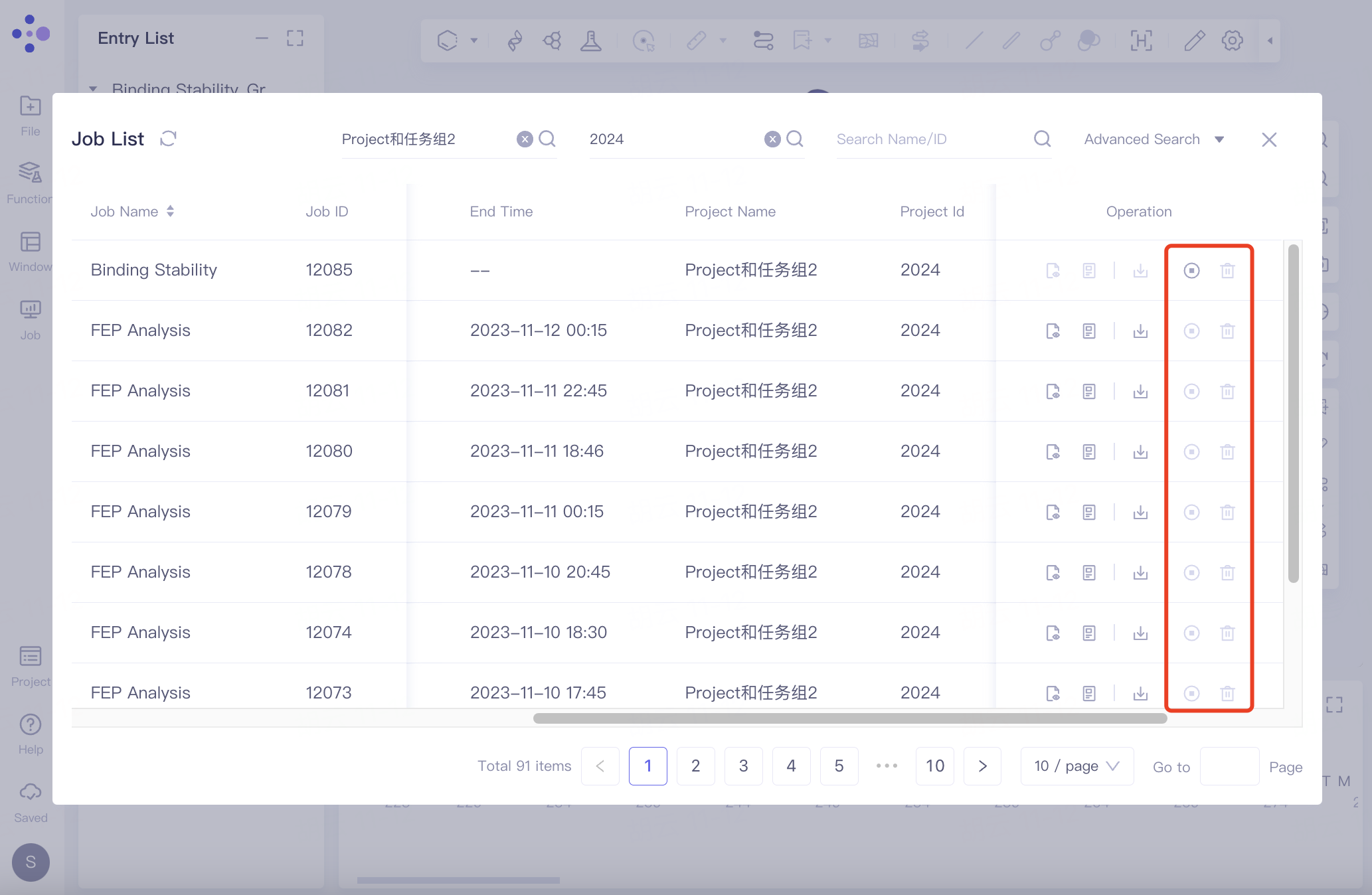
Operations under:
Show: Display the results of this job.
Log & Paramas: Log and parameters, when your task calculation fails and you need to seek help from Hermite official operators, the information provided together, so that we can quickly help you locate the problem;
Download: Download the relevant files for this job.
Cancel: Cancel the task. Canceling the task does not mean suspending it. Once the task is canceled, if you need to run it again, you need to recalculate it in Functions.
Delete: Delete the task.
There are differences in the operations supported by administrators and members on the Job List, as follows:
Administrators and members can both "Cancel" and "Delete" their projects.
Administrators can perform "Cancel" and "Delete" operations on members' projects.
Members cannot perform the "Cancel" and "Delete" operations on the administrator's project.
2.5 Project
2.5.1 New Project
Create an independent Project for the new research computing system. The specific steps are: General toolbar Menu → Project → New Project → Name the new Project → Click Apply → Automatically jump to the new page (4 graphical windows open by default).
When a member is in the invited organization, the new project will be automatically synchronized with the administrator.
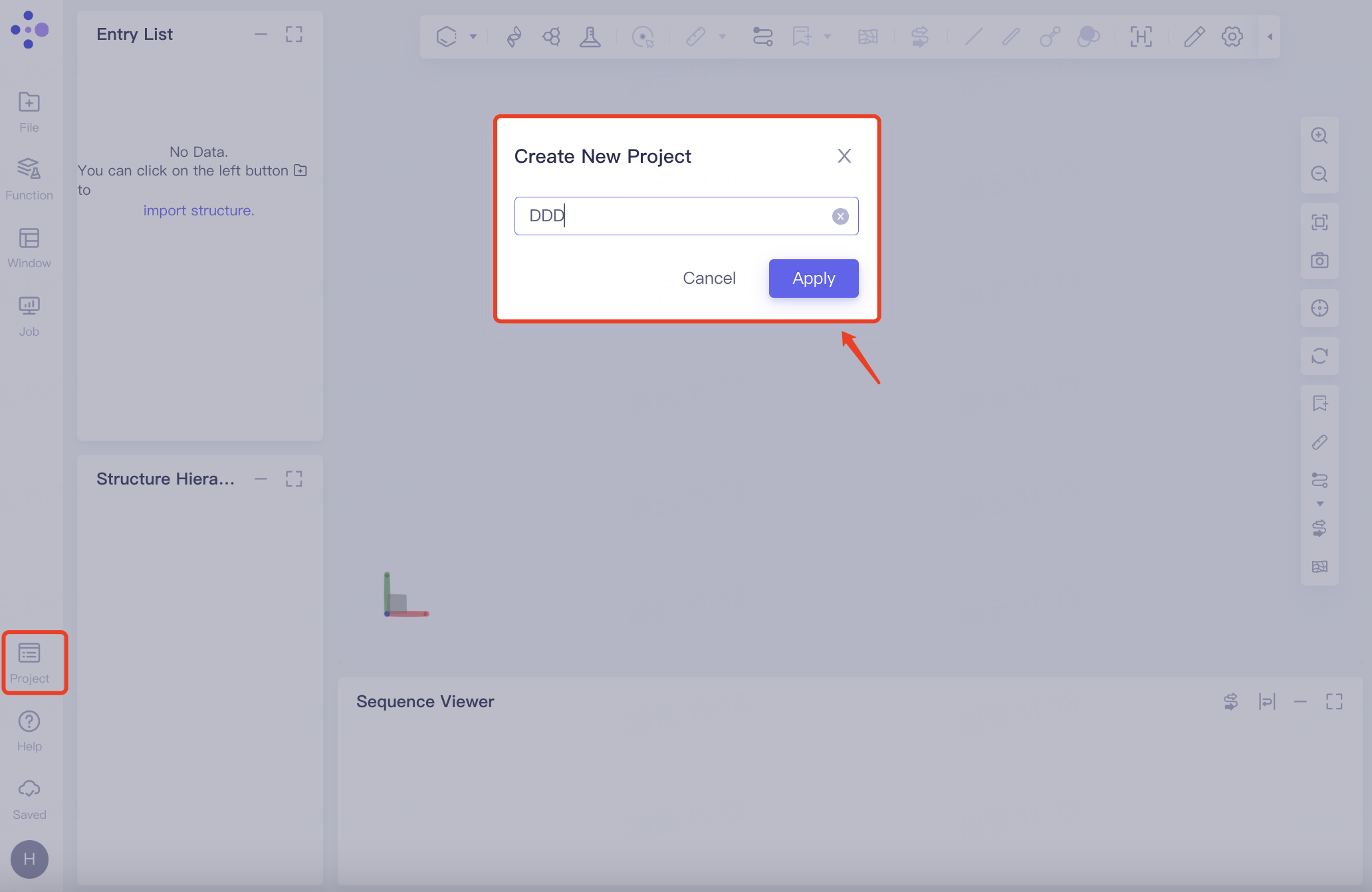 | 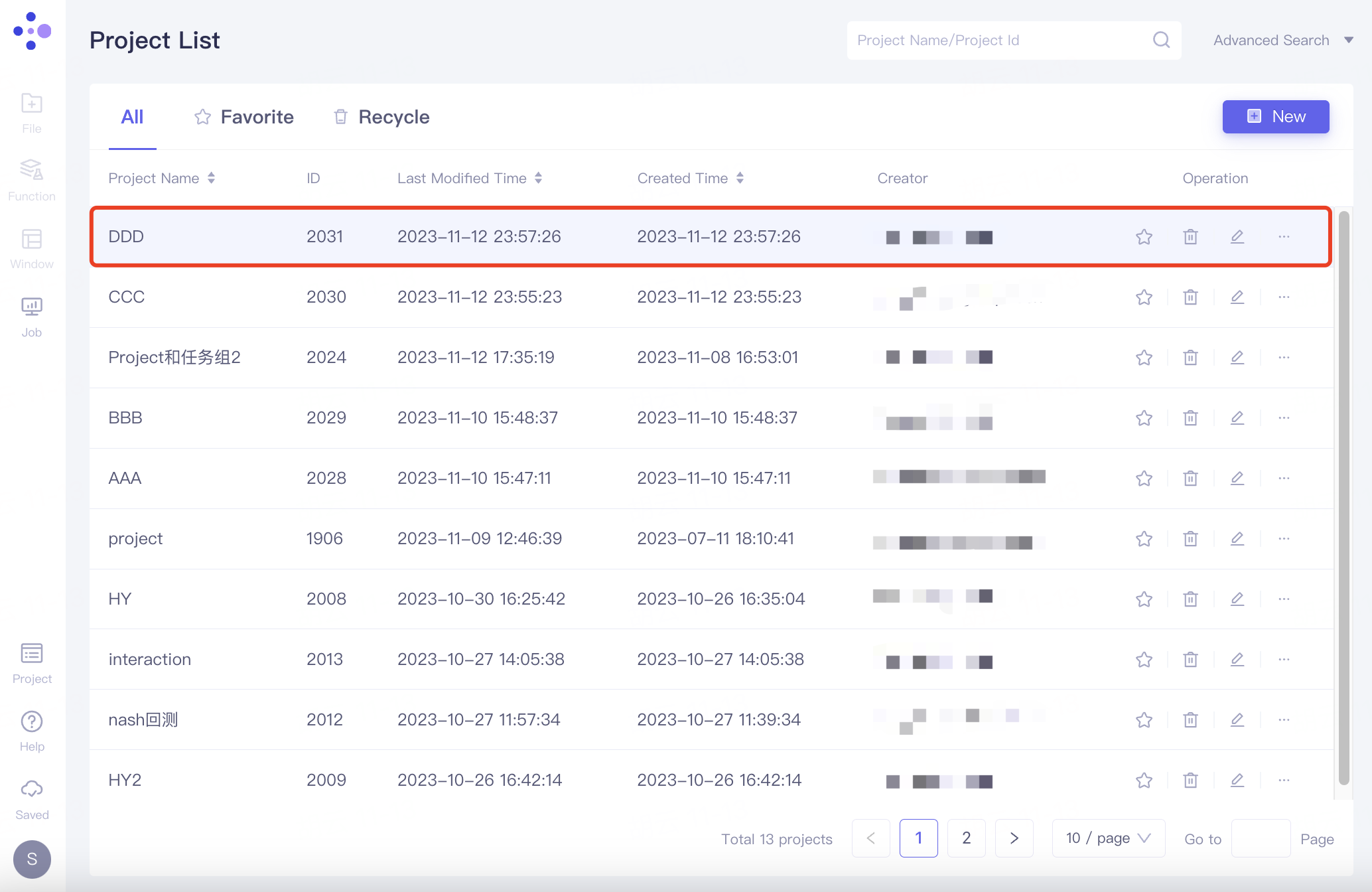 |
2.5.2 Save Project
Manually save the display status of the graphical interface under the current Project and the parameters of various activation functions. Specific steps: General toolbar Menu → Project → Save Project; or shortcut method: Ctrl + S (Win system)/Command + S (OSX system).
Note: After you perform important operations, we hope you can save your data; at the same time, in order to maximize your convenience, Hermite, as a SaaS Cloud Computing Platform, will regularly automatically save your project data in the local browser to your cloud account.
2.5.3 Project List
The Project List provides comprehensive management of your historical projects on the Hermite platform, allowing you to enter the project by clicking on the project name under the Project Name column.
Support searching and sorting by Project Name, Last Modified Time, and Create Time.
Operations under:
Add to Favorite: Favorite this project. The favorited projects can be viewed in the Favorite category.
Delete: Delete this item. Deleted items can be viewed in the Recycle category.
Rename: Rename the project.
Member: Add organization members.
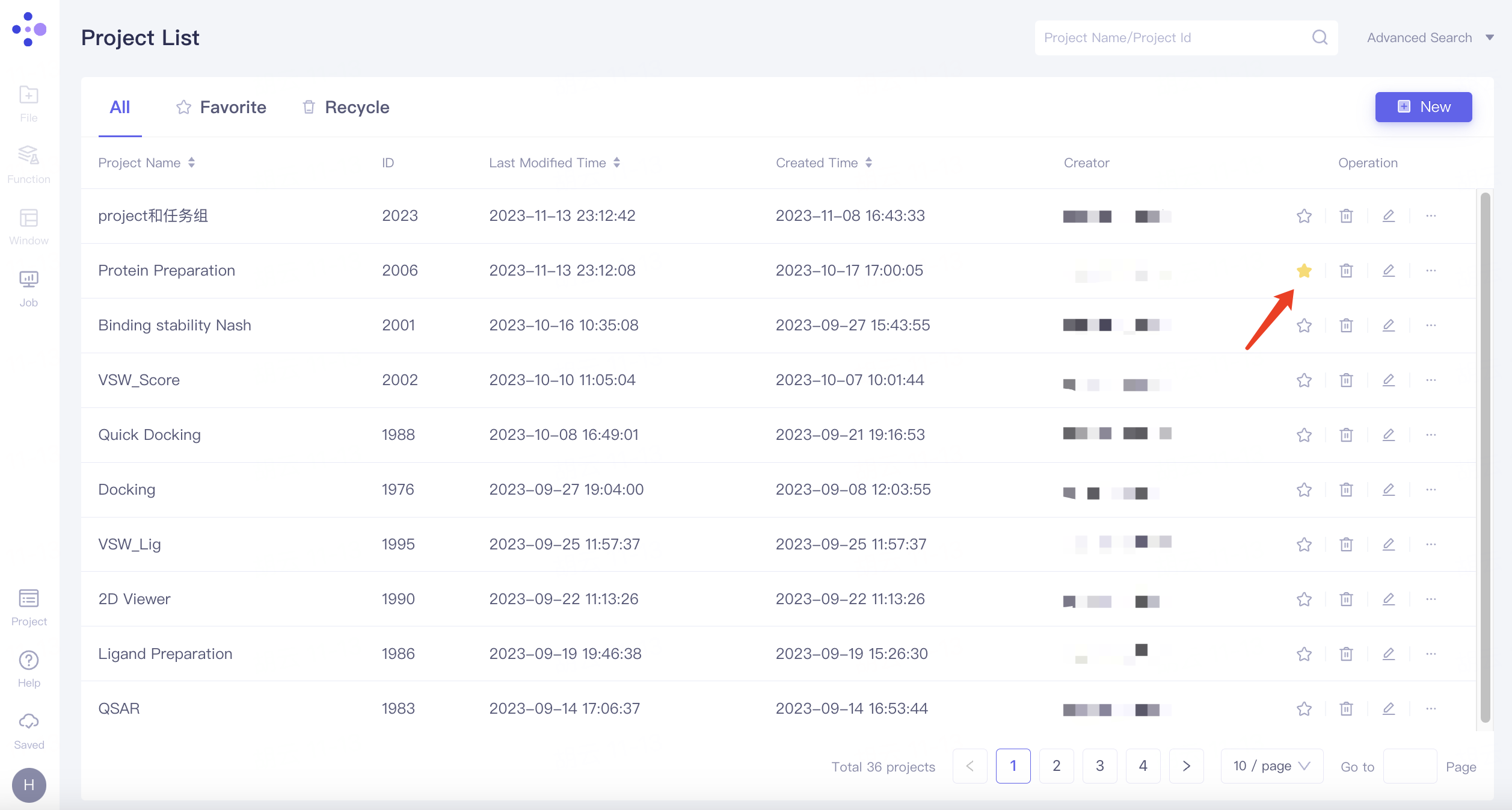
Add members to Project
When a project requires multiple participants, the following methods can be used to achieve project sharing among multiple people. The specific operations are as follows:
Administrators share projects
Find the project you want to share in the expanded project list, click "..." under Operation on the right, and click "Member" in the expanded options.
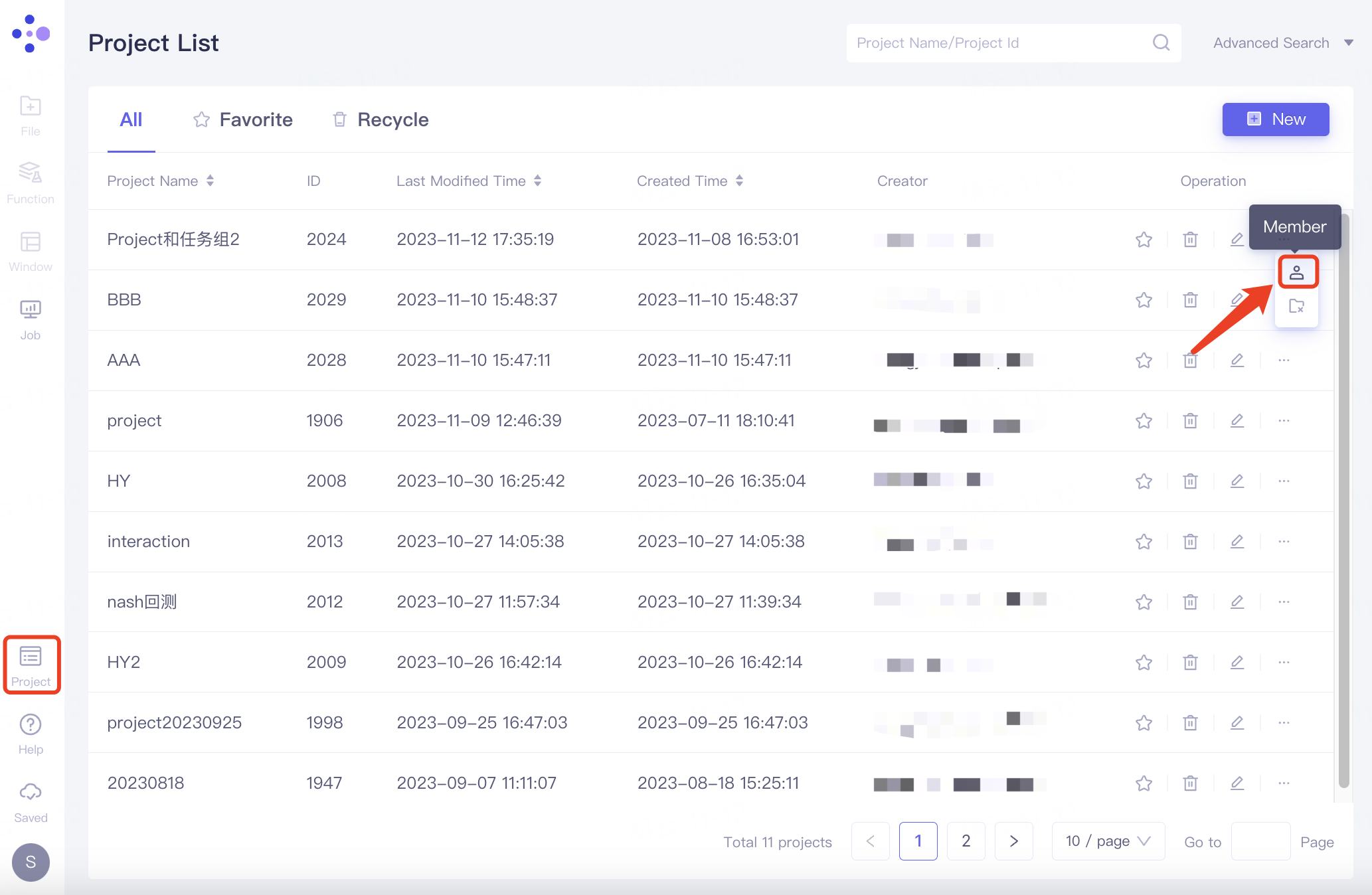
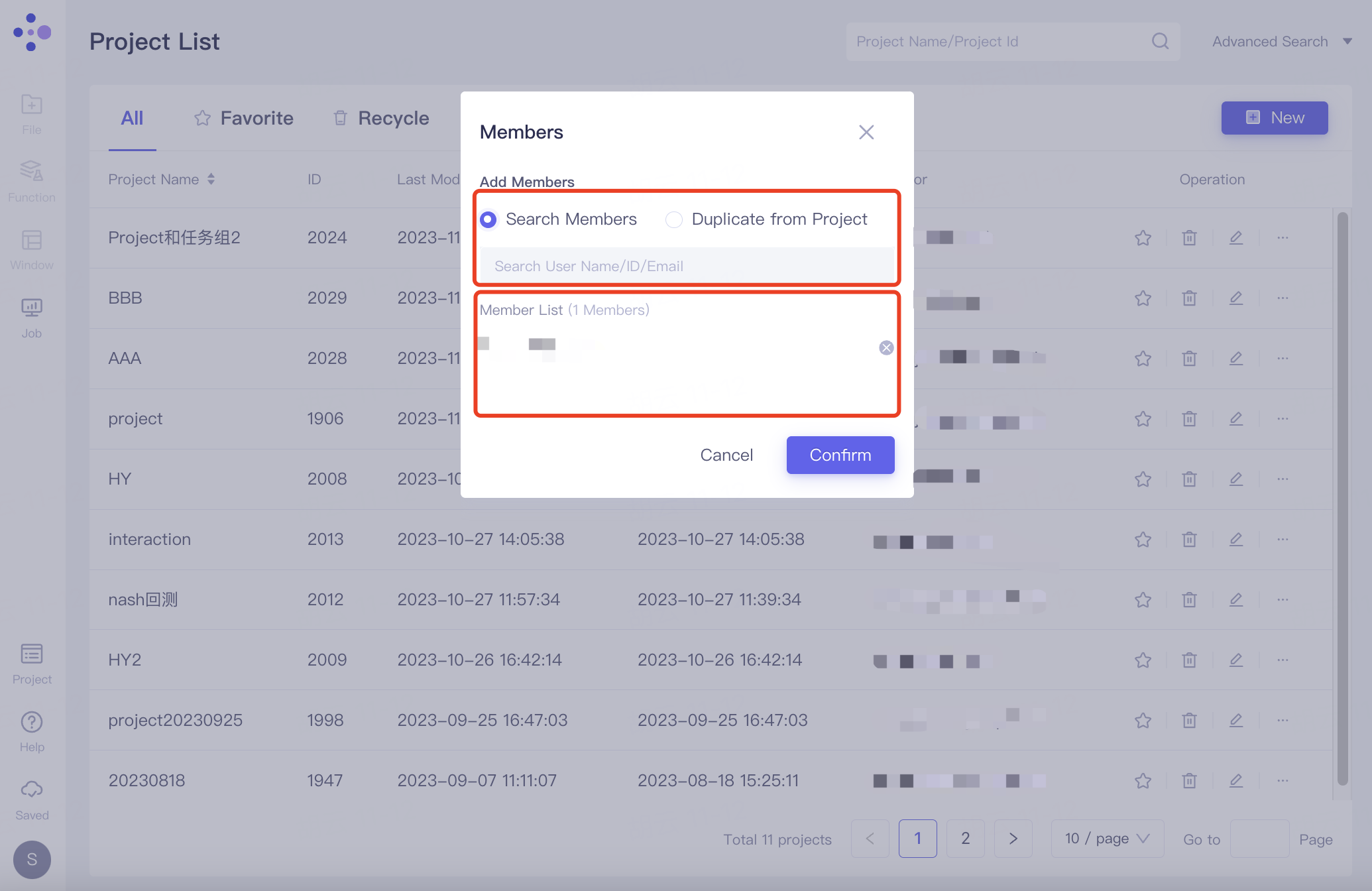
The "Members" interface pops up. After adding members, the member email is displayed under the Member List. Click "Confirm" to confirm the addition.
Members can be added in two ways:
Search Members: Invite users to enter the current project by searching their username/ID/email.
Duplicate from Project: Invite users into the current project based on their project name/ID.
Note: Only members already in the organization can be added. For details on adding members to the organization, see User Center.
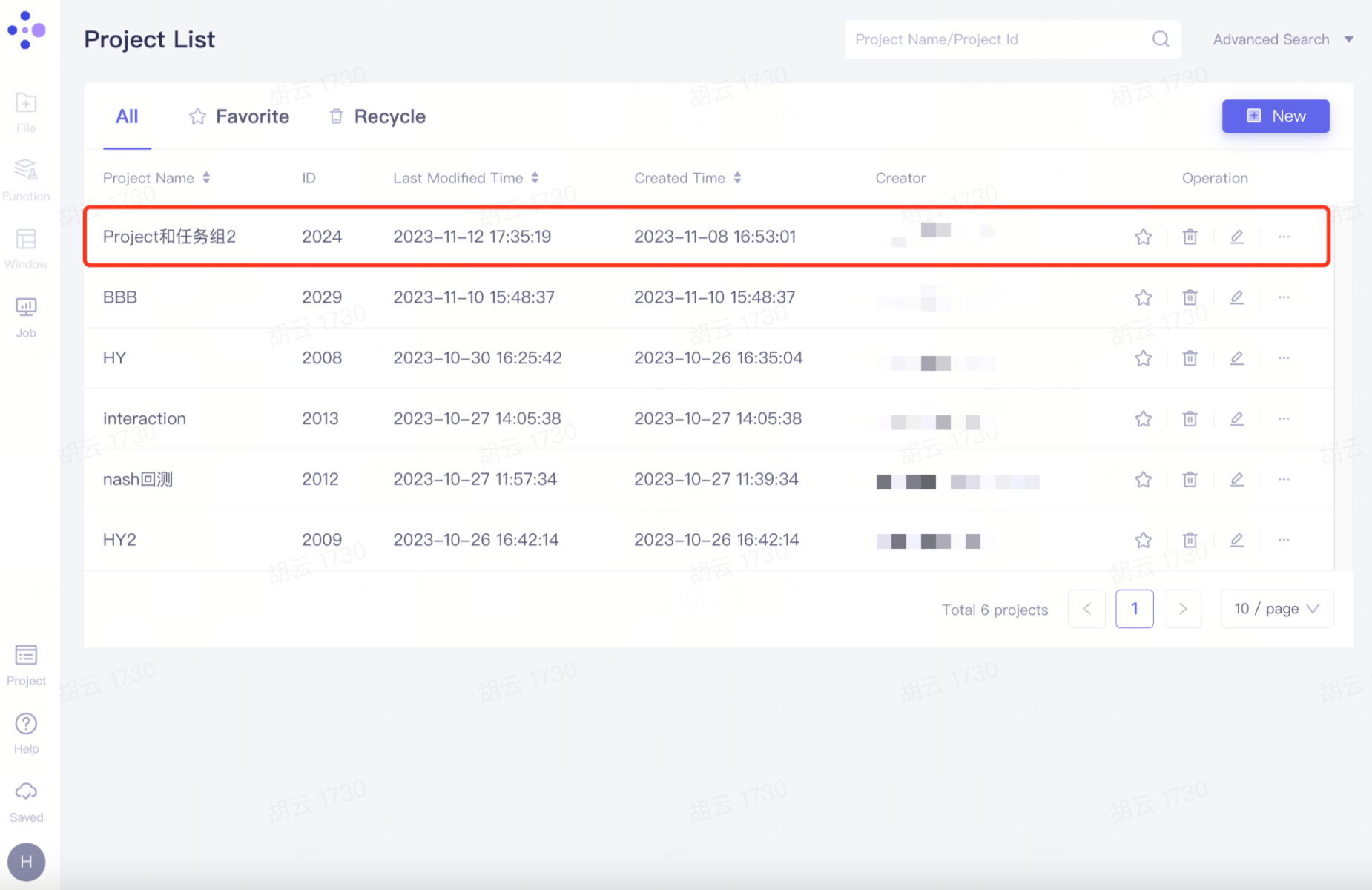
Members synchronize projects
The project appears synchronously in the member account
2.6 Help Center
Help document, including homepage, dynamics and announcements, software cases, operation manuals, get more help.
2.7 Saving to the Cloud
Save the current task to cloud space.
2.8 Account Center
2.8.1 Organization
When you have not joined any organization, you are the administrator of an independent organization by default; After you join an organization, you can switch organizations as follows.
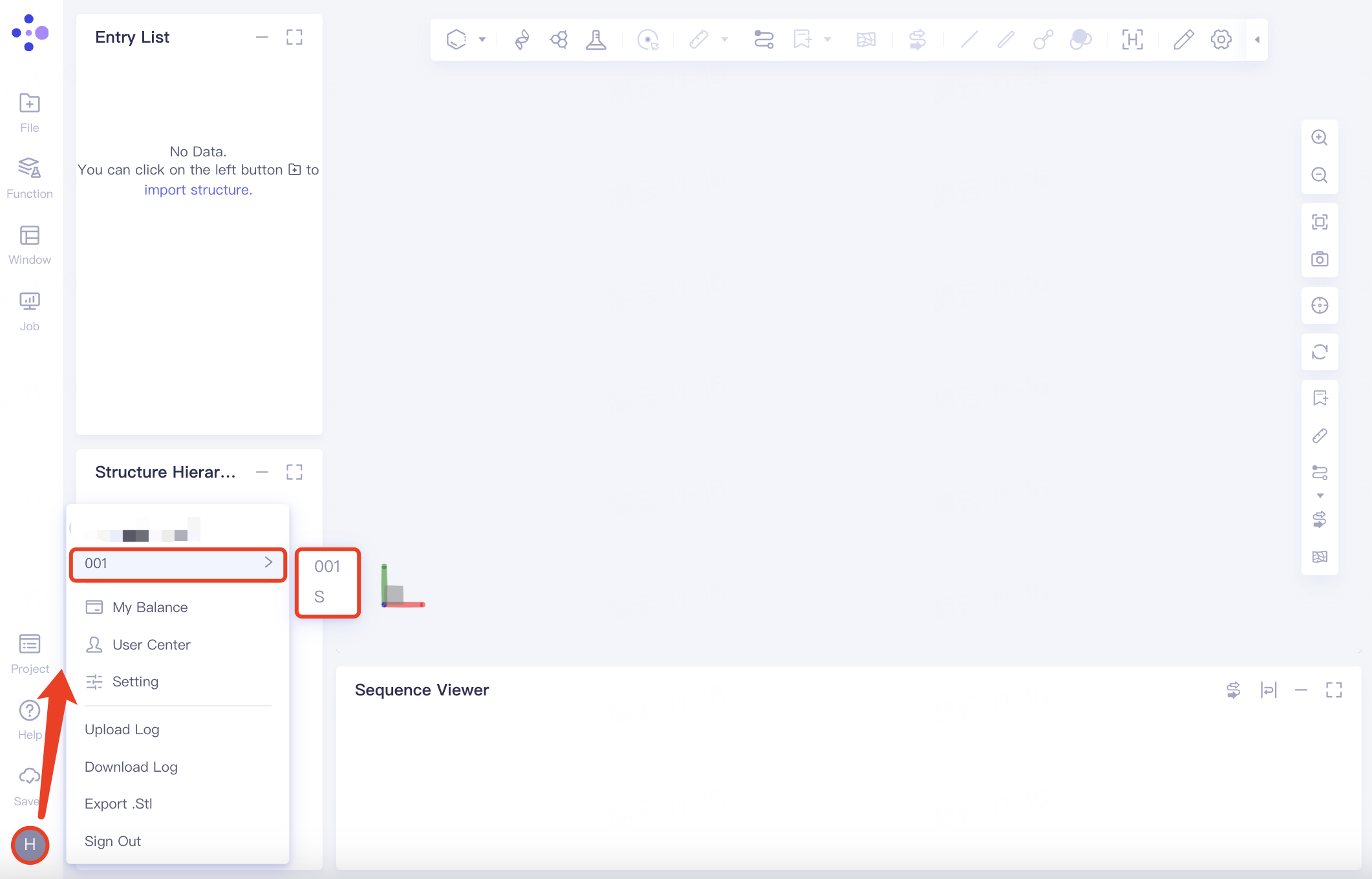
2.8.2 My Balance
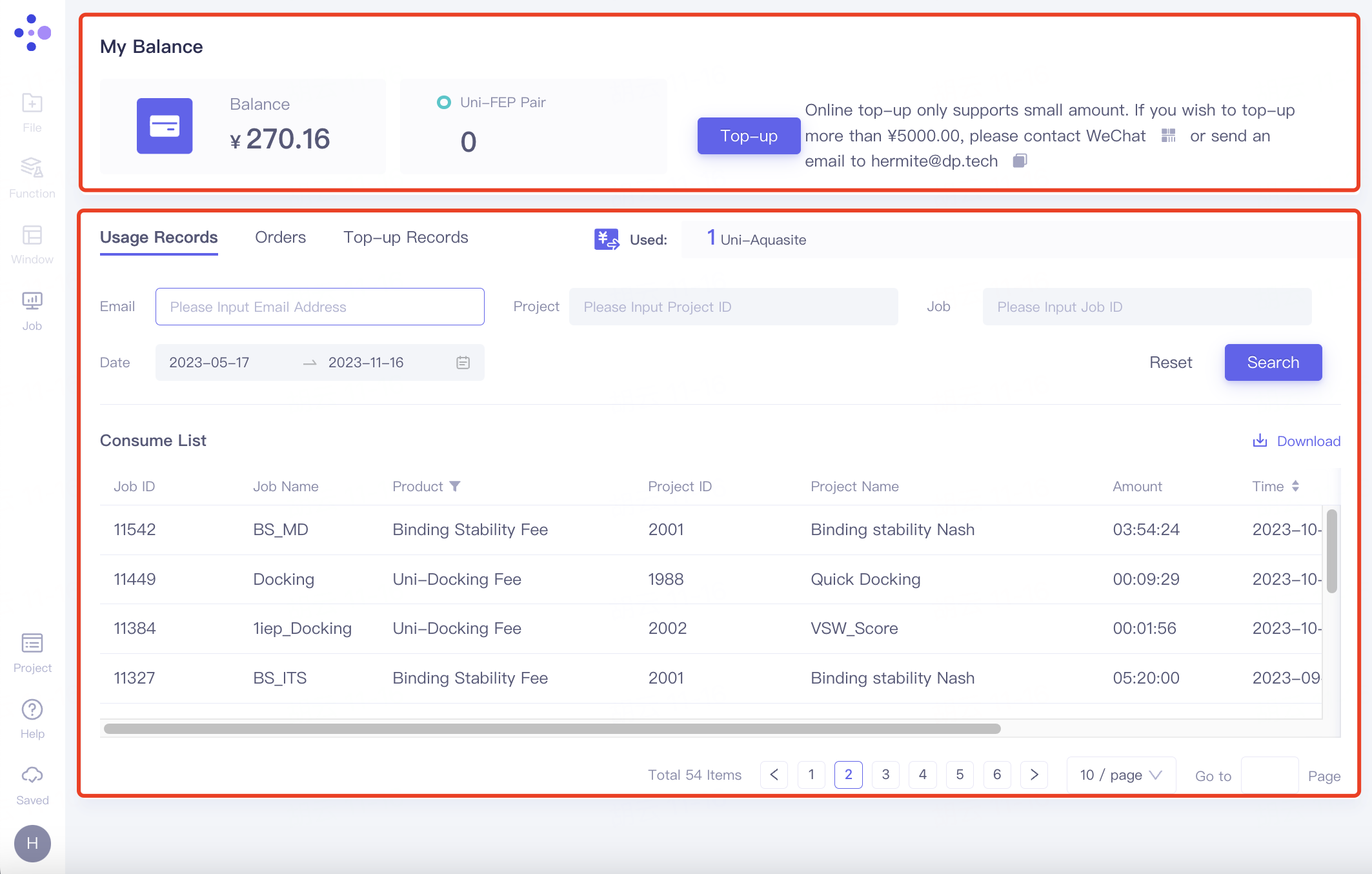
At the top red box, Blance shows the balance, and Uni-FEP Pair shows the remaining number of FEP Pair. Click "Top-up" to make a small recharge of less than 5000 yuan.
At the red box below:
Usage Records: Records the use of effect packages;
Orders: Recorded the amount of effect package usage.
Top-up Records: Recharge records.
2.8.3 User Center
In the user center interface, the top is personal information, and the bottom User List records the information of other members
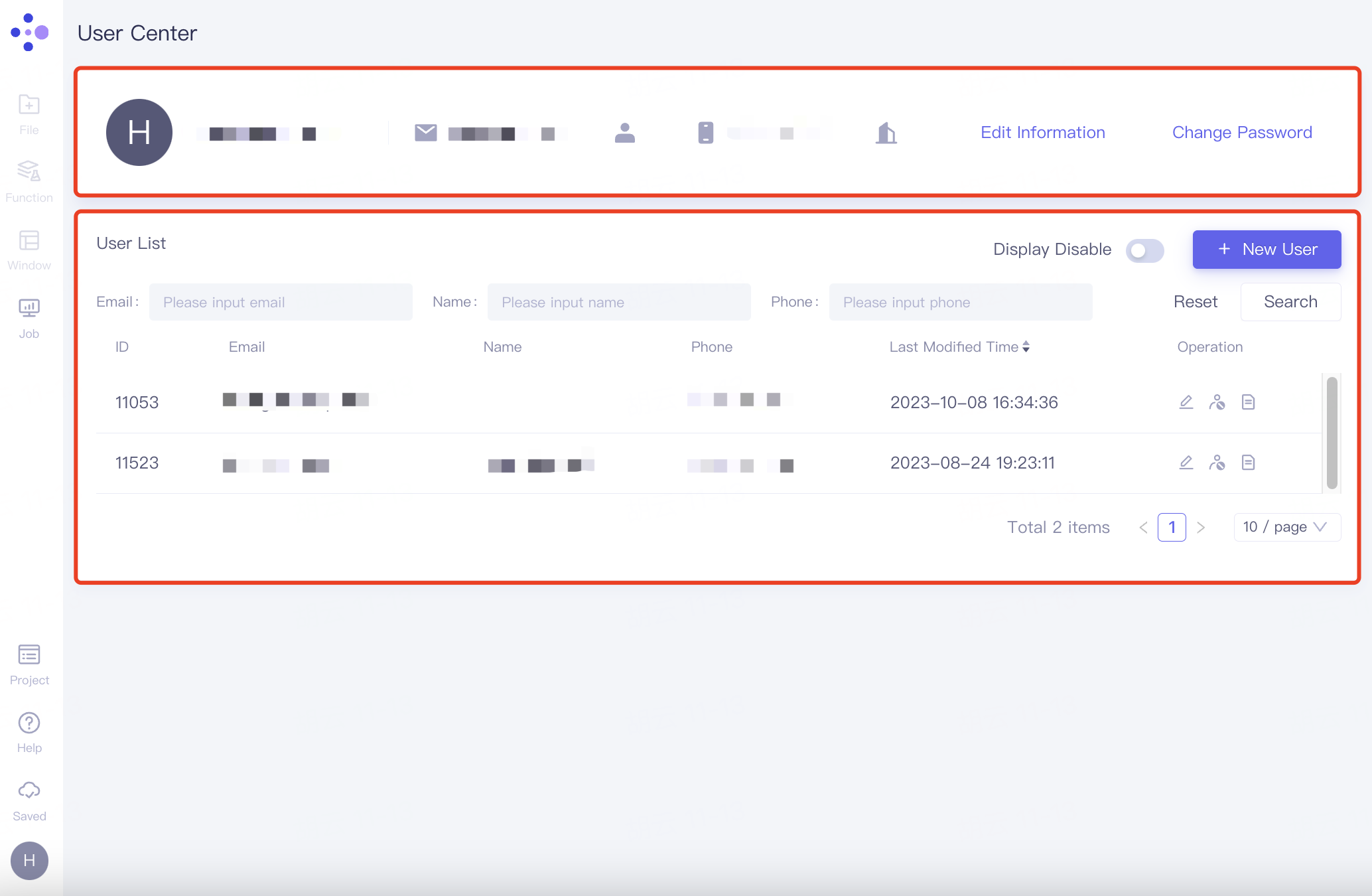
2.8.3.1 Personal information
Edit personal/organizational information
In the personal information field, click "Edit Information" on the right to expand the "Edit Personal Information" window. Fill in the user name, phone number, verification code, organization name and other information below, and click "Submit".
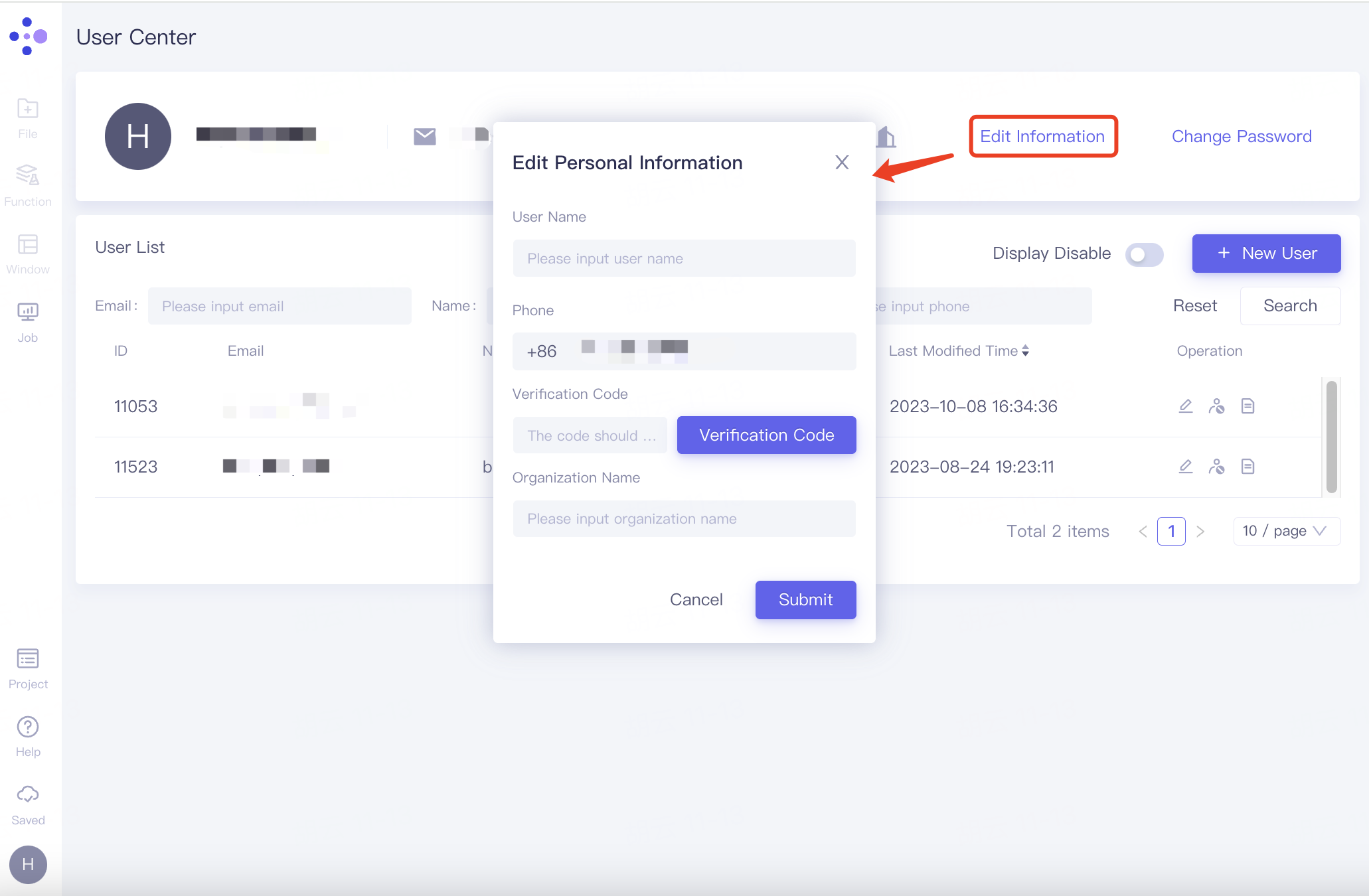
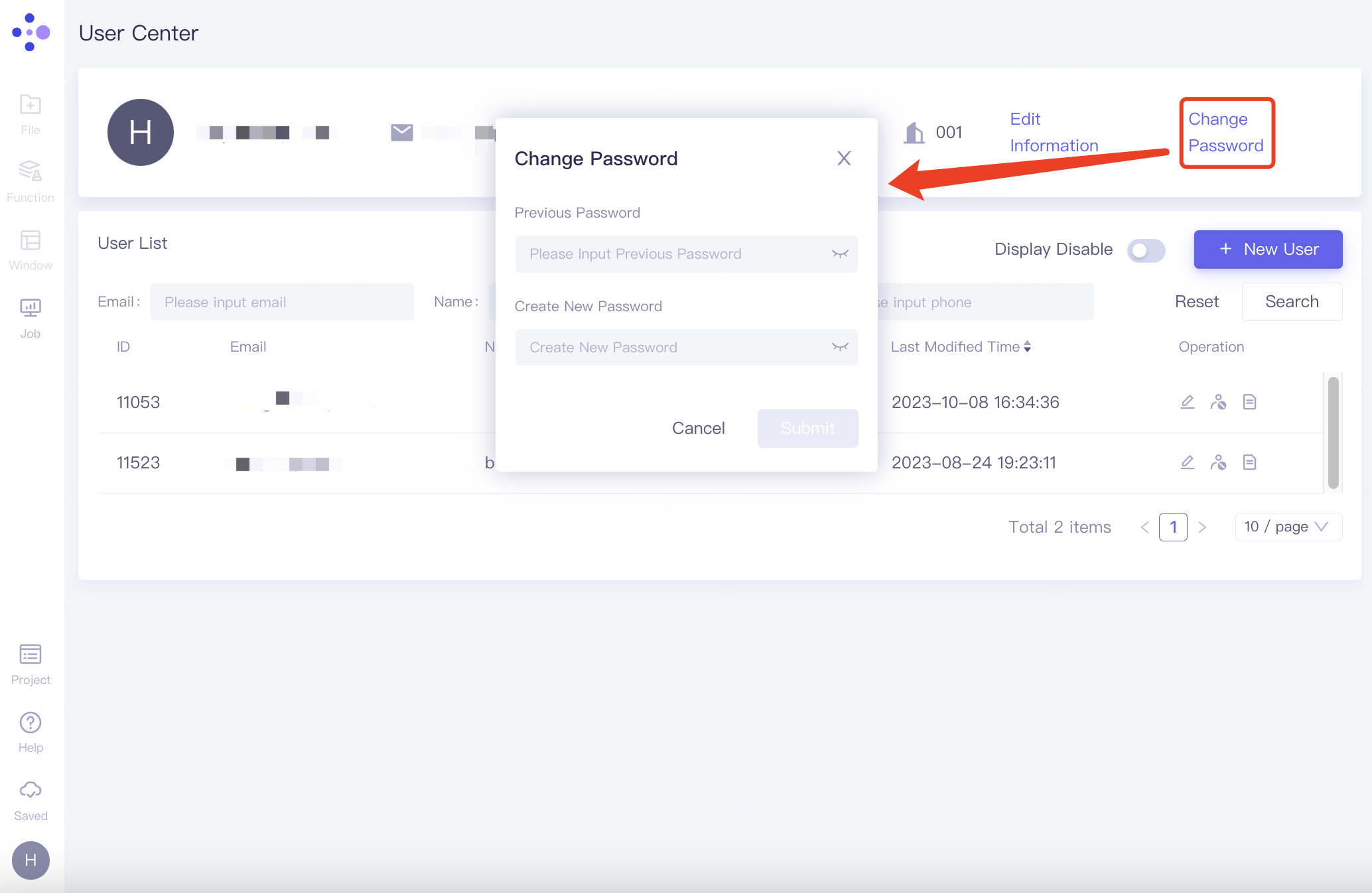
Change password
In the personal information field, click "Change Password" on the right, and the "Change Password" window will pop up. Fill in the original password and the modified password, and click "Submit".
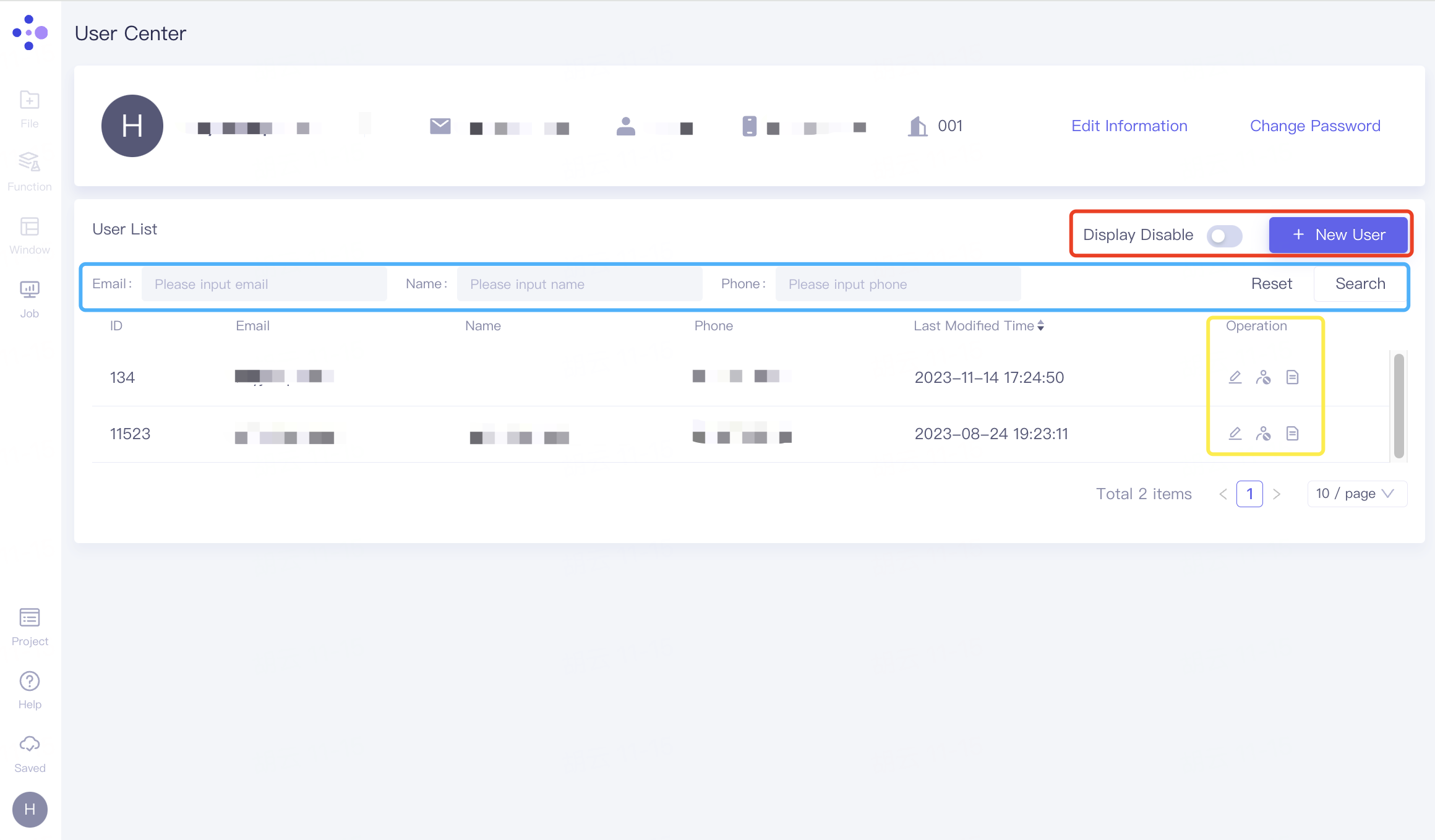
2.8.3.2 User List
Basic operation
Red box:
Display Disable: The button can adjust the display and concealment of the disabled user.
+New User: Add members to the current organization;
Blue box: According to information such as Email, Name, Phone, etc., click "Search" to search for the desired user, and click "Reset" to reset.
Yellow frame:
Edit: Adjust the Projects this member participates in;
Disabled User: Remove members from the organization.
Job Record: View members' task records.
Organizational management
- Administrators invite into the organization
Click Account, select "User Center" in the expanded list, and enter the User Center.
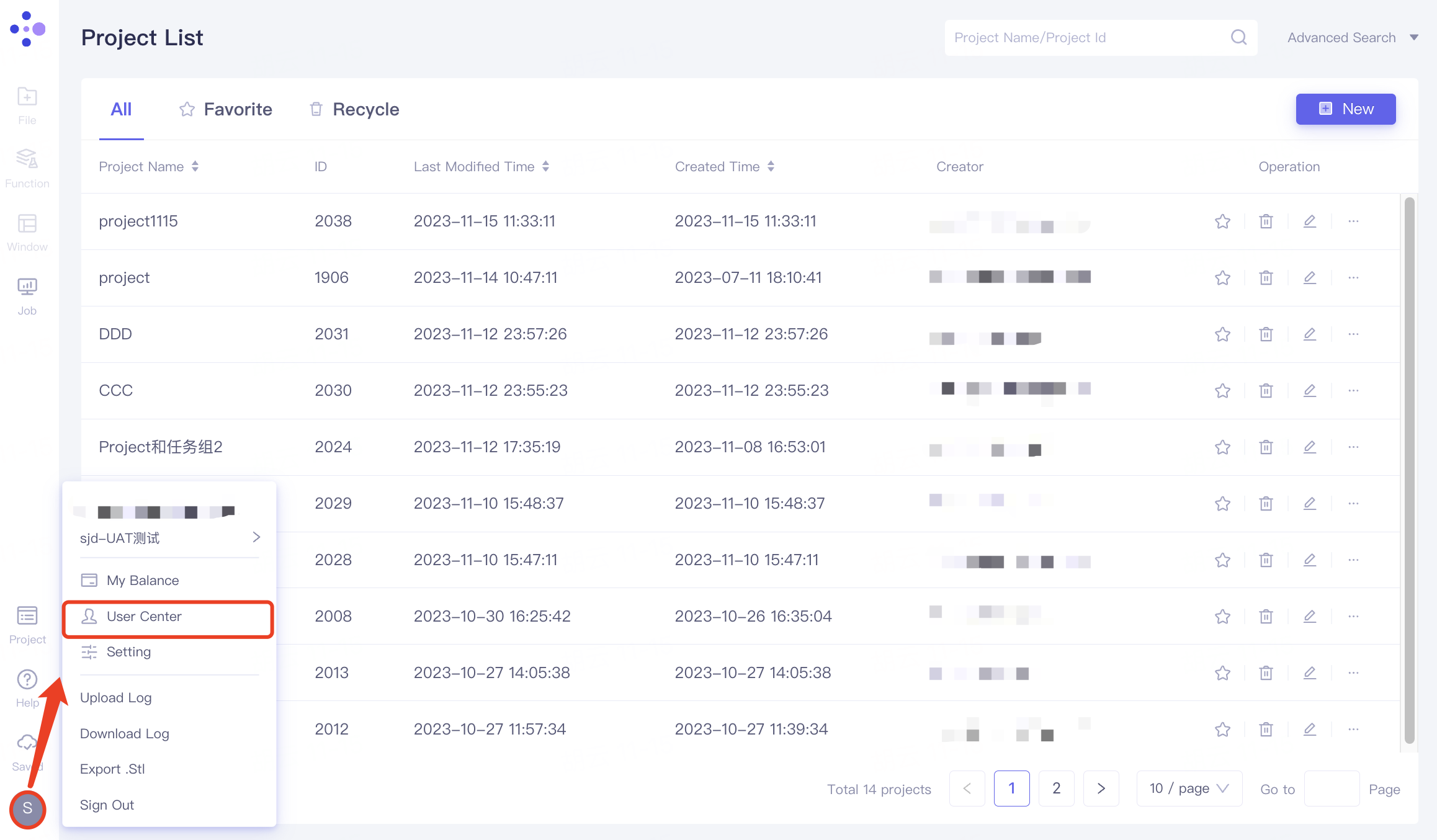
Click Edit Information, name the organization at Organization Name in the pop-up window, and click "Submit".
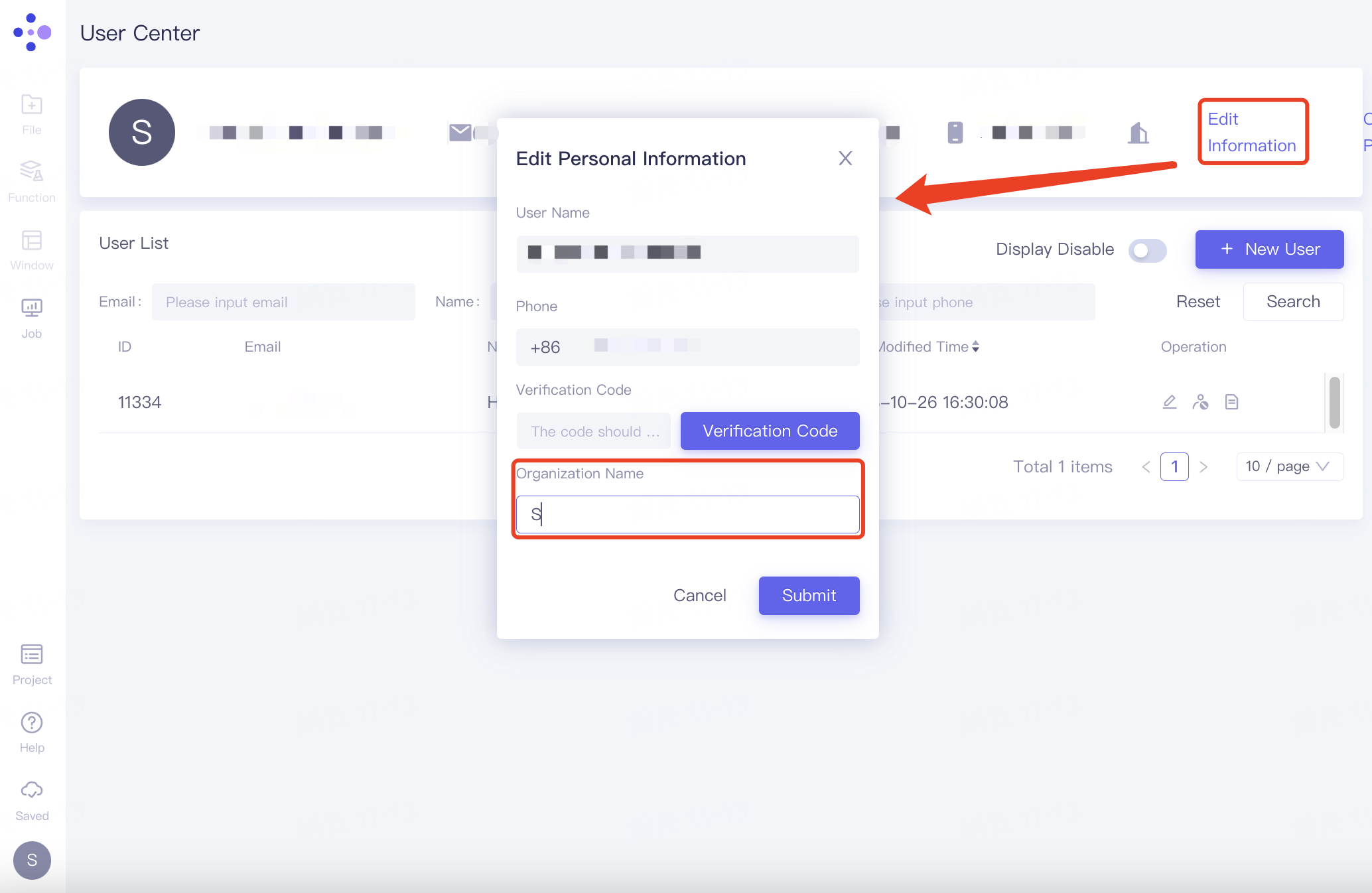
The administrator invites others to join the organization through "+ New User". Click "+ New User" to pop up the "Edit User" window, fill in the email address of the invitee in "* Email", select the shareable project under "Project", the selected project will be listed under "Project List", click "Submit" to submit for processing.
Project search and processing:
1)Search Projects: Search for projects based on Project Name/ID. Projects that meet the conditions will appear in the drop-down interface. Click to load them into the Project List.
2) Duplicate from Member: Search for a certain person based on User Name/ID/Email, click on the person on the drop-down interface to display all the projects of the person in the Project List.
3) Click "All" on the right side of Project List to load all projects under Project List.
4) Click "Clear" on the right side of Project List to clear all items under Project List.
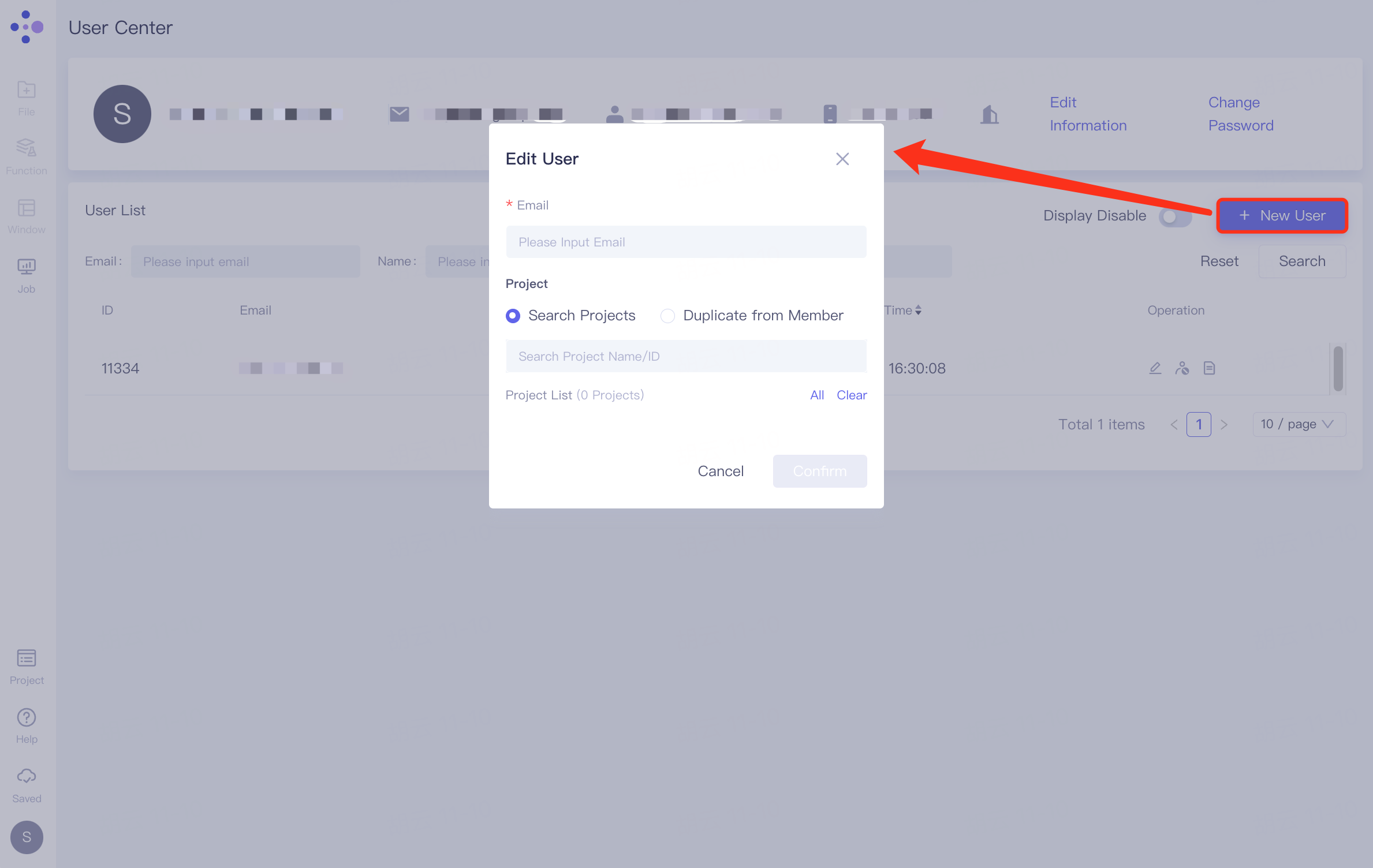
- Members are invited to join the organization
After the administrator completes the relevant operation, a confirmation email will be received in the member's mailbox. Follow the email prompts to operate;
Log in to Hermite, click Account, and switch organizations under Email number in the expanded list.
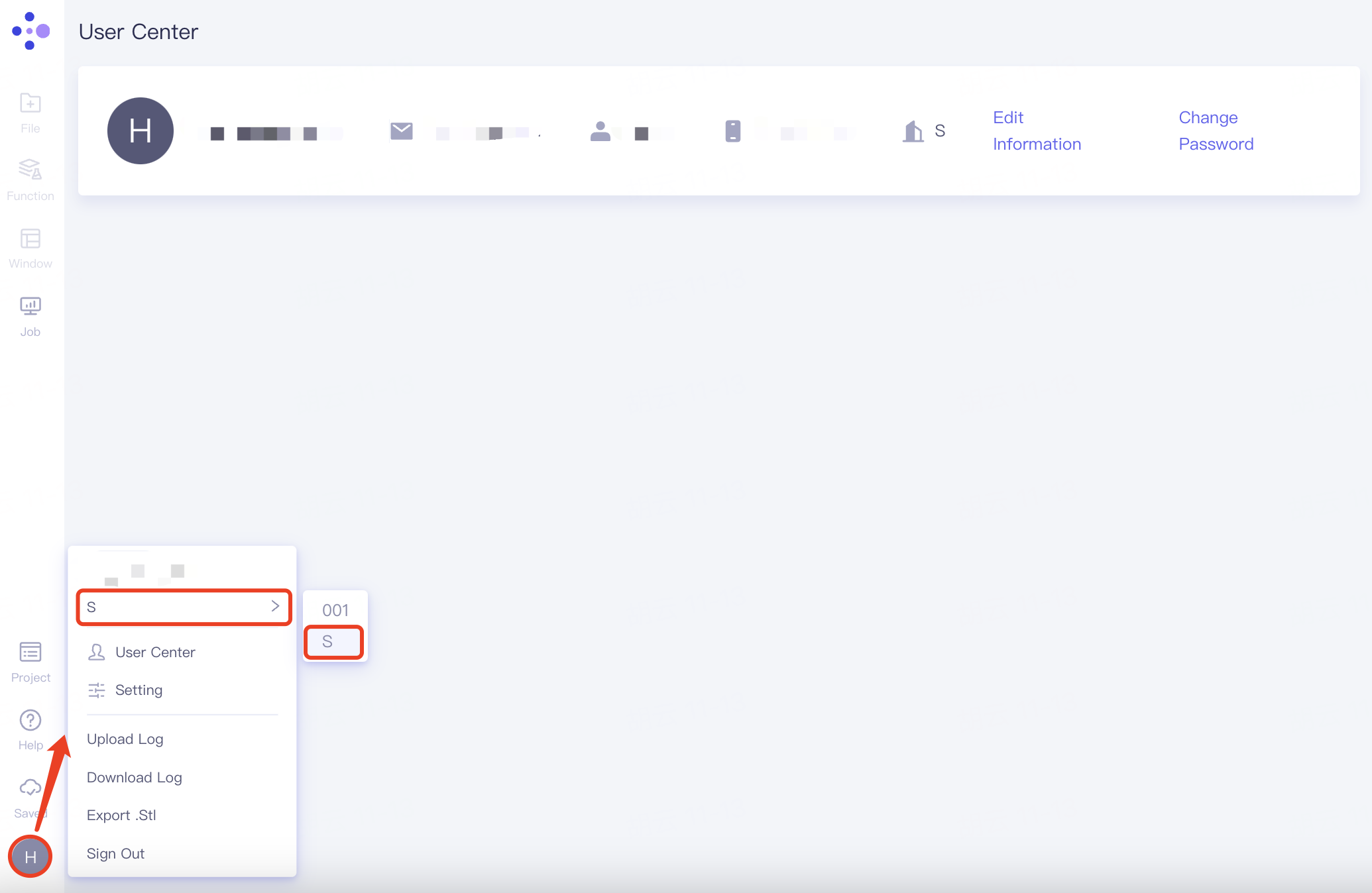
2.8.4 Setting
Setting: Support setting the color, size, font type of labels in 3D Workspace, as well as the background color of the 3D Workspace interface. Click Apply to change the settings.
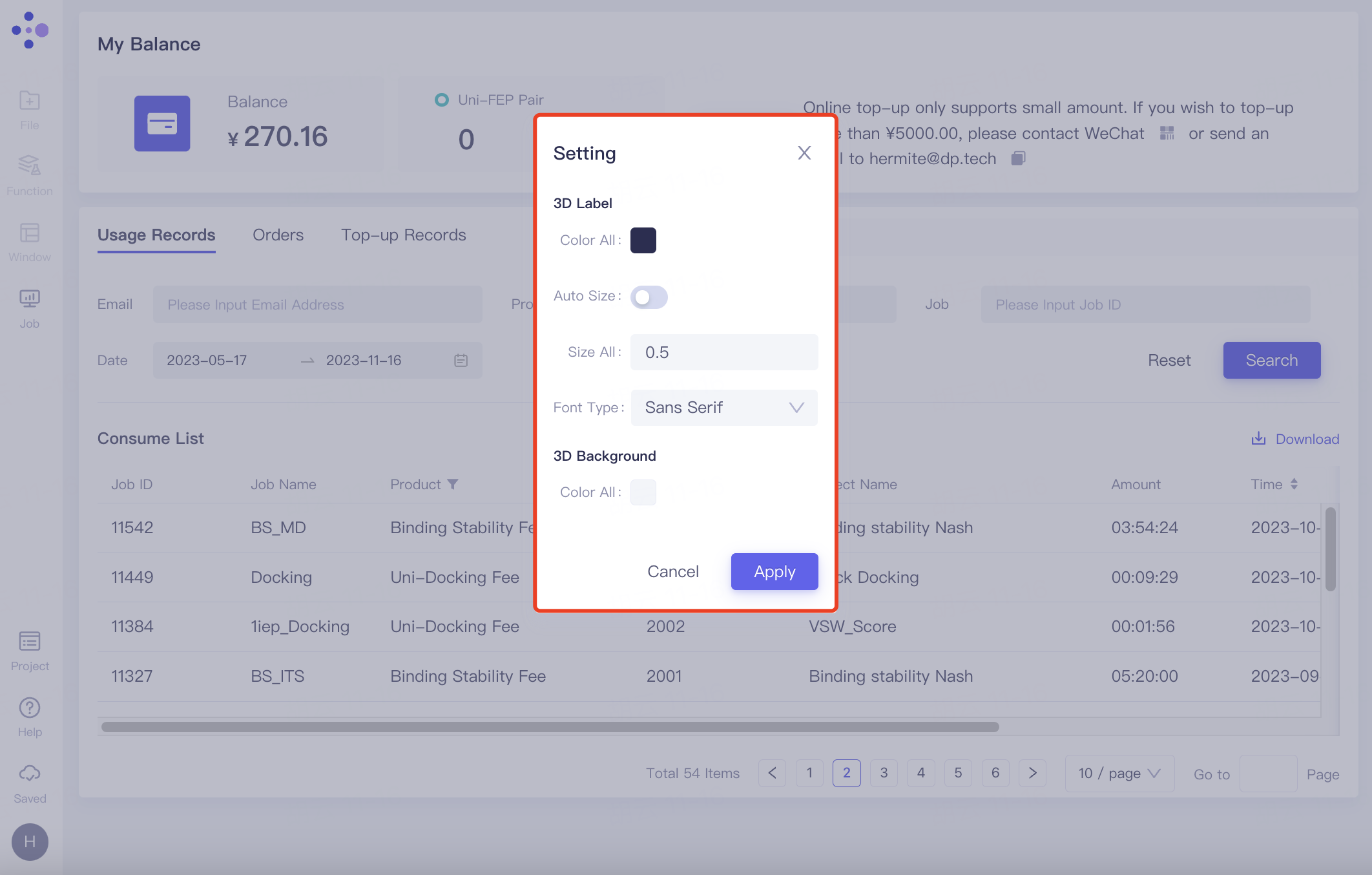
2.8.5 Upload Log
Upload Log.
2.8.6 Download Log
Download Log.
2.8.7 Export. Stl
Export to .stl format file.
2.8.8 Sign Out
Log out.