EM Structure Fitting
Introduction
In recent years, along with a series of breakthroughs in the hardware and algorithms of cryo-electron microscopy (cryo Electron Microscopy, cryo-em) technology, the applicability and results resolution of cryo-em technology have been steadily improved, and it has become an important research tool in the field of structural biology. In the process of processing cryo-em data, it is very important to use the reconstructed three-dimensional density map to model biomolecules and further optimize the atomic coordinates. The EM Structure Fitting module (Hermite ® Uni-EM structure fitting) of the Hermite ® platform provides the flexible building function of the all-atom structure model based on the cryo-em density map. With the friendly operation interface, optimized algorithm and flexible cloud computing resources of Hermite platform, you can easily and quickly obtain high-quality cryo-em molecular structure model consistent with the density map.
7 XXB is the crystal structure of Arabidopsis thaliana auxin transporter PIN3. In this course, the protein structure predicted by the 7XXB sequence and the electron microscope density map of 7 XXB are selected to optimize the predicted structure based on the EM Structure Fitting module. The protein structures before and after optimization were analyzed and compared by calling the Protein Alignment module.
The files required for this tutorial are as follows:
1. EM Structure Fitting
1.1 Entrance
- The left general menu bar 'Function' → Structure Modeling → EM Structure Fitting.
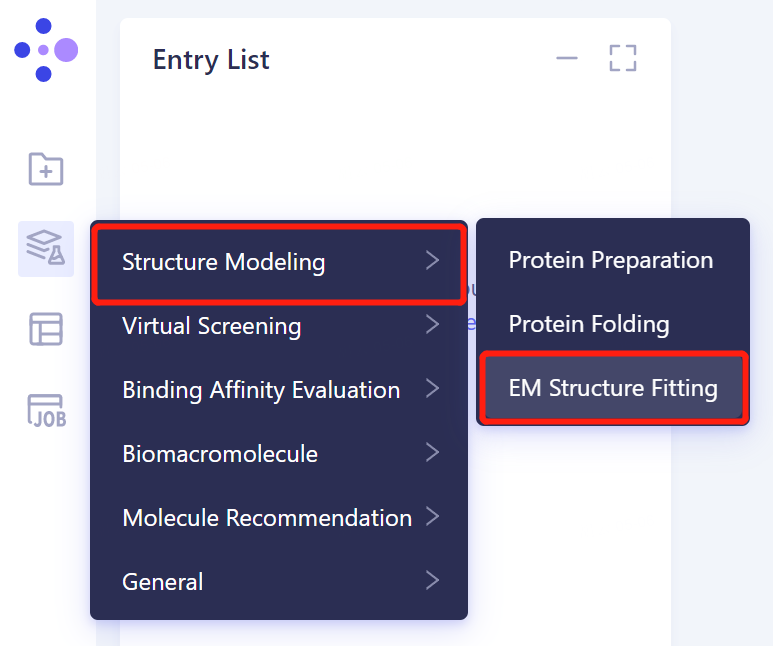
- The operation box of the EM Structure Fitting (shown in the red box) appears on the right side, and the overall interface is as follows:
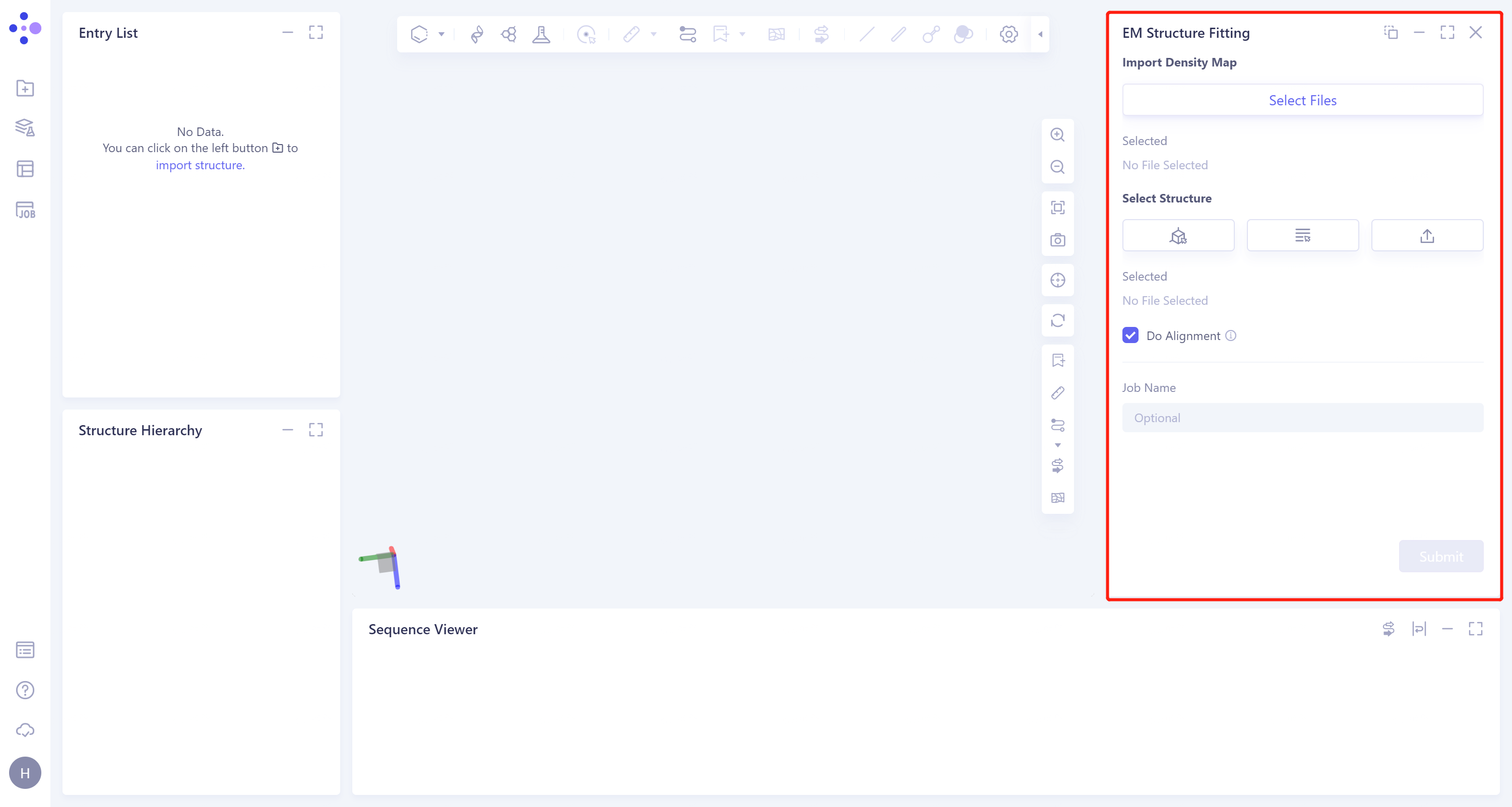
1.2 Operation
-
Click 'Select File' in the 'Import Density Map' to upload the electron microscope density map. After uploading, the uploaded file (7xxb_EMD_33500.ccp4) will be displayed at the bottom. The electron microscope density map will be displayed in the 3D Workspace window;
-
Under 'Select Structure', click 'Select File' to upload the protein structure. After uploading, the uploaded file (7XXB-Fold.pdb) will be displayed at the bottom, and the protein structure will be displayed in the 3D Workspace window;
-
Check 'Do Alignment' to compare the input structure with the electron microscope density map;
-
Name the task as 'EM_7XXB' at 'Job Name'.
-
Click 'Submit' to submit the task.
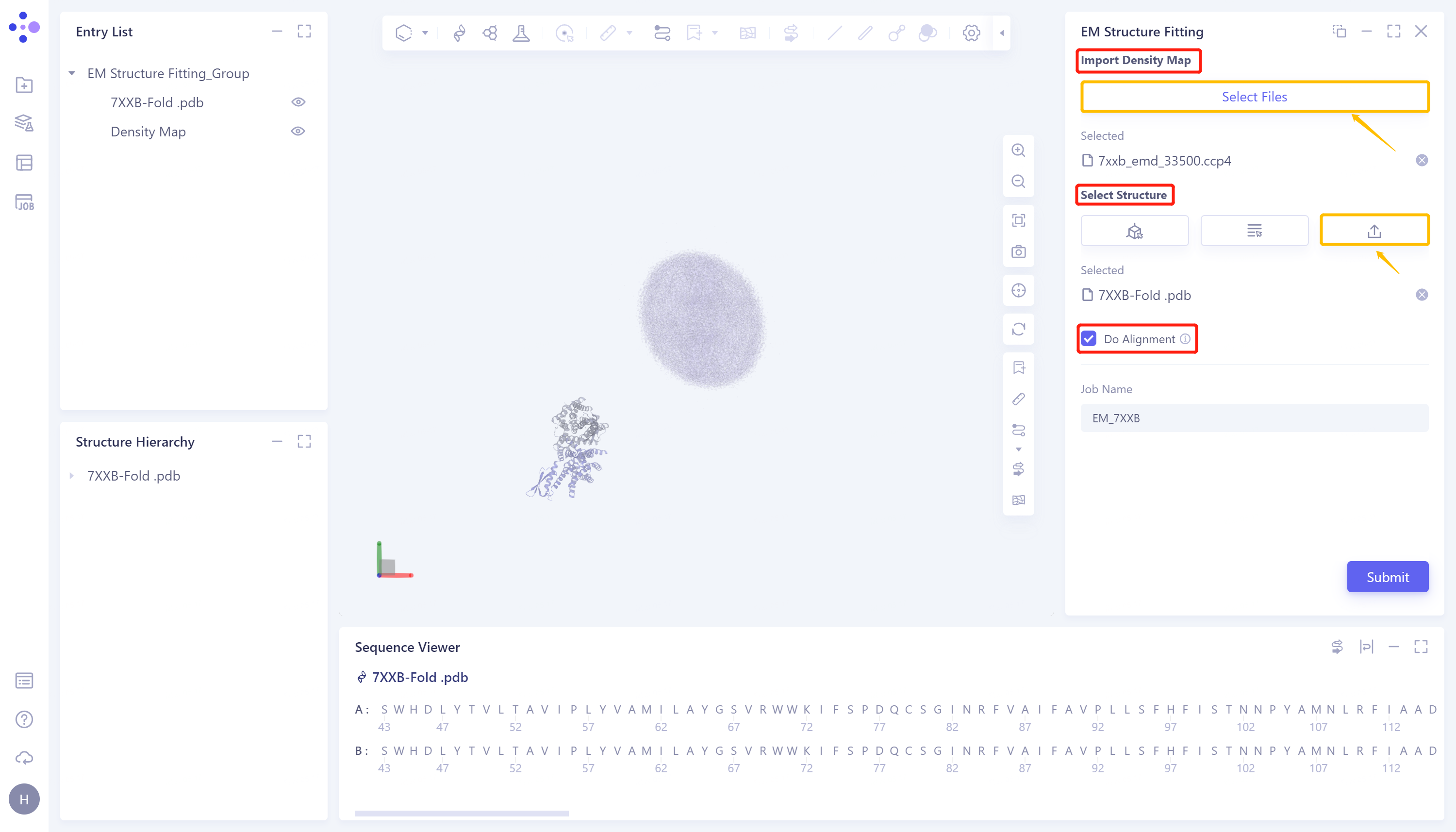
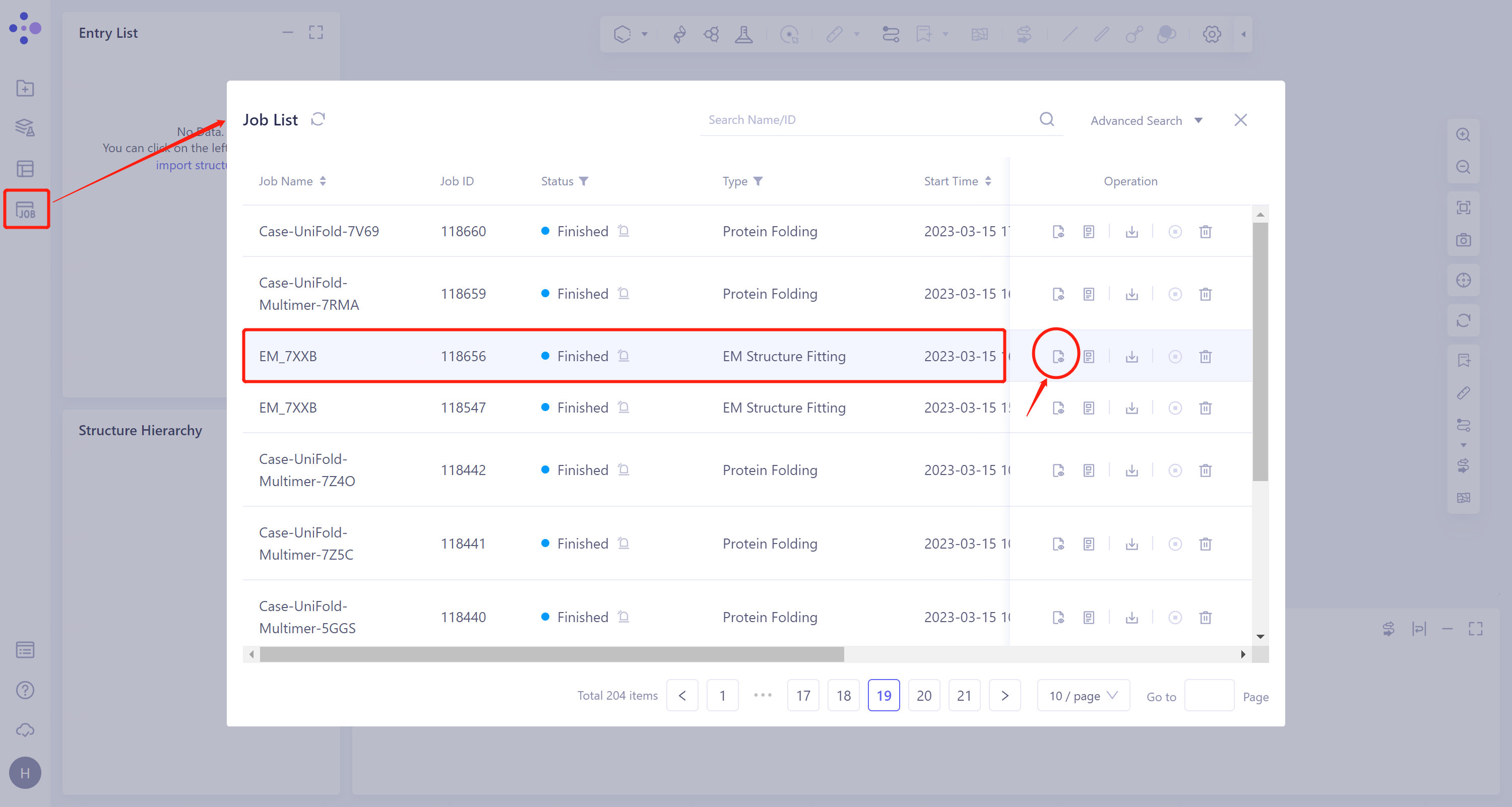
1.3 Display of results
- General menu bar on the left: Job → Find the task 'EM_7XXB' → Click 'Show' under the Operation column to display the results of the task, as shown in the figure below.
 | 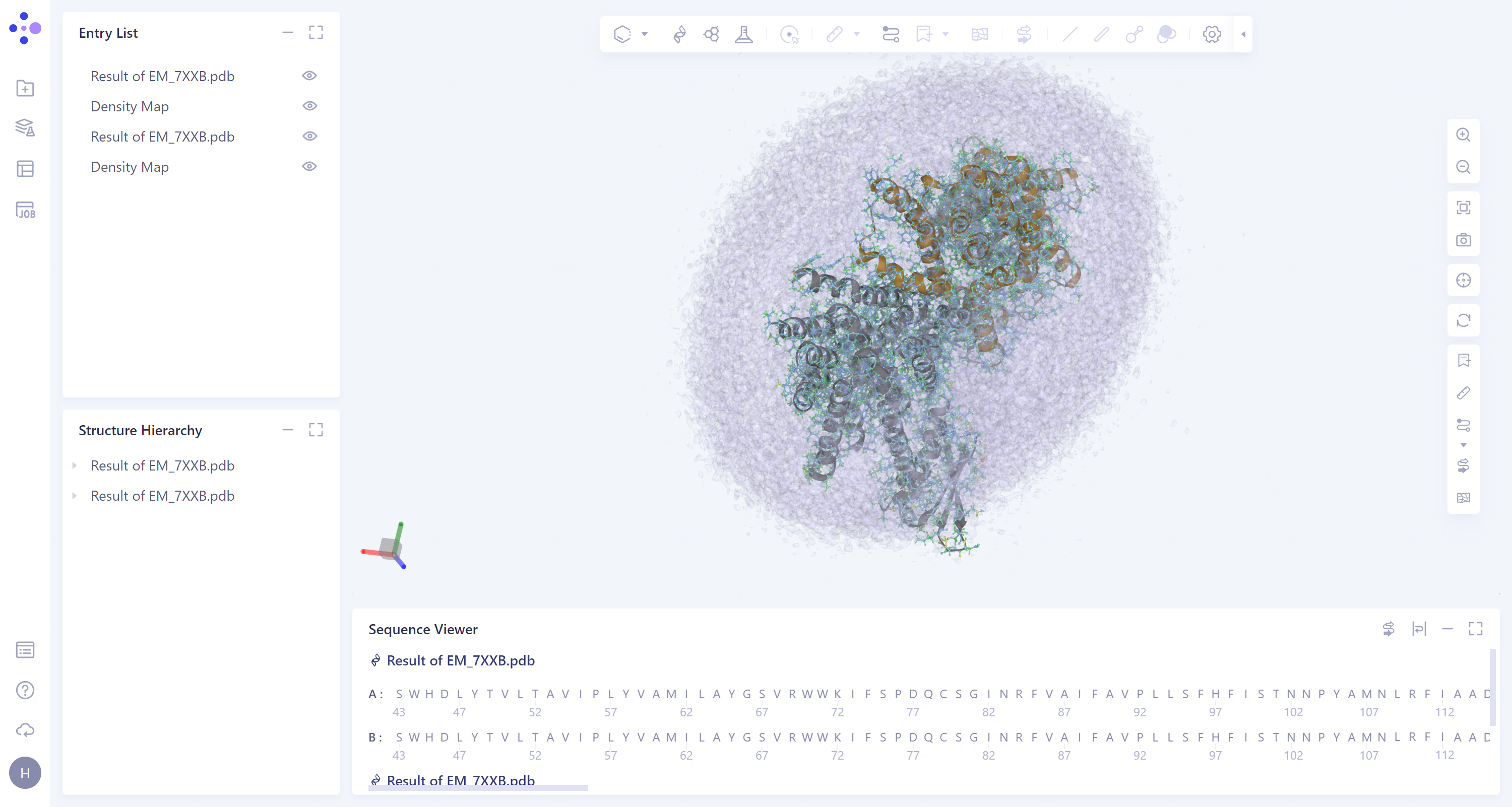 |
- The results are displayed in the 3D Workspace area. Turn off the display of the Protein Ribbon by clicking on the 'Ribbon' icon on the right.
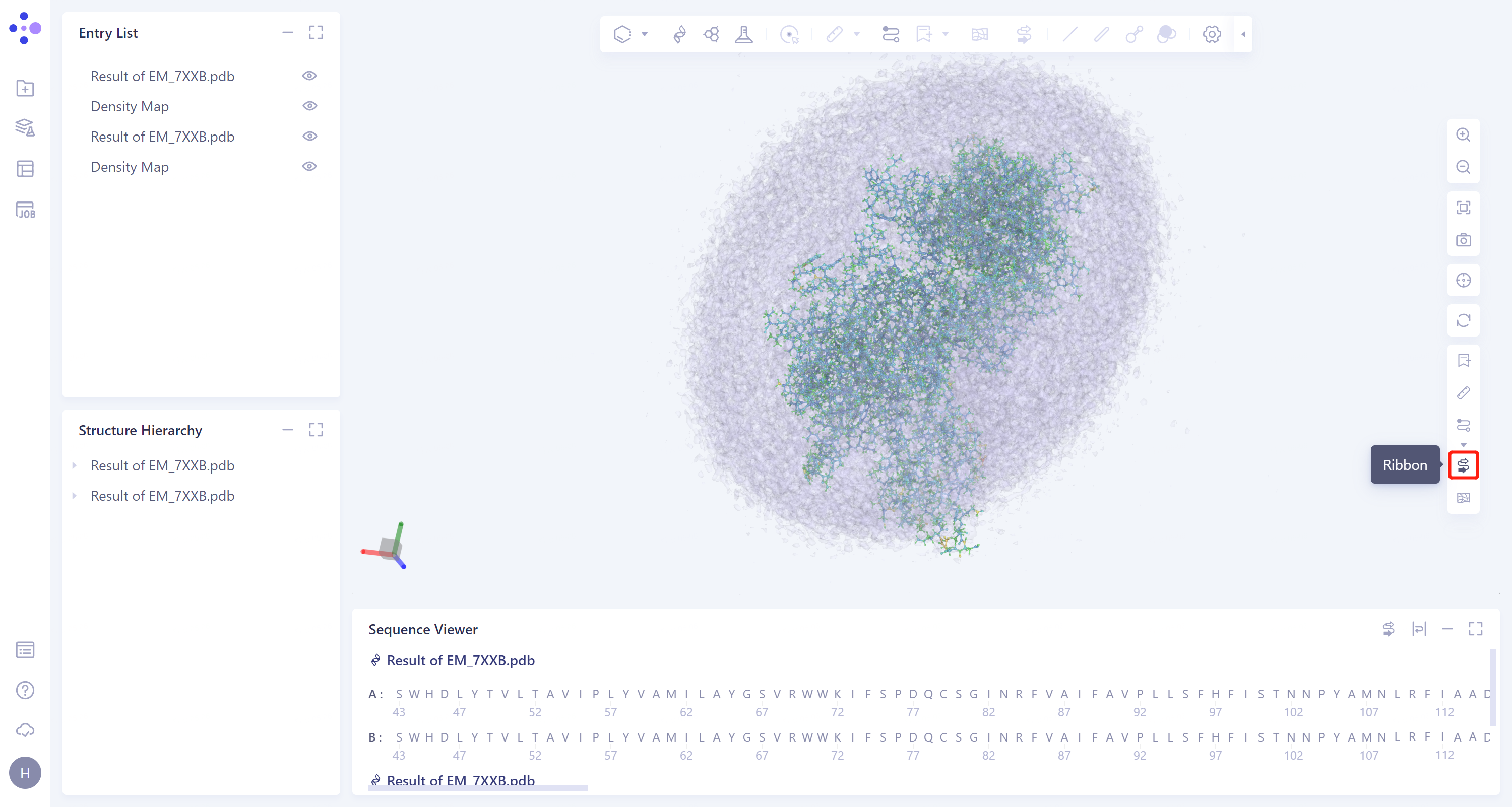
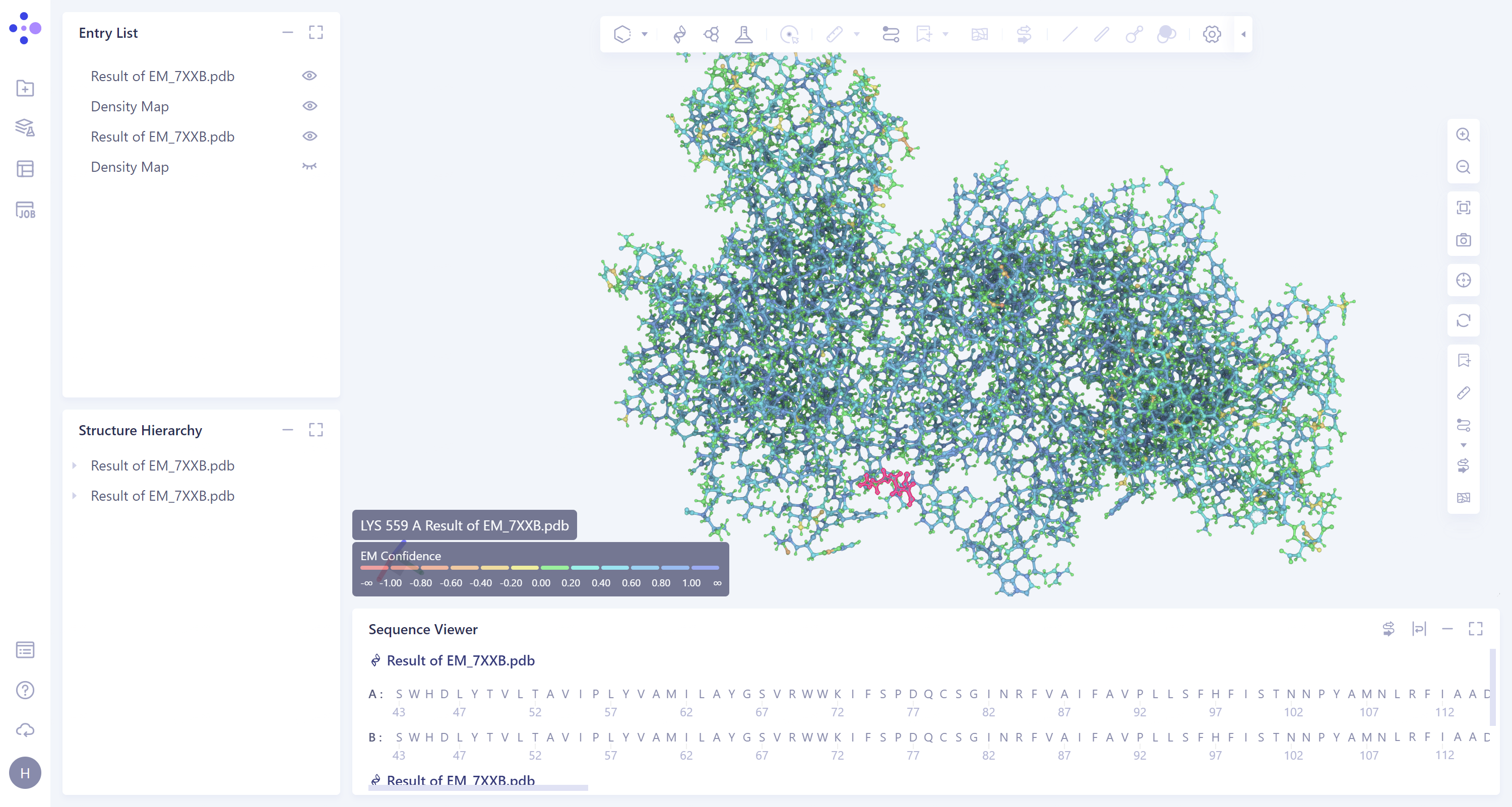
- According to the color information of protein residues, it can be seen that the confidence of the protein structure is high.
2. Protein Alignment
2.1 Entrance
- Left general menu bar Menu → Function → General → Protein Alignment.
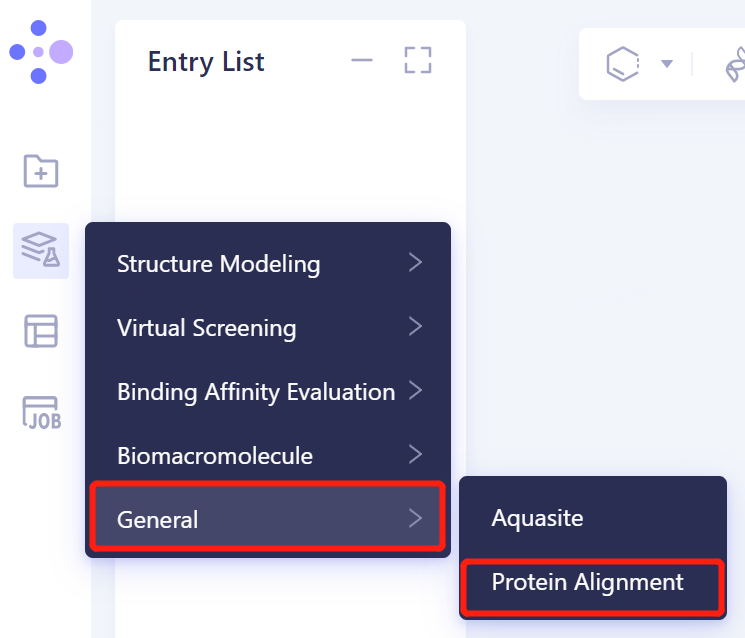
- The operation box of Protein Alignment (shown in the red box) appears on the right side, and the overall interface is as follows:
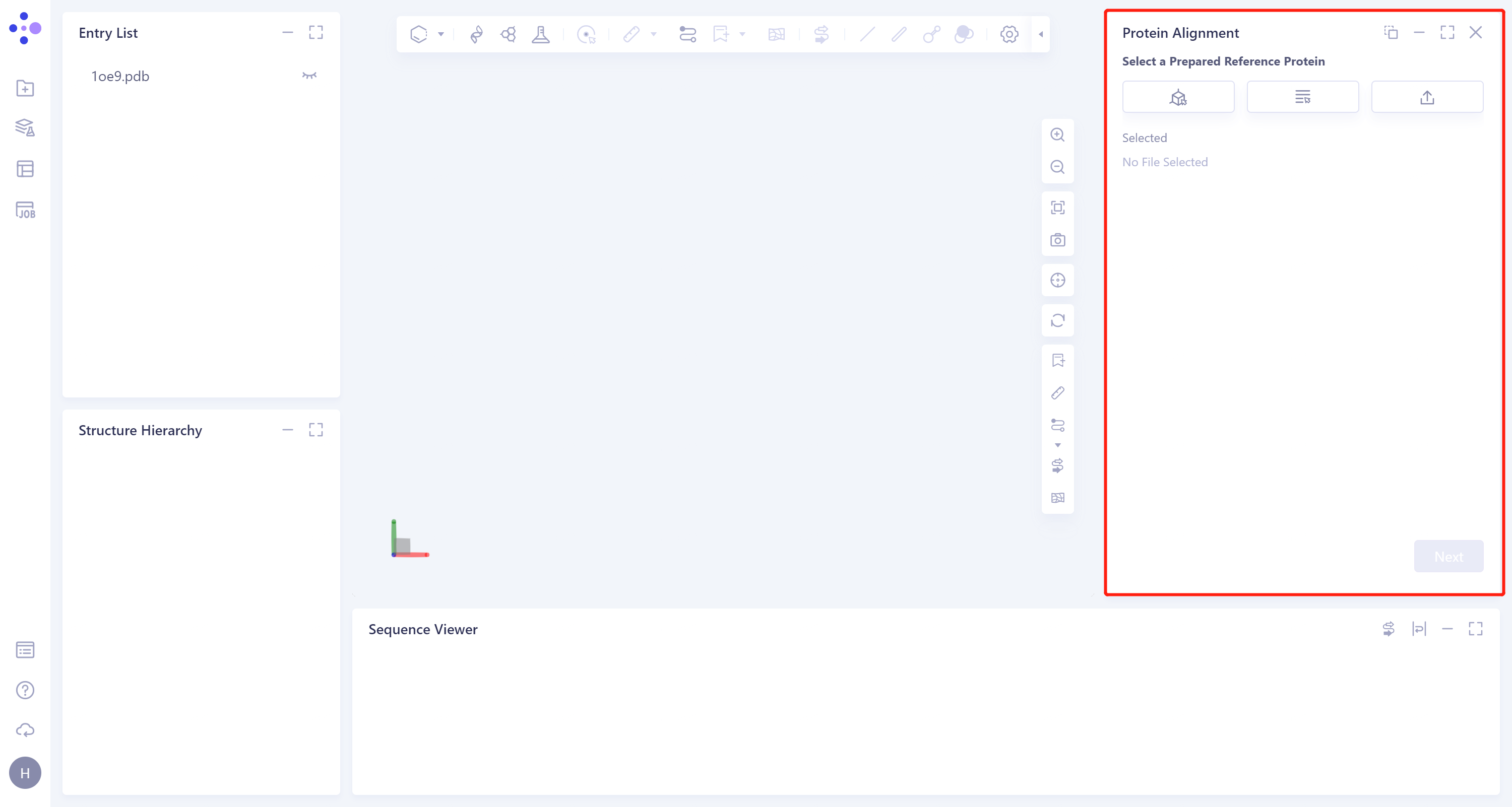
2.2 Operation
2.2.1 Protein structure alignment after EM Structure Fitting
2.2.1.1 Introduction of reference protein
- In the same project, click the left general menu bar 'File' → Get PDB → Input '7XXB' in the pop-up window to enter the PDB ID: 7XXB → Import.
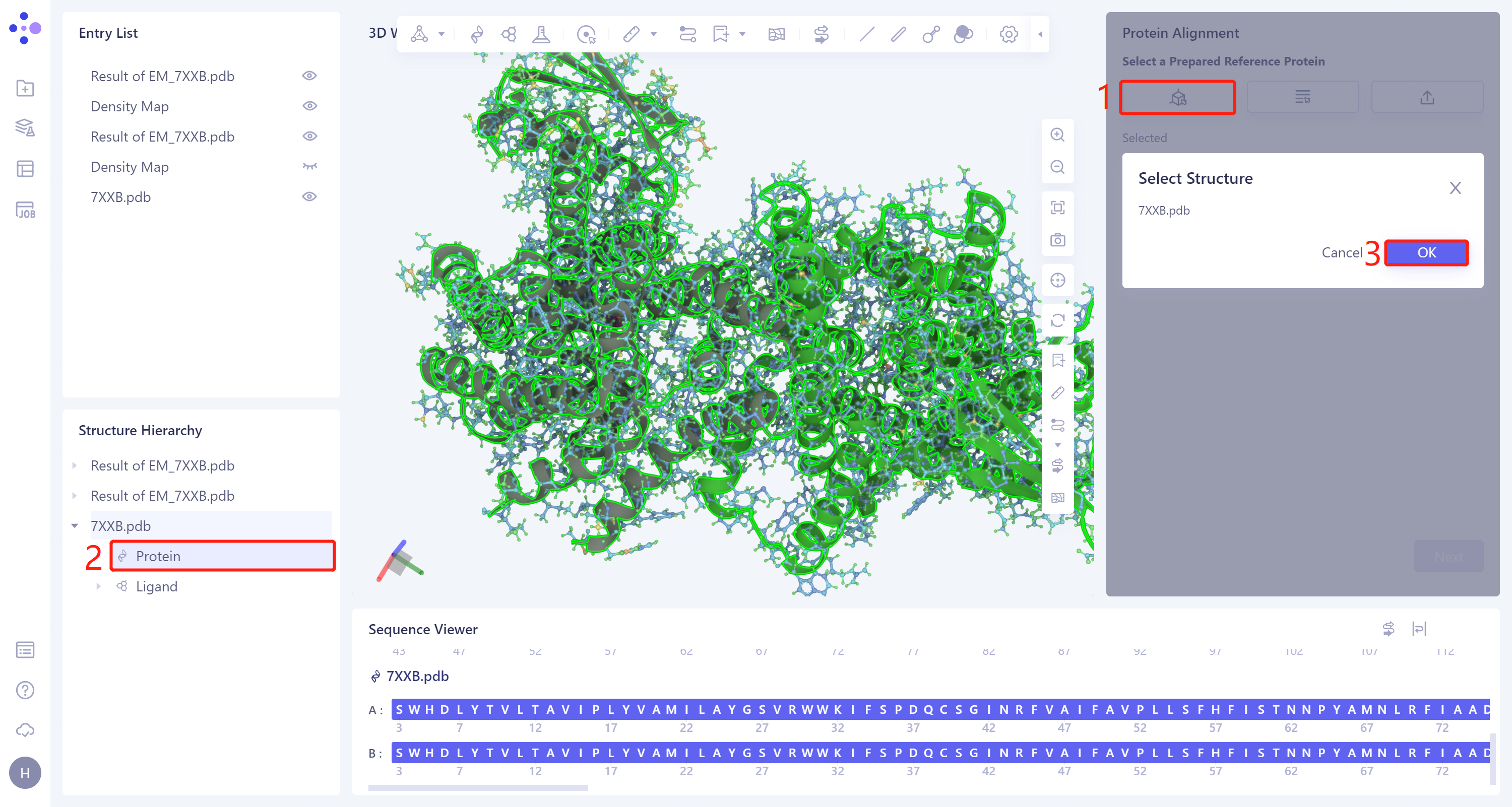
- The 'Protein Alignment' interface appears on the right side of the interface. First, import the reference protein structure: click 'Select Structure', the 'Select Structure' window pops up → Select the protein structure of '7XXB' in the Structure Hierarchy → Click 'OK' in the 'Select Structure' window.
2.2.1.2 Introduction of proteins to be compared
- Click 'Next' to enter the comparison protein selection window and import the protein structure to be compared: click 'Select from Project', The 'Select from Project' window pops up → Select Result of 'EM_7XXB.pdb' in the 'Select from Project' window → Click 'OK'.
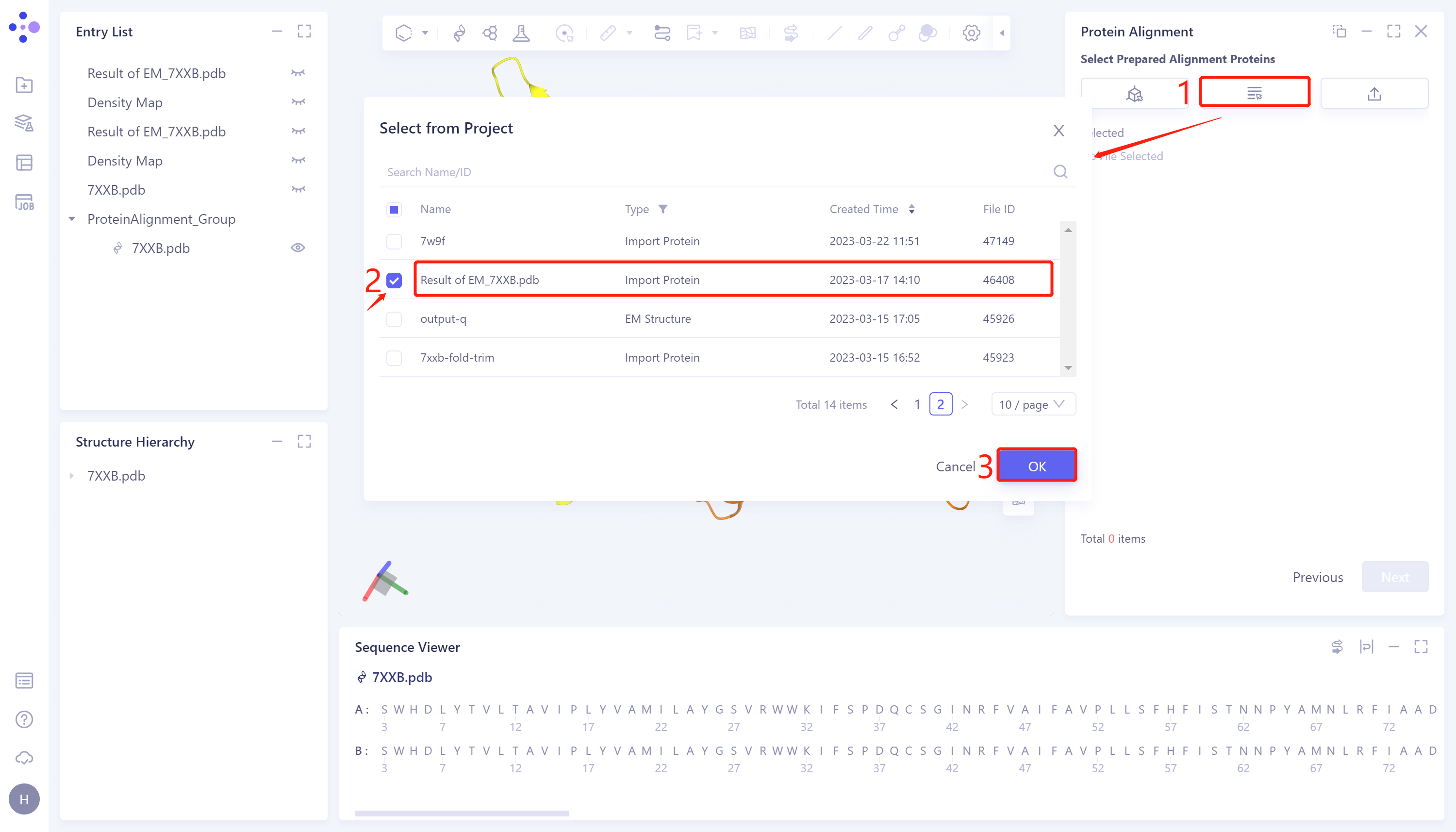
2.2.1.3 Parameter setting
- Click 'Next' to enter the parameter setting page. 'Alignment Atoms' selects 'Cα' to perform alignment based on the α-carbon atoms between proteins. Since there is no special comparison requirement, the parameter setting of 'Select Binding Site' is skipped. Name the task as '7XXB_EM_PA' and click 'Submit' to submit the task.
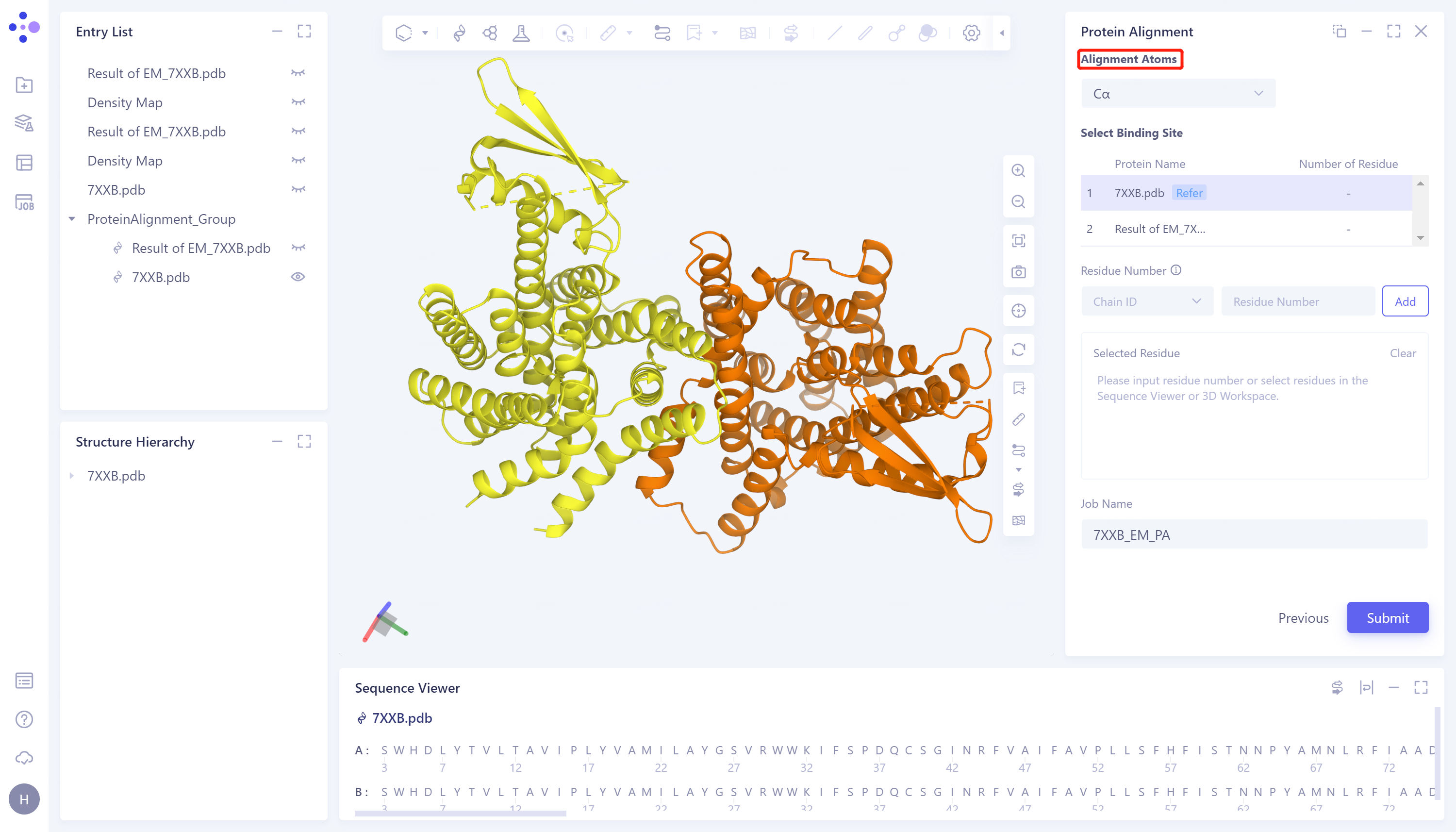
2.2.2 Protein structure alignment before EM Structure Fitting
- The operation steps are the same as those in 2.2.1 (select '7XXB-Fold.pdb' for the 'Select Prepared Alignment Proteins'), and the task is named as '7XXB-Fold PA'.
2.3 Analysis of results
2.3.1 Inlet
-
Job → Find Task '7XXB_EM_PA' → Click 'Show' to display the results of the task, as shown in the figure.
-
The results of '7XXB-Fold PA' task are viewed as above.
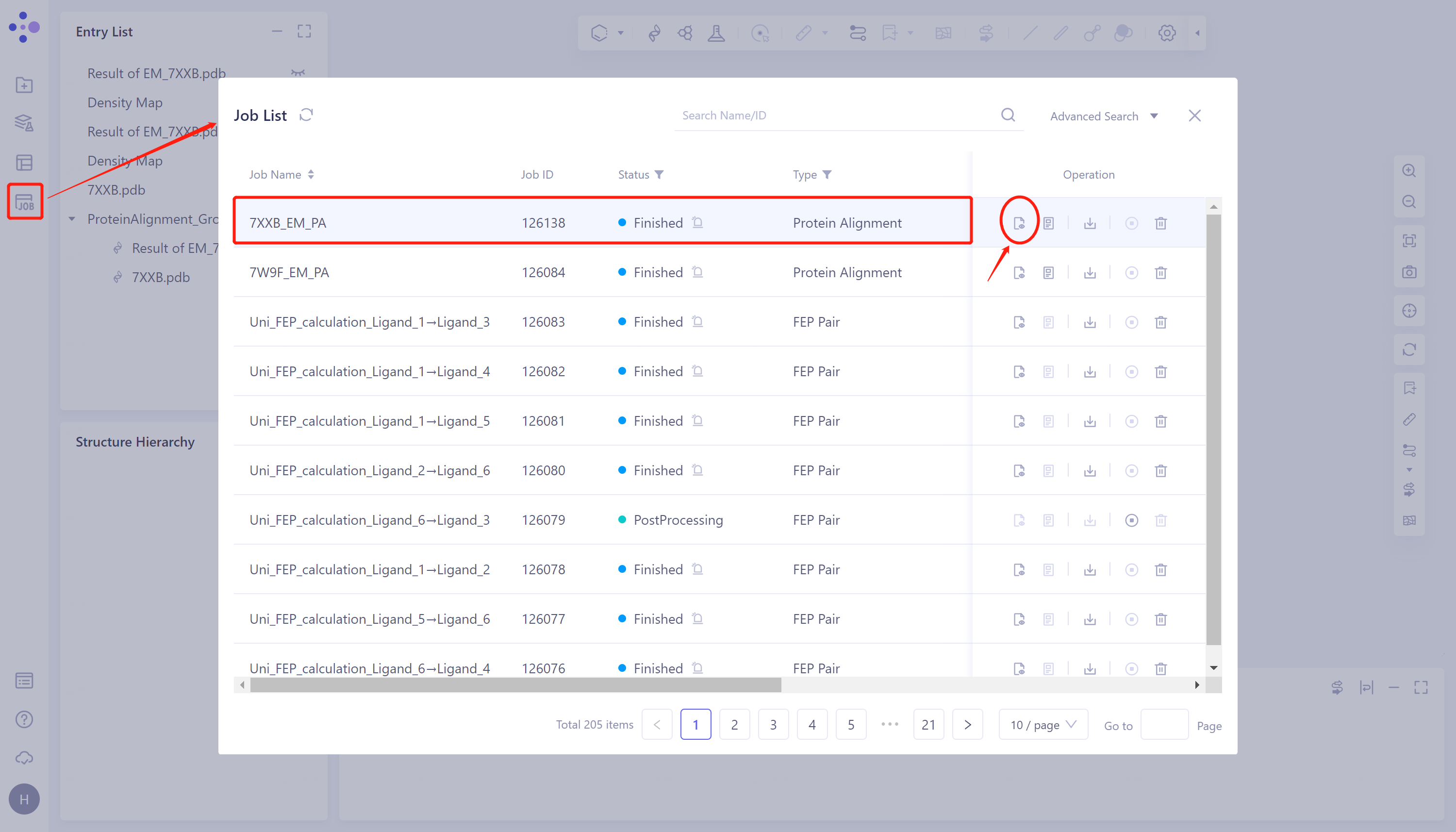 | 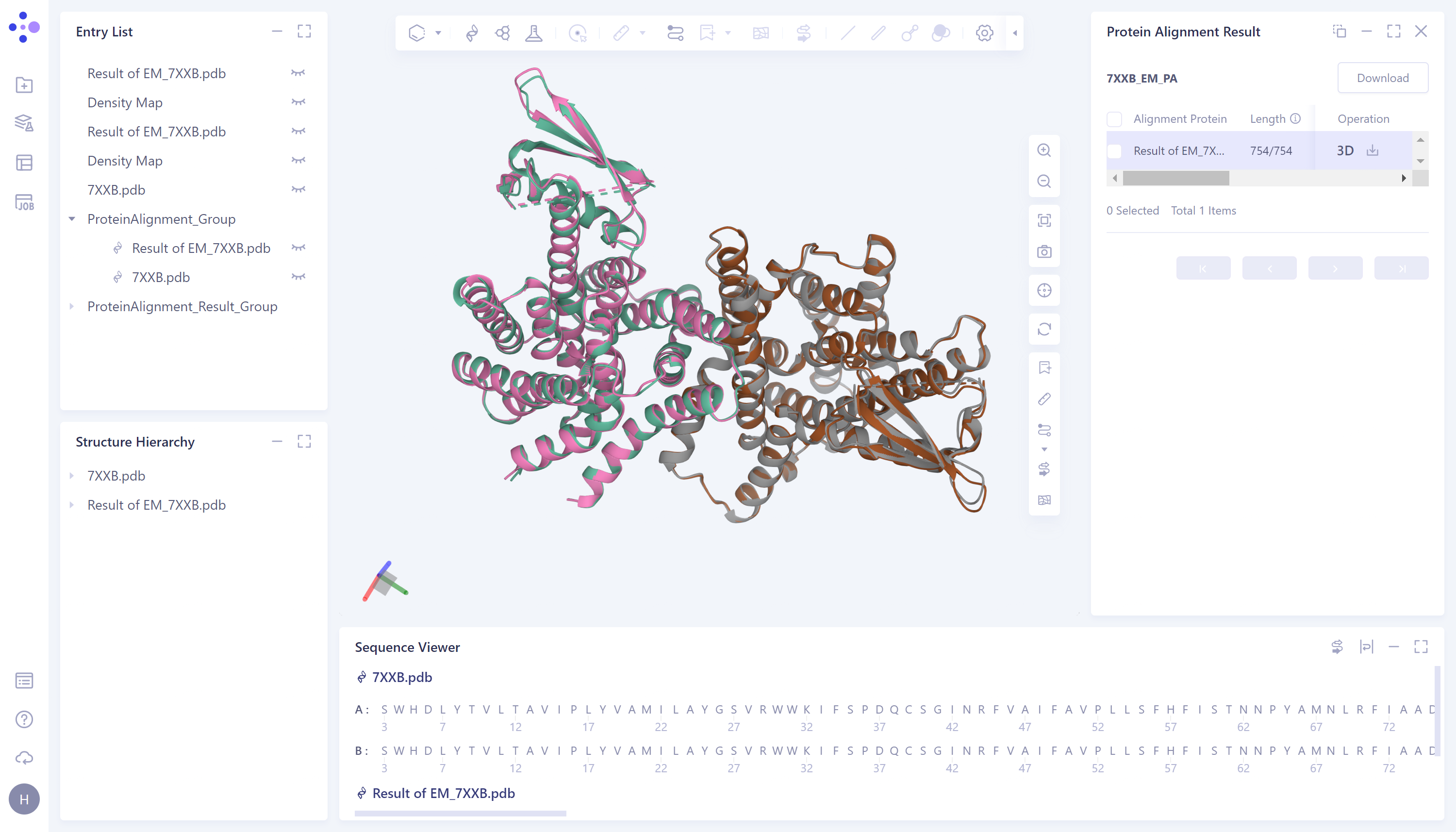 |
2.3.2 Comparison of results
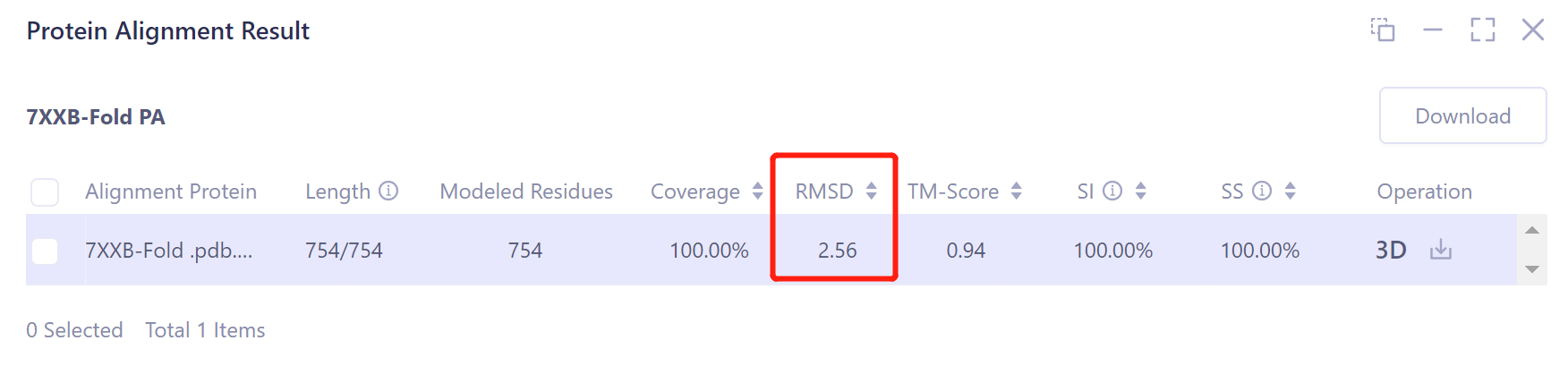
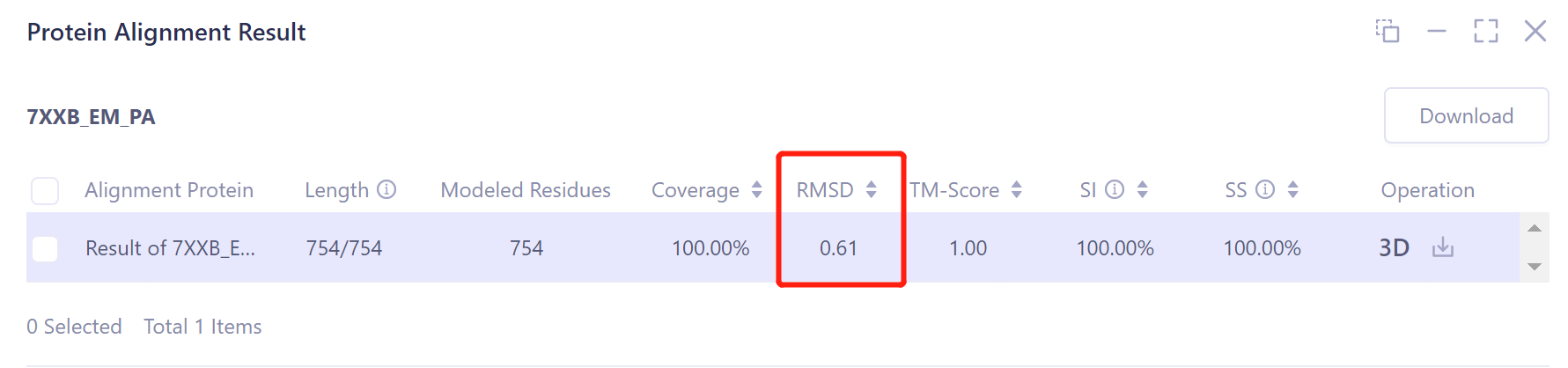
- According to the comparison, the structural deviation (RMSD) between the protein structure before EM Structure Fitting optimization and its experimentally measured crystal structure 7XXB is 2.5656 Å (upper panel). The structural deviation (RMSD) of the protein structure optimized by EM Structure Fitting from that of 7XXB is only 0.6161 Å (the figure below), which means that the optimization effect of EM Structure Fitting on the initial structure of the protein according to the electron microscopy density map is significant.
-
The visualization result of protein alignment is displayed in the 3D workspce window. It can be seen from the figure that the original input protein structure 7XXB-Fold.pdb contains a part that cannot be accurately predicted, and the information of this part is not recorded in the electron microscope density map, so this part is omitted after EM Structure Fitting.
-
After the optimization of the EM Structure Fitting, the structure of the protein can be better overlapped with the crystal structure obtained from the experimental observation.
Before 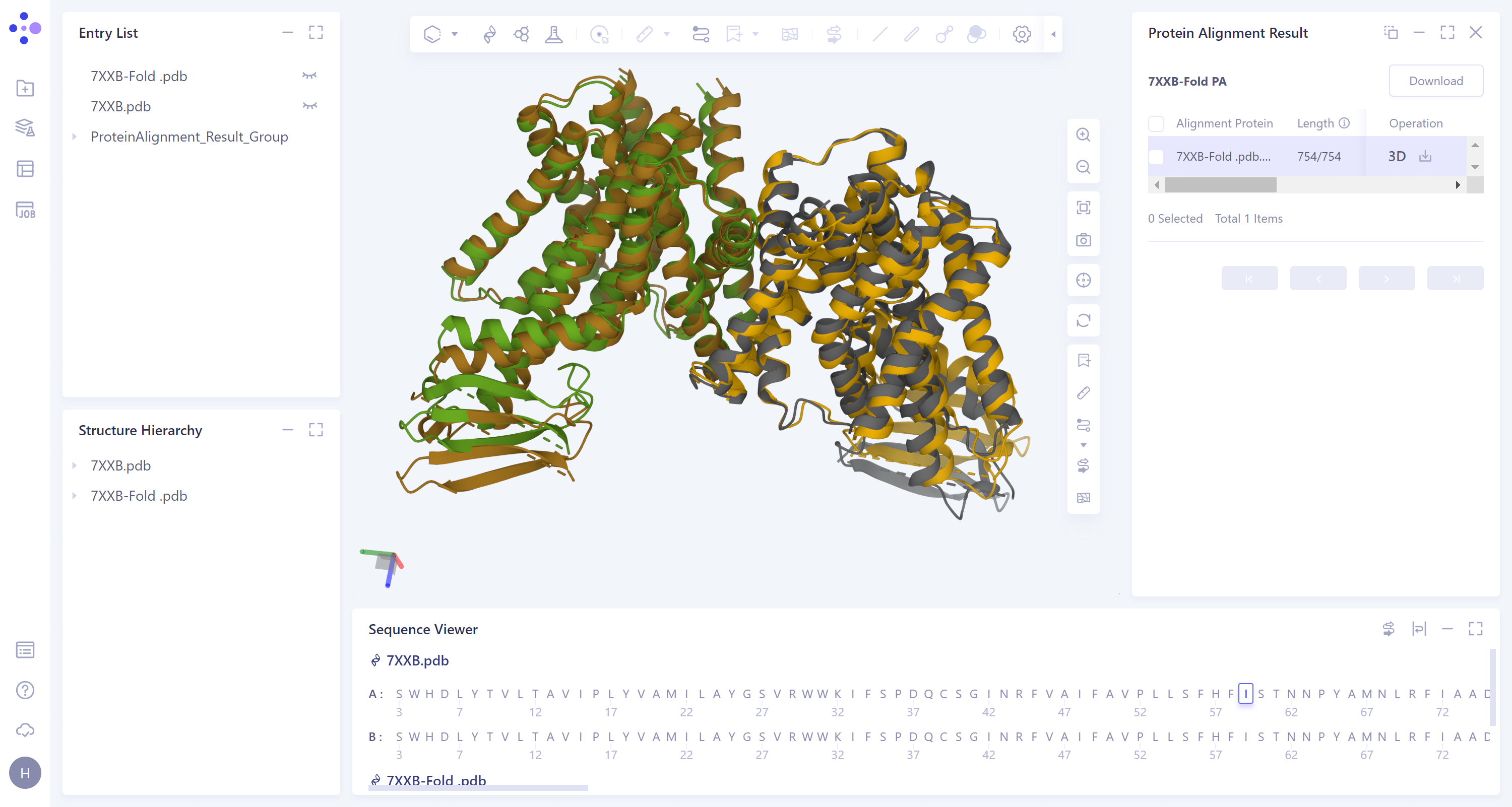
EM Structure Fitting optimization
After 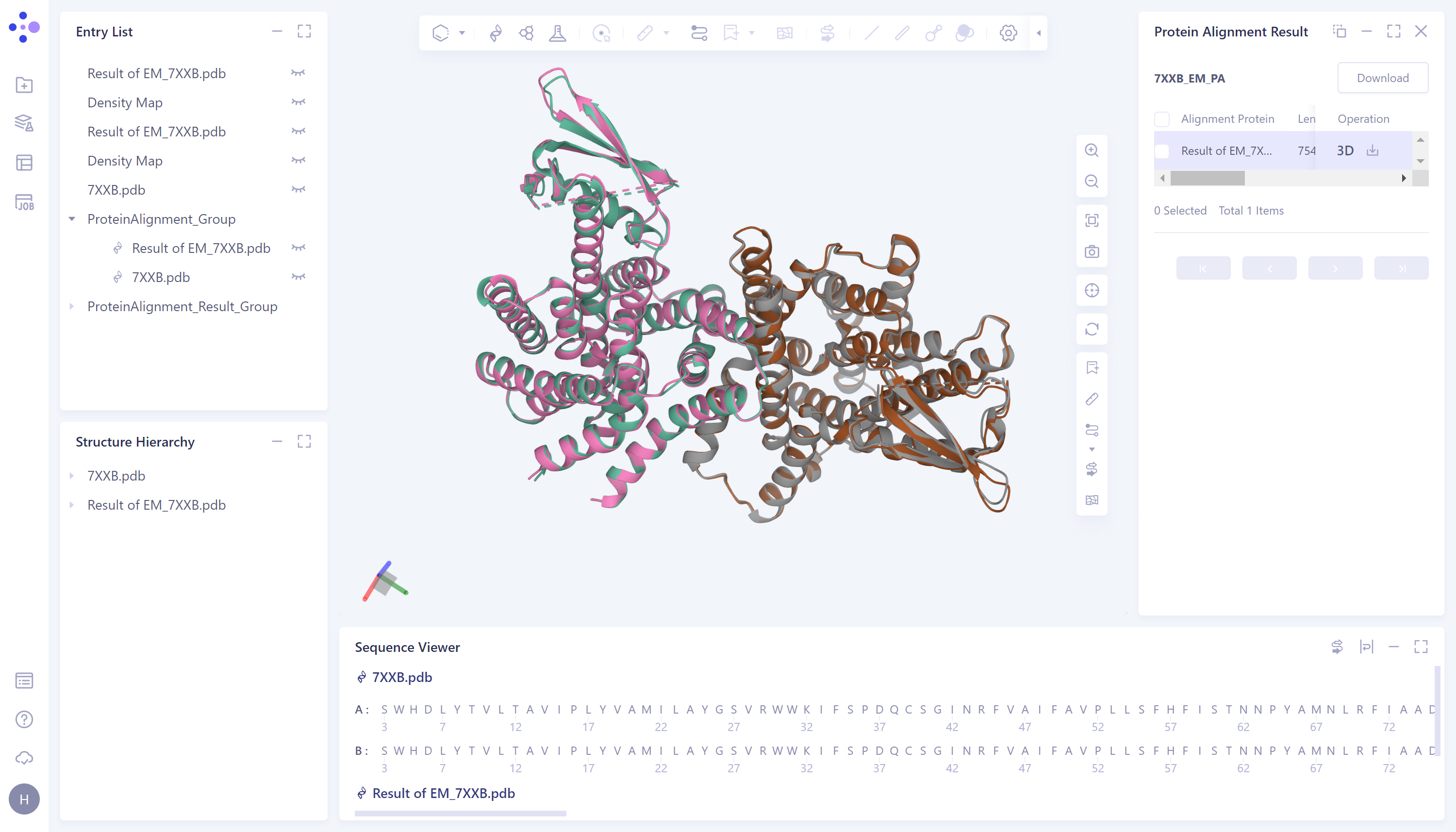
EM Structure Fitting optimization