Antibody Humanization
Introduction
Non-human antibodies can cause severe body rejection when they enter the human body, and then affect the safety and therapeutic effect of antibodies in clinical application. Therefore, it is necessary to humanize the antibody to reduce the heterology of the antibody as much as possible, and to keep its specificity and affinity unchanged.
The novel coronavirus epidemic caused by SARS-CoV-2 virus has swept the world, and studies have found that SARS-CoV-2 Spike protein can be used as a target for drug design. SARS2-57 is a murine antibody targeting the N-terminal region of the SARS-CoV-2 Spike. This tutorial is based on the variable region sequence of SARS2-57 to design its antibody humanization.
The Antibody Humanization module of Hermite ® platform provides the function of antibody humanization. Based on the user-friendly graphical task submission and result analysis interface, you can easily and quickly complete the design of antibody humanization.
The data required for this tutorial is as follows:
7SWW_2|Chain B[auth H]|SARS2-57 Fv heavy chain|Mus musculus (10090)EVQLQQSGAELVRPGALVKLSCKASGFNIKDYFVHWVKQRPVQGLEWIGWIDPENGNTIYGPKFQGKASLAADTSSNTGYLQLSSLTSEDTAVYYCARWDGYETLDYWGQGTSVTVS
7SWW_3|Chain C[auth L]|SARS2-57 Fv light chain|Mus musculus (10090)DIVMSQSPSSLAVSVGEKVTMSCQSSQSLLYSNNEKNYLAWYQQKPGHSPKLLLYWASTRESGVPDRFTGSGSGTAFTLTISSVKAEDLAVYYCQQYYNYPYTFGGGTKLEI
1. How to use
1.1 Entrance
- The left general menu bar Menu → Function → Biomacromolecule → Antibody Humanization.
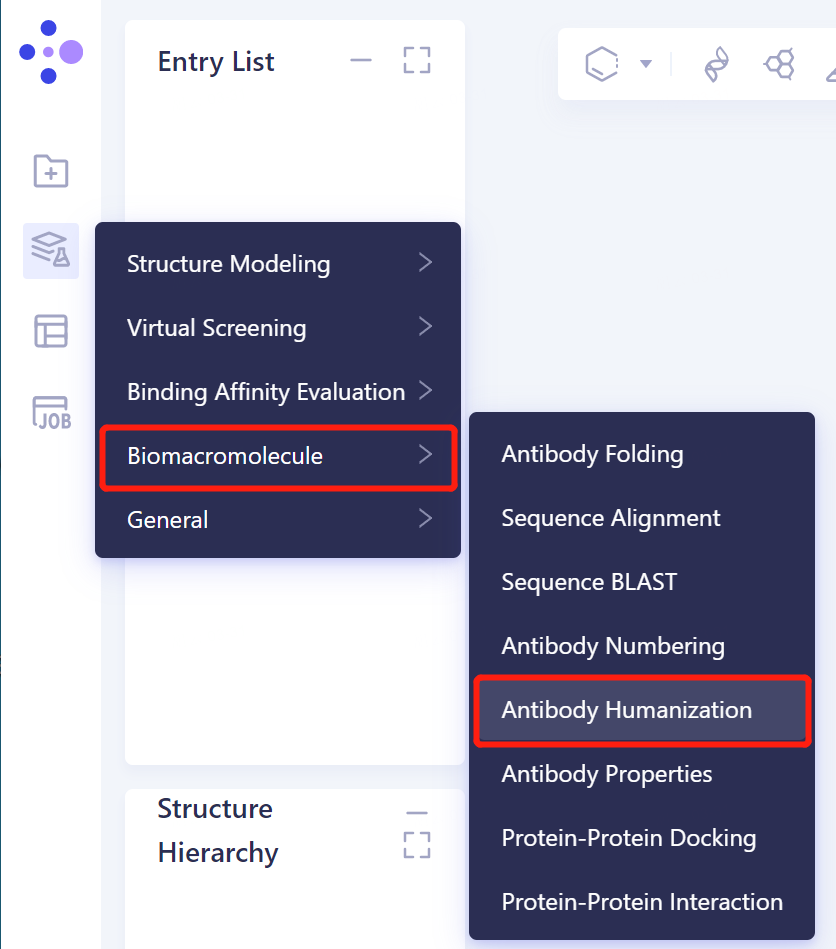
- The operation box of the Antibody Humanization (shown in the red box) appears on the right side, and the overall interface is as follows:
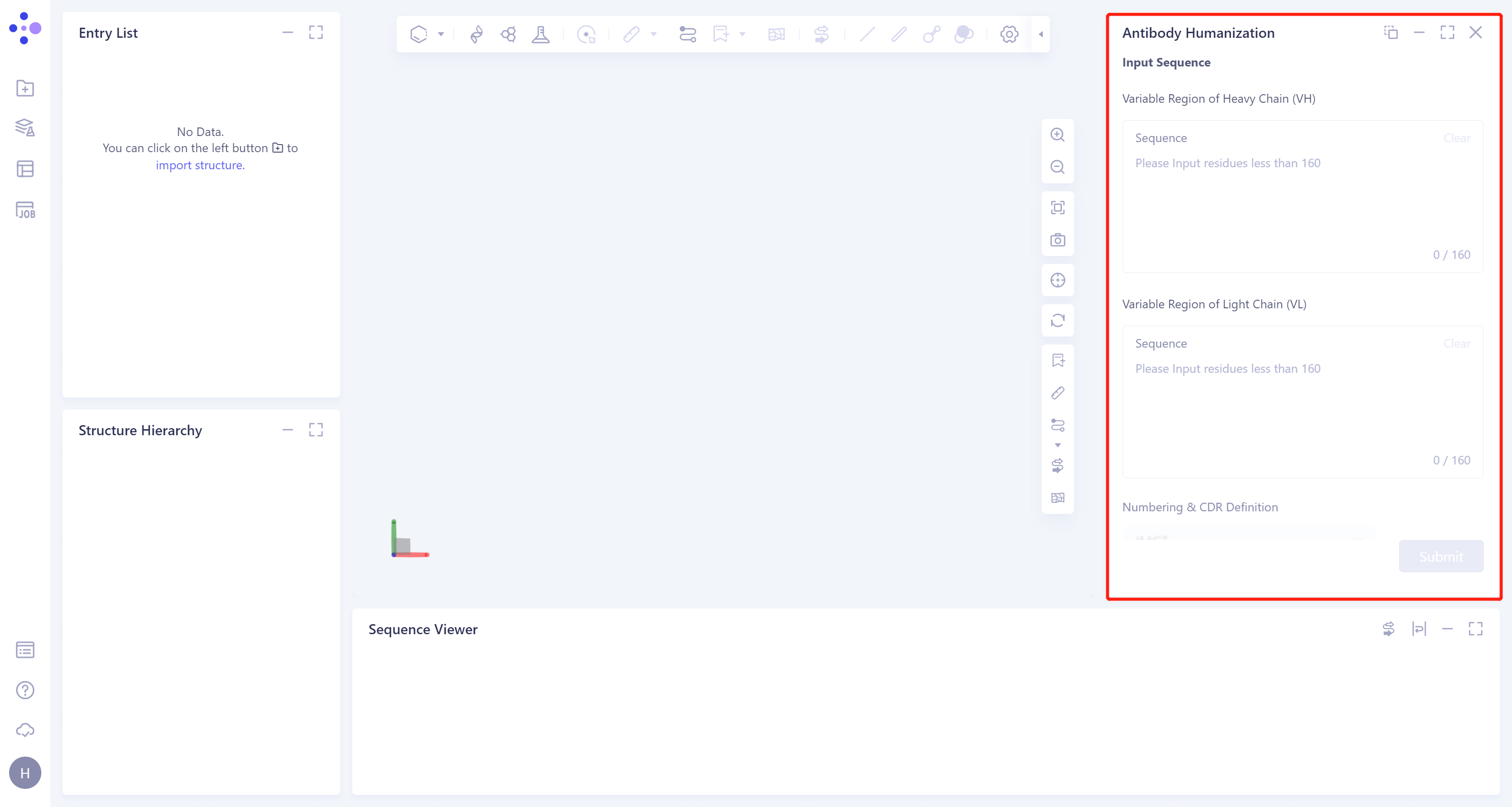
1.2 Operation
-
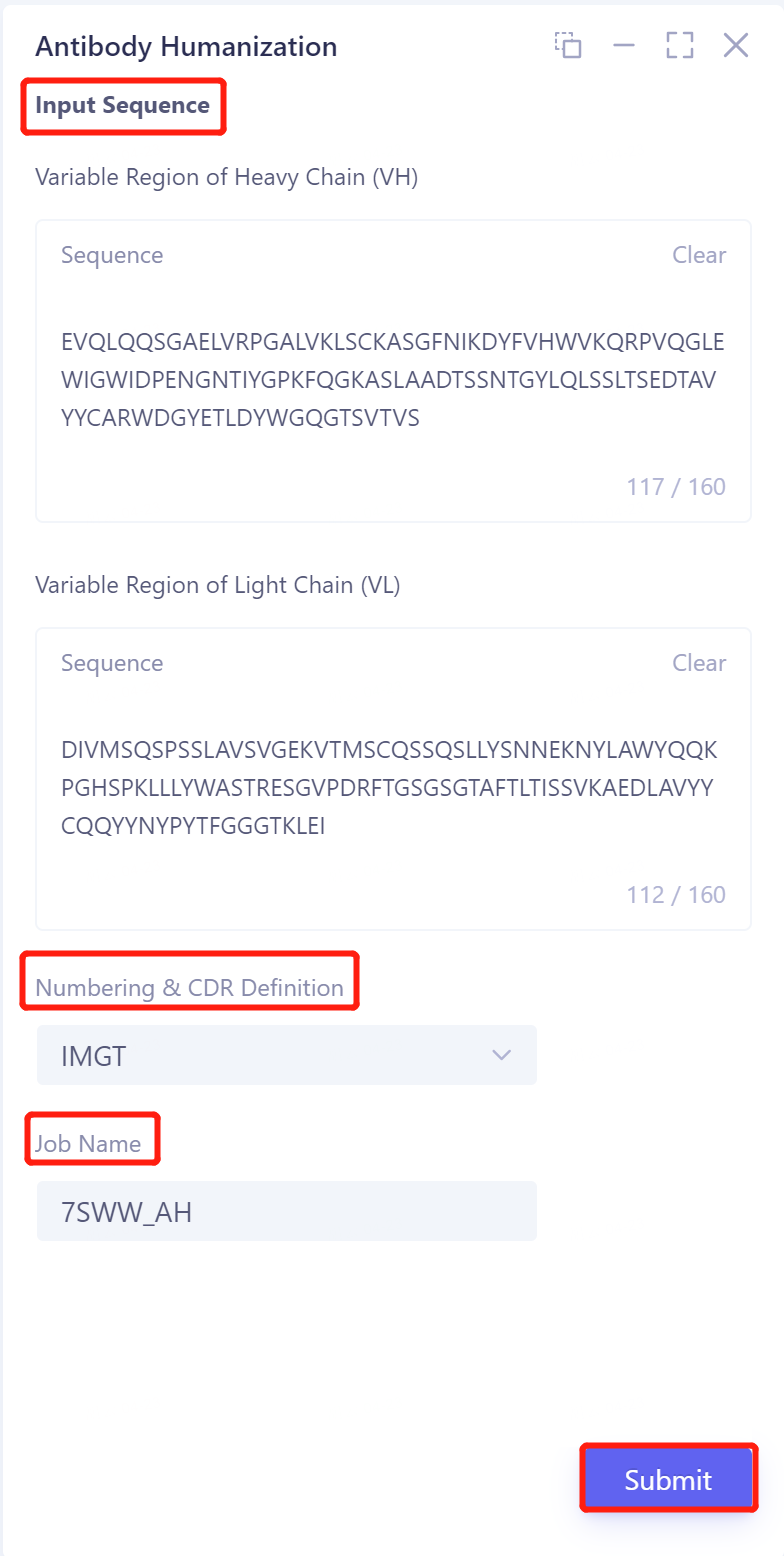
Input Sequence: Input the sequence of the variable region of the antibody
-
Enter the variable region sequence of the antibody heavy chain in the sequence box under Variable Region of Heavy Chain (VH);
-
Enter the variable region sequence of the antibody light chain in the sequence box under Variable Region of Light Chain (VL).
-
-
Numbering & CDR Definition: here select 'IMGT'.
-
Name the task at 'Job Name'.
-
Click 'Submit' to submit the task.
2. Analysis of results
2.1 Entrance
-
General menu bar on the left: Job → Find the required task.
-
Select the task to be viewed and click 'Show' in the Operation column to display the result of the task, as shown in the figure below.
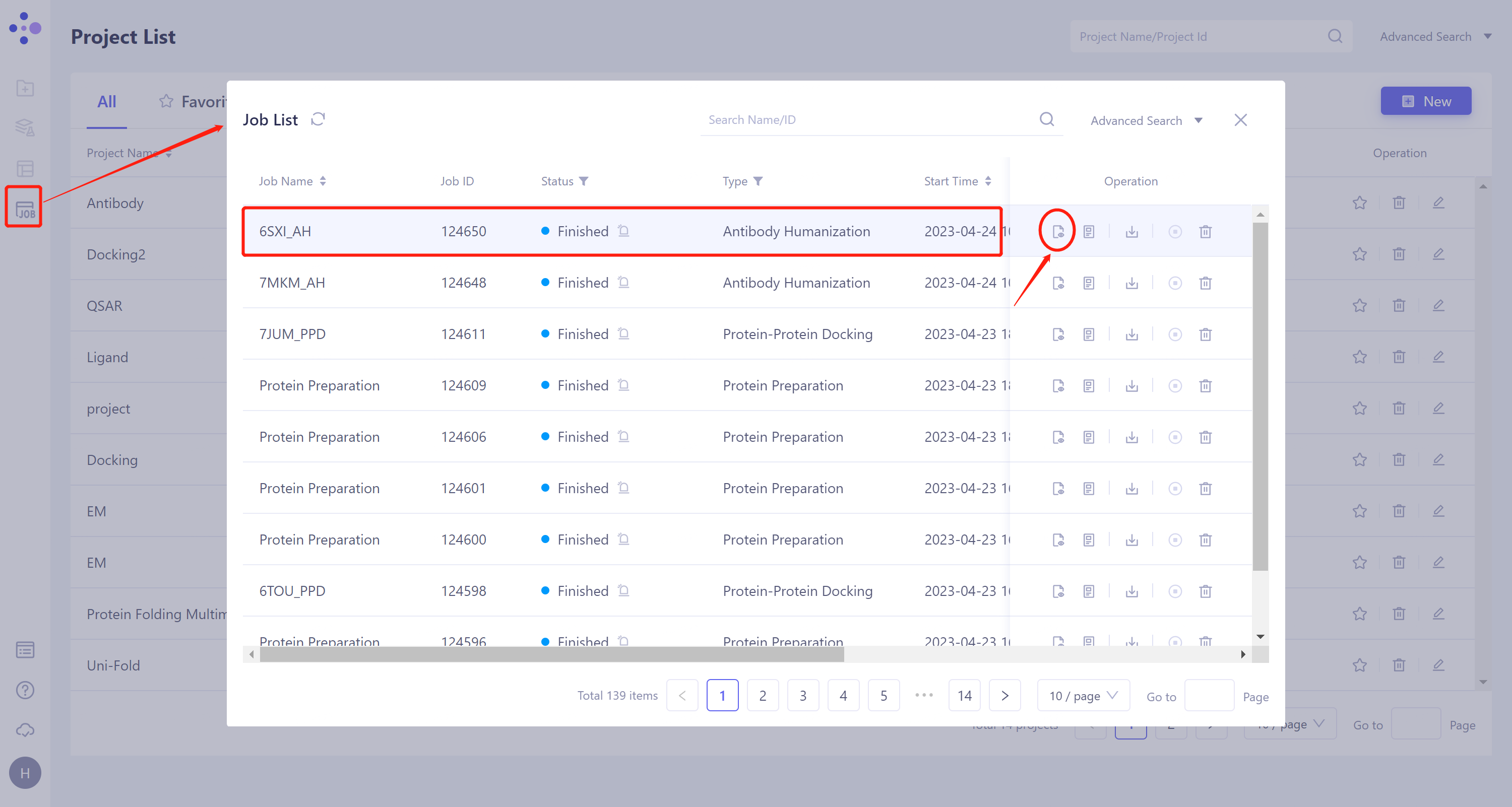 | 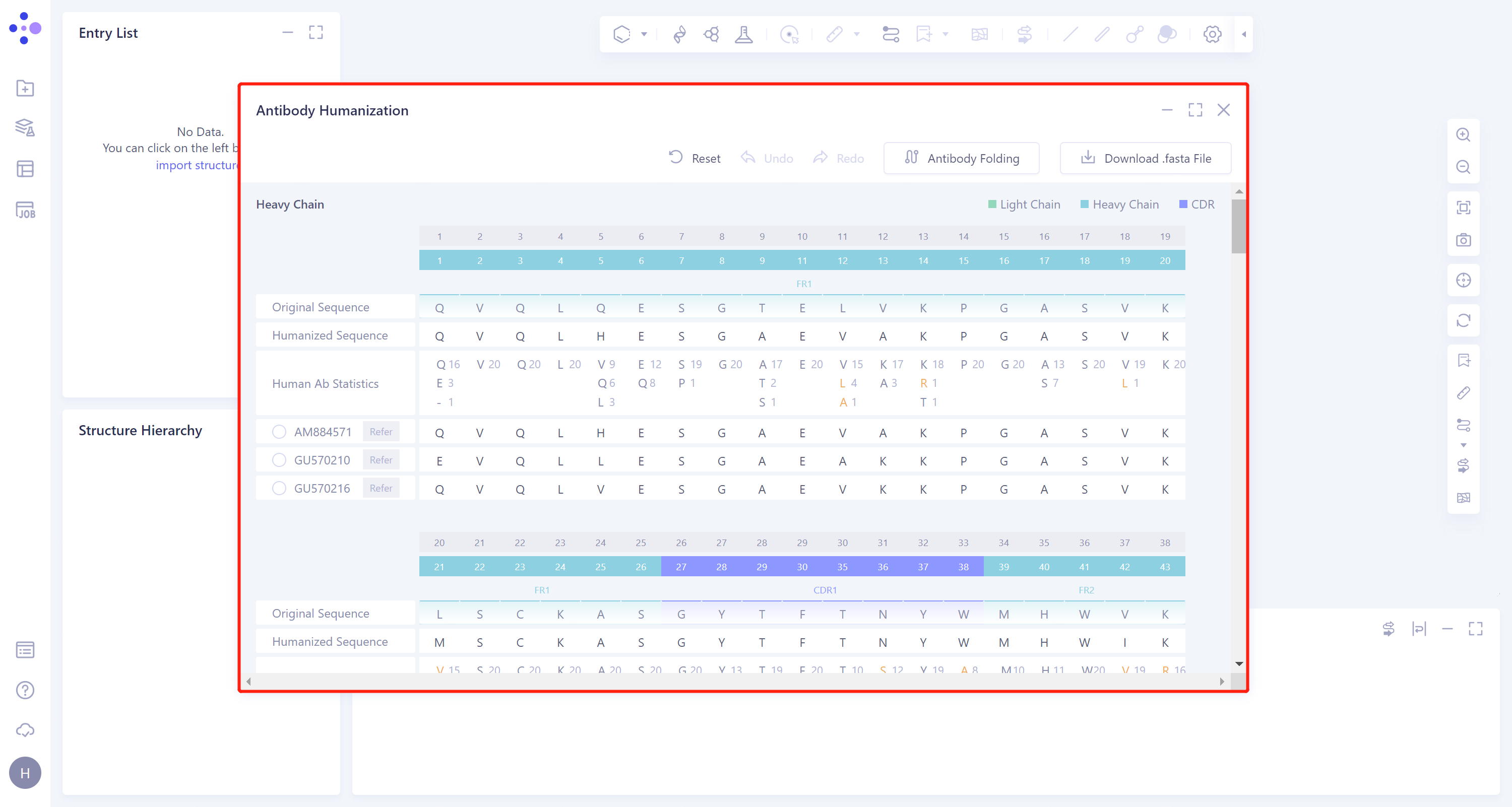 |
2.Description of results and operation of antibody mutation
- The heavy chain of the antibody sequence is marked in light blue, the light chain in light green, and the CDR region in blue-violet.
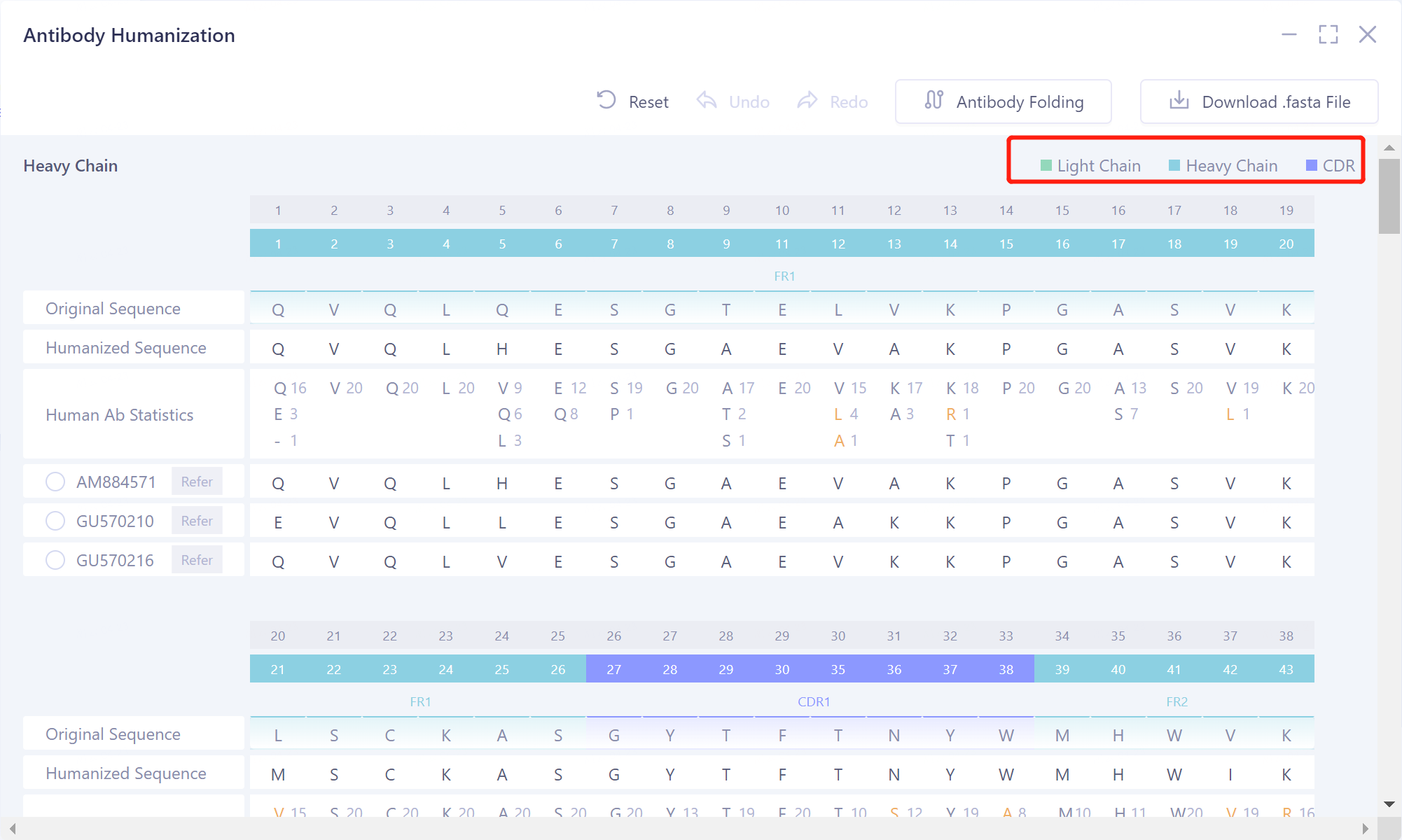 | 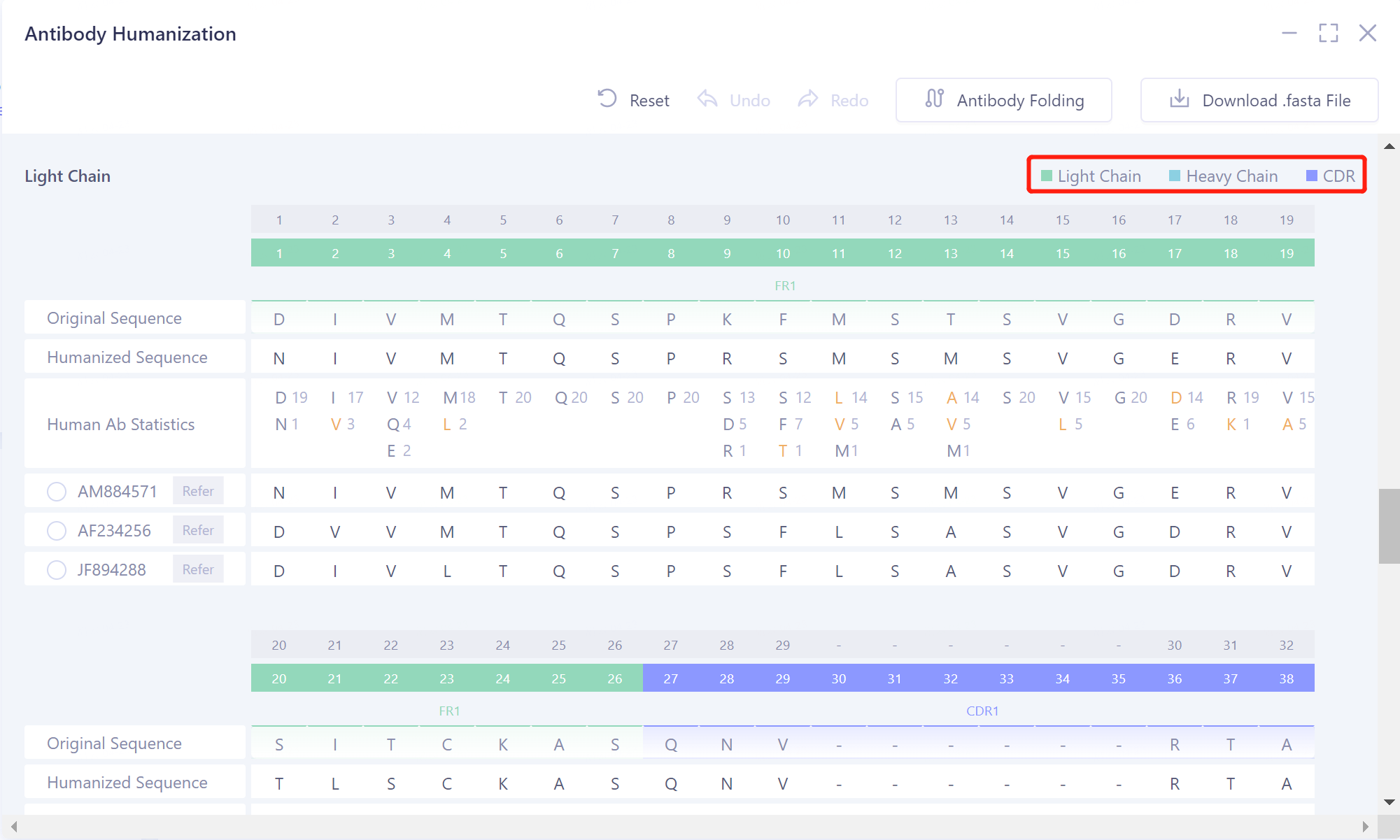 |
-
The Original Sequence line shows the amino acid sequence of the original input antibody, and the Humanized Sequence line shows the humanized sequence, which is modified from the original antibody sequence:
-
To construct a recombinant sequence, select 'AM884571', and use the framework region of this sequence to construct the framework region of the humanized antibody, while retaining the CDR region of the original antibody sequence to ensure the affinity between the antibody and the antigen. The specific operation is as follows: Click 'AM884571', and select 'Keep original CDR' in the pop-up 'Confirm' window. Click 'OK'.
-
According to the statistical data of the 20 most matched human sequences displayed in the Human Ab Statistics line, the amino acids marked in red are the amino acid residues recommended for mutation (amino acid residues of the same type), and the above recombinant sequences are further mutated to mutate the 20th M to V. Specific steps: In the Humanized Sequence line, click the amino acid residue M at the 20th position, select V from the expanded 20 selectable amino acid residues, and the manually adjusted amino acid will be marked in red.
-
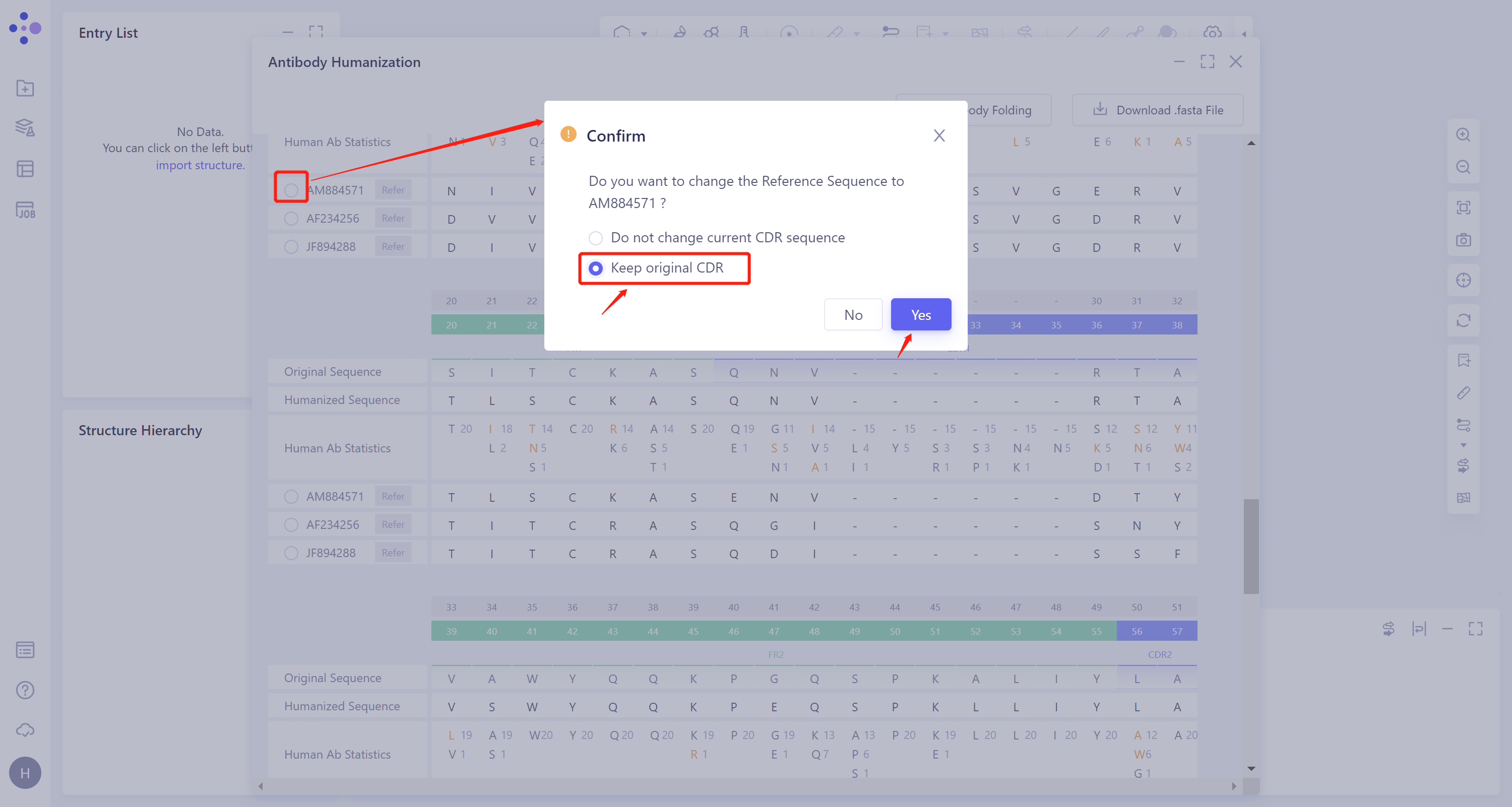
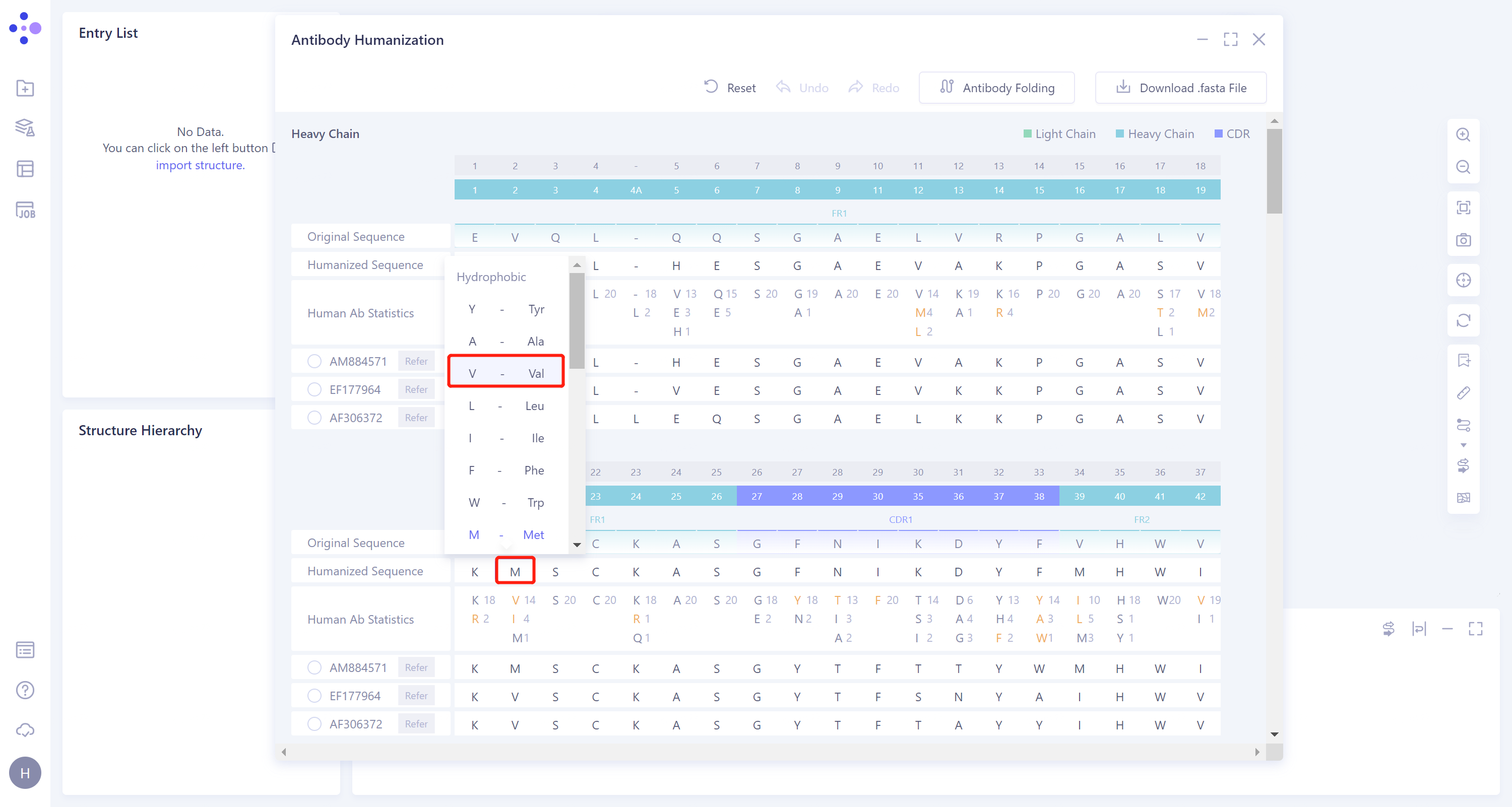
-
Download .Fasta File: Download the humanized antibody sequence (shown in the Humanized Sequence line) and save the file in '.fasta' format.
-
For the Antibody Folding function, see the corresponding section in Tutorials.