Antibody Folding
Introduction
The monoclonal antibody pembrolizumab is an FDA-approved PD-1 blocker used to treat several types of cancer. 5GGS is the complex crystal structure of PD-1 and pembrolizumab.
In this tutorial, you will learn the structure prediction of antibodies and evaluate the accuracy of antibody structure prediction through protein structure alignment. Antibody structure prediction calls the Antibody Folding module in Hermite. Protein alignment calls the Protein Alignment module in Hermite.
The data used in this tutorial is as follows:
5GGS_1|Chains A, C|heavy chain|Homo sapiens (9606)QVQLVQSGVEVKKPGASVKVSCKASGYTFTNYYMYWVRQAPGQGLEWMGGINPSNGGTNFNEKFKNRVTLTTDSSTTTAYMELKSLQFDDTAVYYCARRDYRFDMGFDYWGQGTTVTVSS
5GGS_2|Chains B, D|light chain|Homo sapiens (9606)EIVLTQSPATLSLSPGERATLSCRASKGVSTSGYSYLHWYQQKPGQAPRLLIYLASYLESGVPARFSGSGSGTDFTLTISSLEPEDFAVYYCQHSRDLPLTFGGGTKVEIK
1. Antibody Folding
1.1 Entrance
- The left general menu bar Function → Biomacromolecule → Antibody Folding.
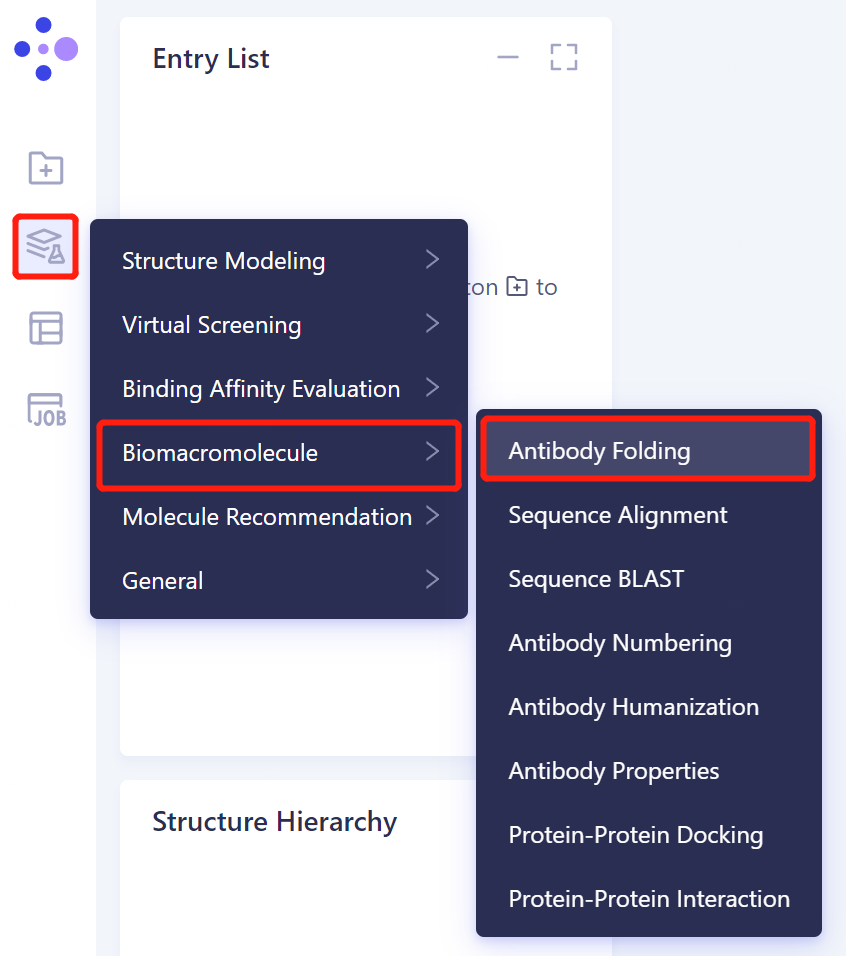
- The right side of the interface appears:
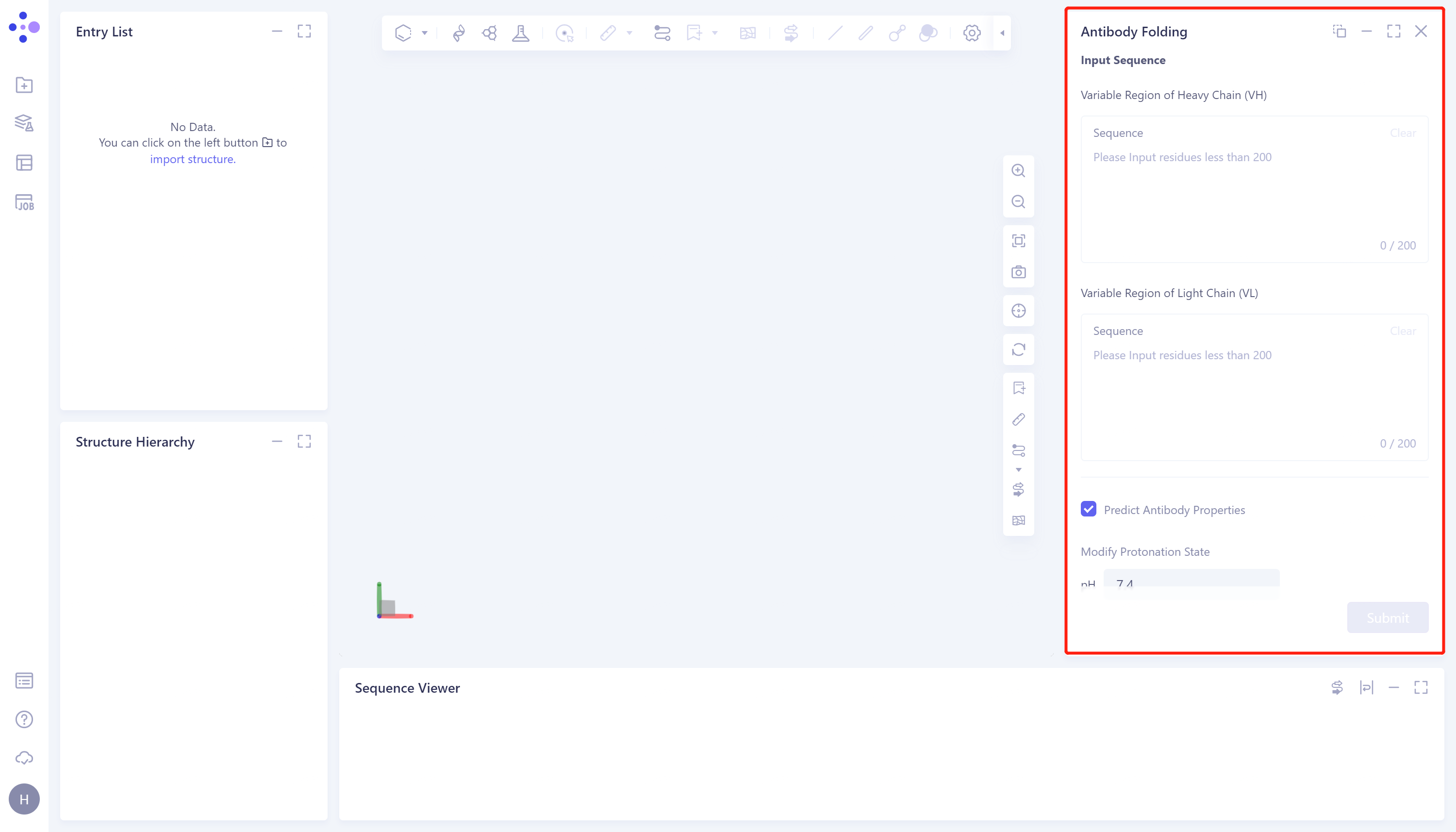
1.2 Sequence input and parameter adjustment
- Input the heavy chain and light chain of the antibody in the 'Input Sequence' → Check the 'Predict Antibody properties' to predict the properties of the antibody → Adjust the pH to 7.4 to make the protein reach the protonated state at this pH → Name the task in the Job Name as 'Case-Antibody Fold-5GGS' → Click 'Submit' to submit the task.
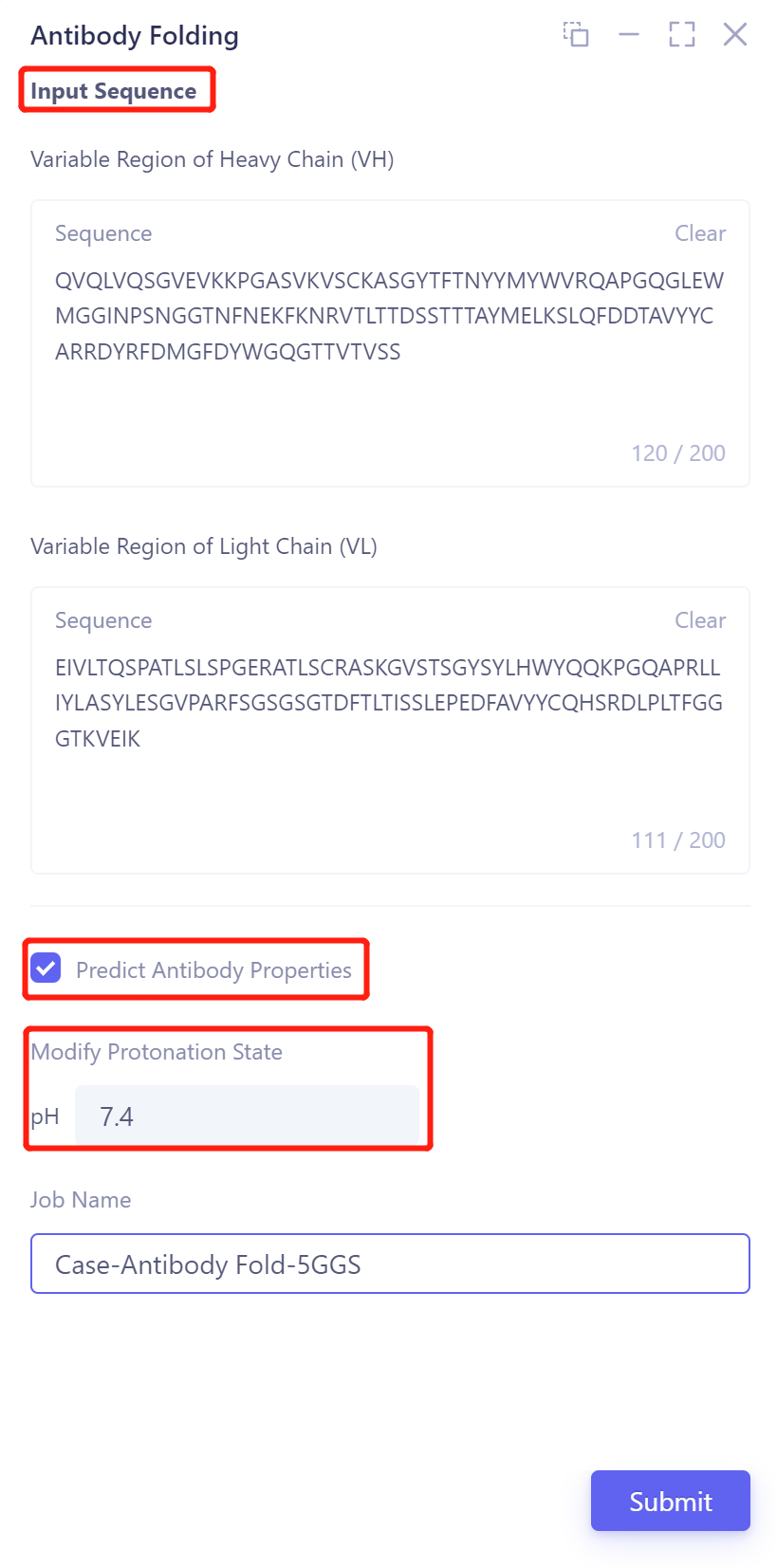
1.3 Display of prediction results
- Click 'Job' in the left general menu bar → find the 'Case-Antibody Fold-5GGS' task in the pop-up 'Job List' interface → click 'Show'.
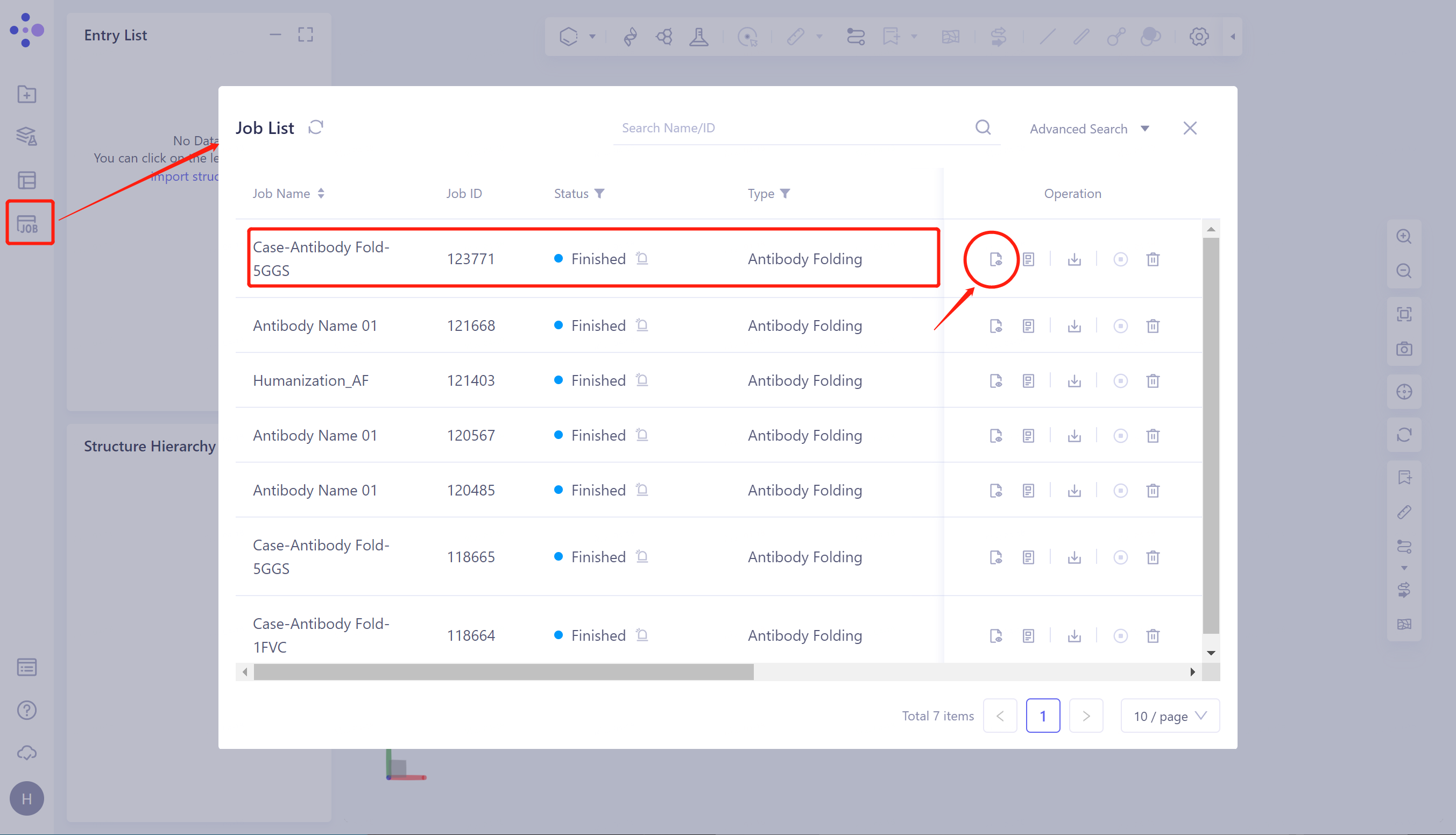
1.3.1 Display of antibody structure prediction results
- Protein prediction results are displayed in the 3D Workspace interface. Antibody structure regions are marked with different colors, blue is heavy chain, green is light chain, purple is non-H3 CDR region, and orange is CDR-H3.
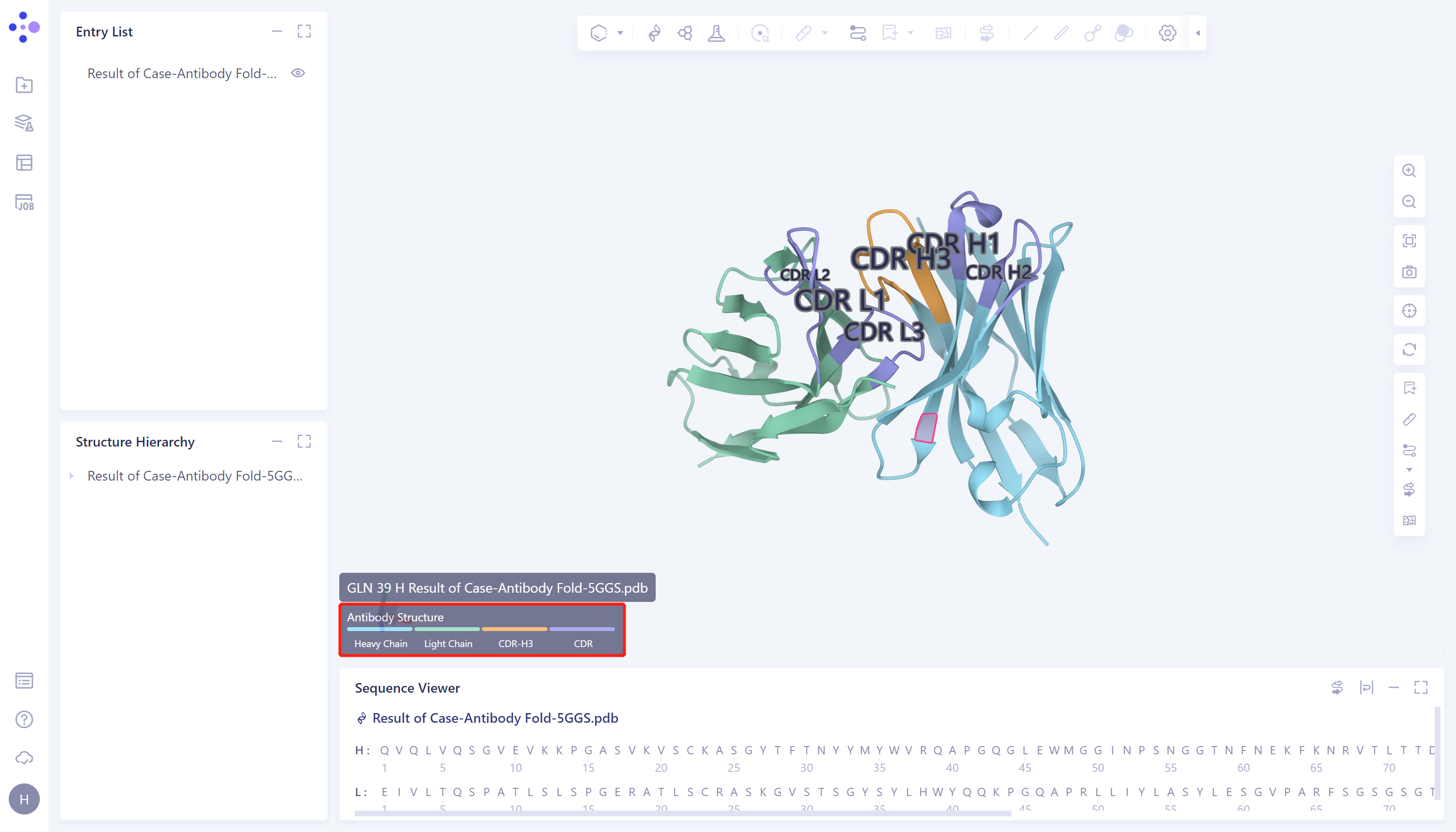
1.3.2 Display of Antibody Property Prediction Results
- The result of antibody property prediction is shown in the figure, which shows that the viscosity of the antibody is appropriate, the clearance risk is low, the aggregation is acceptable, and the PSH, PPC PNC and SFvCSP Score of the antibody are also distributed in the statistical interval, which conforms to the TAP principle. In conclusion, the biophysical properties of the antibody are appropriate, indicating that Antibody Properties can accurately predict the properties of 5GGS antibodies.

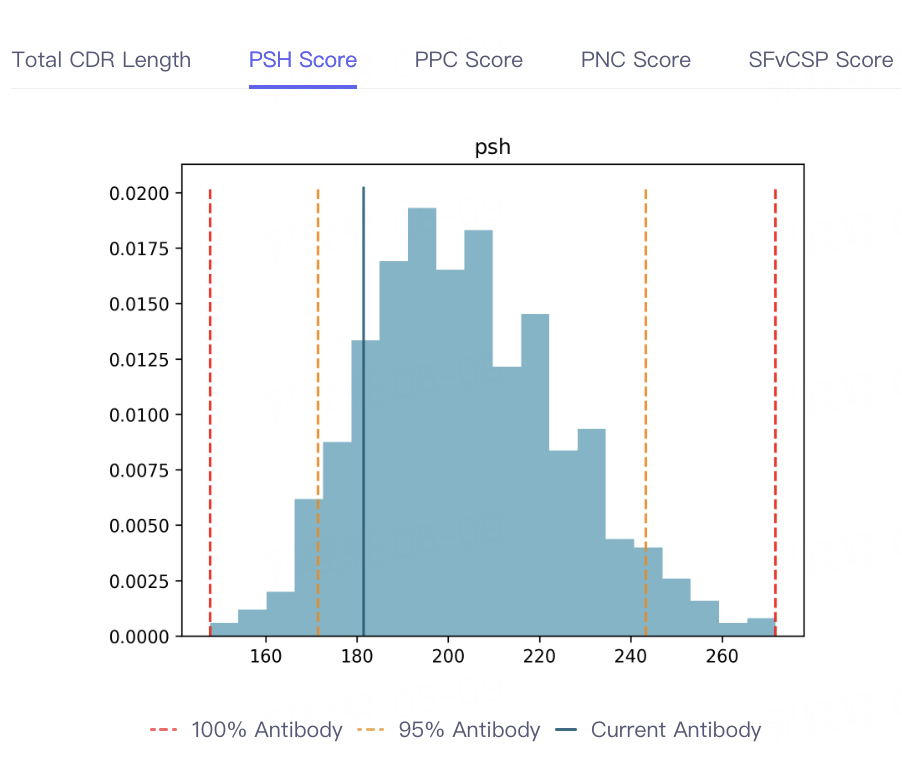 | 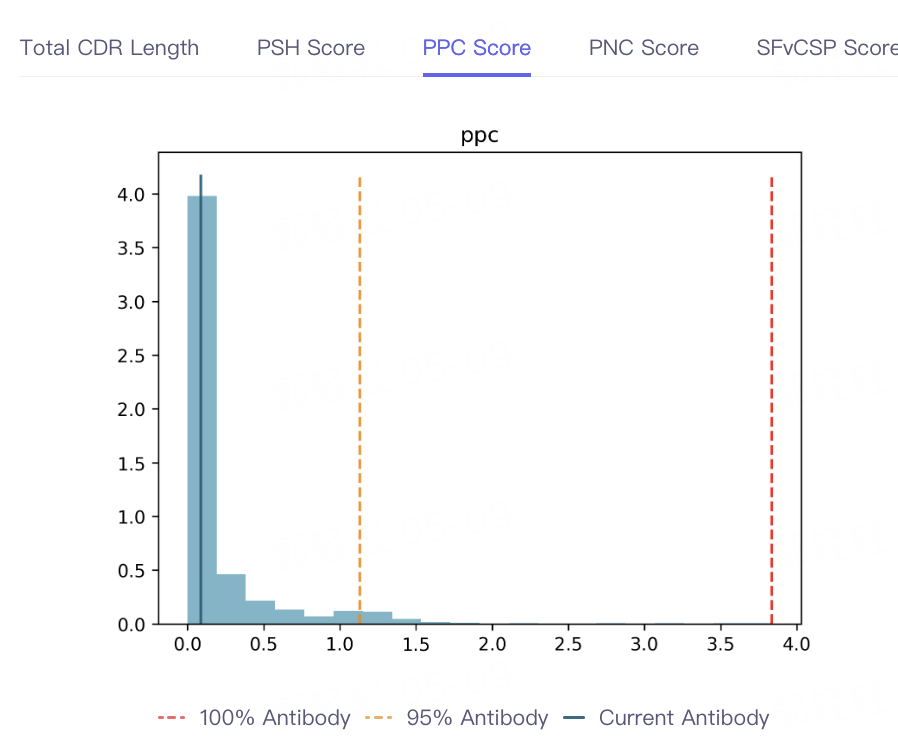 | 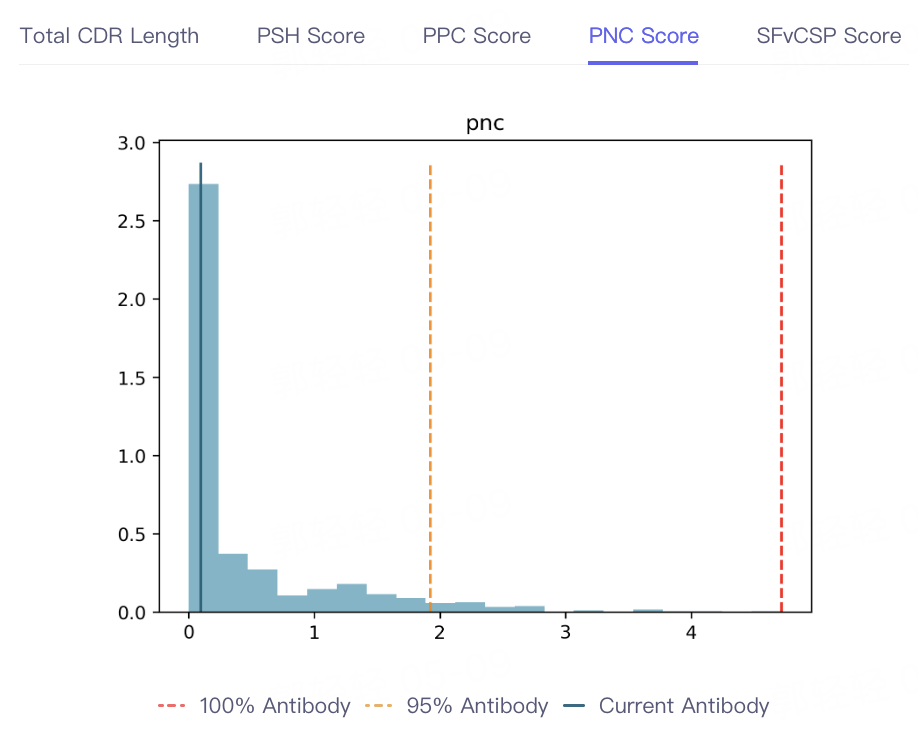 | 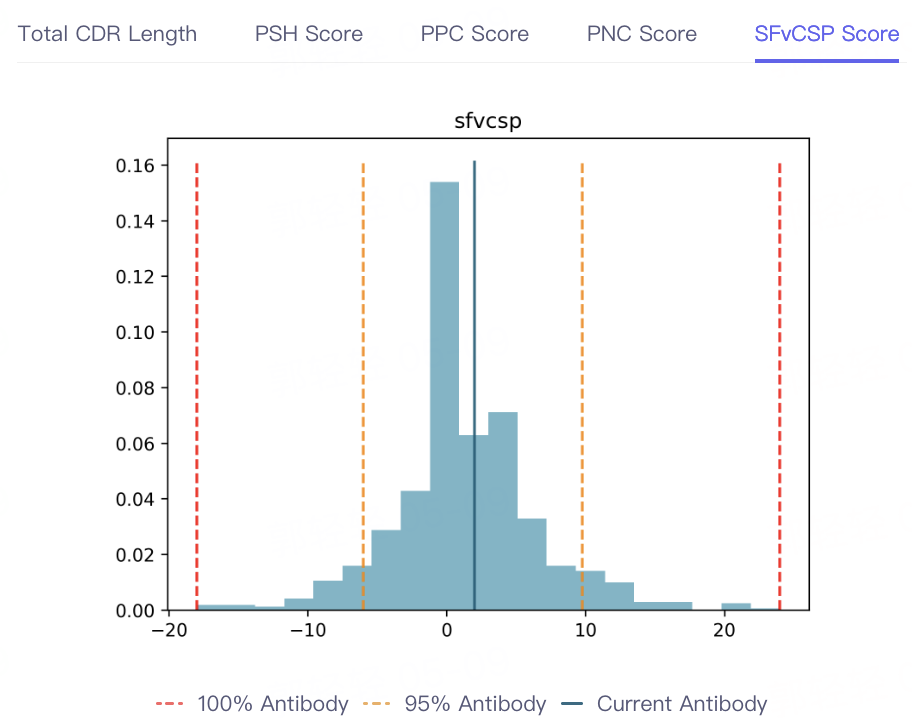 |
2. Protein Alignment
- In order to test the accuracy of antibody structure prediction, the Protein Alignment module was called to compare the predicted protein structure with the 5GGS structure.
2.1 Import protein structures into the 3D Workspace window
-
The left general menu bar 'File' → 'Get PDB'.
-
Enter PDB ID: 5GGS in the pop-up window and click 'Import' to import the protein structure.
 |  |
2.2 Invoke Protein Alignment module
2.2.1 Inlet
- In the same project, click 'Function' → 'General' → 'Protein Alignment'.
2.2.2 Introduction of reference protein structures
- In the 'Protein Alignment' window on the right side of the interface, first import the reference protein structure: click 'Select from 3D Workspace', pop up the 'Select Structure' window → Select Chain A (antibody heavy chain) and Chain B (antibody light chain) of 5GGS protein in the Structure Hierarchy → Click 'OK' in the 'Select Structure' window.
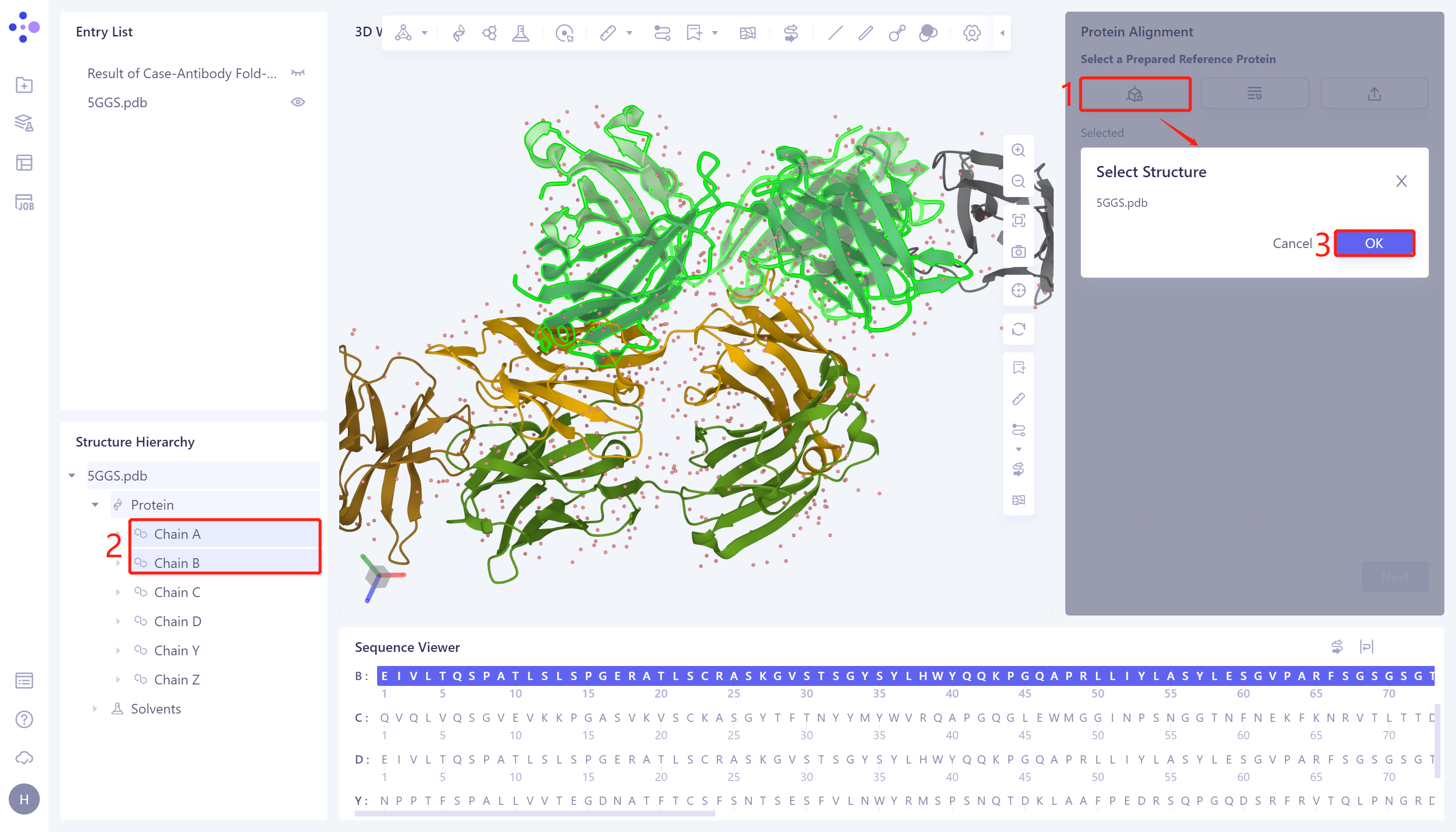
2.2.3 Introduction of protein structures to be compared
- Click 'Next' to enter the selection window of protein to be compared, and import the protein structure to be compared: click 'Select from 3D Workspace', the 'Select Structure window pops up' → Select the predicted 5GGS protein structure 'Case-Antibody Fold-5GGS' in the Structure Hierarchy → Click 'OK'.
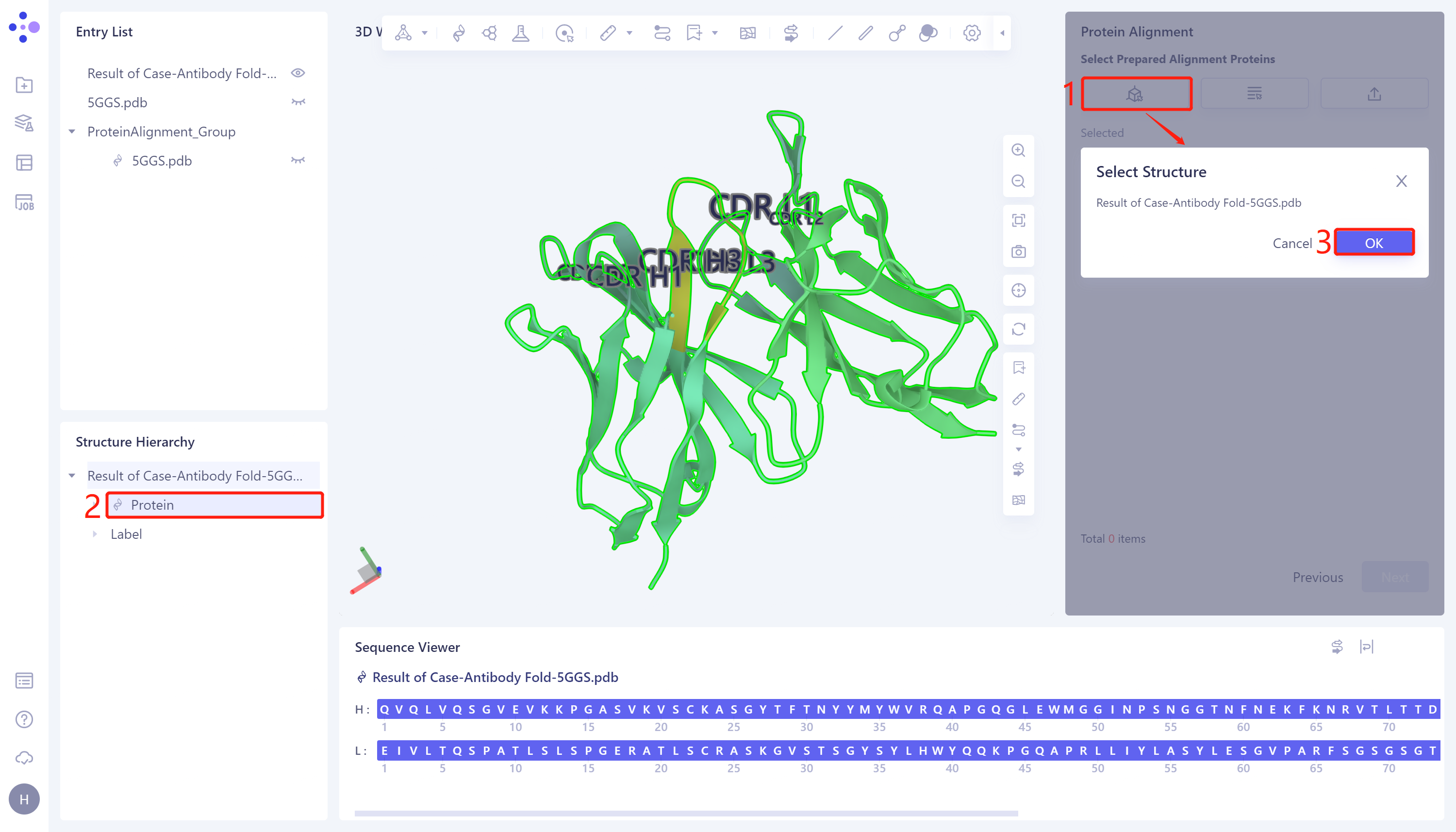
2.2.4 Comparison parameter setting
- Click 'Next' to enter the parameter setting page. Alignment Atoms selects 'Cα' to perform alignment based on the α-carbon atoms between proteins. Since there is no special comparison requirement, the parameter setting of Select Binding Site is skipped. Name the task as '5GGS_PA' at the 'Job Name', and click 'Submit' to submit the task.
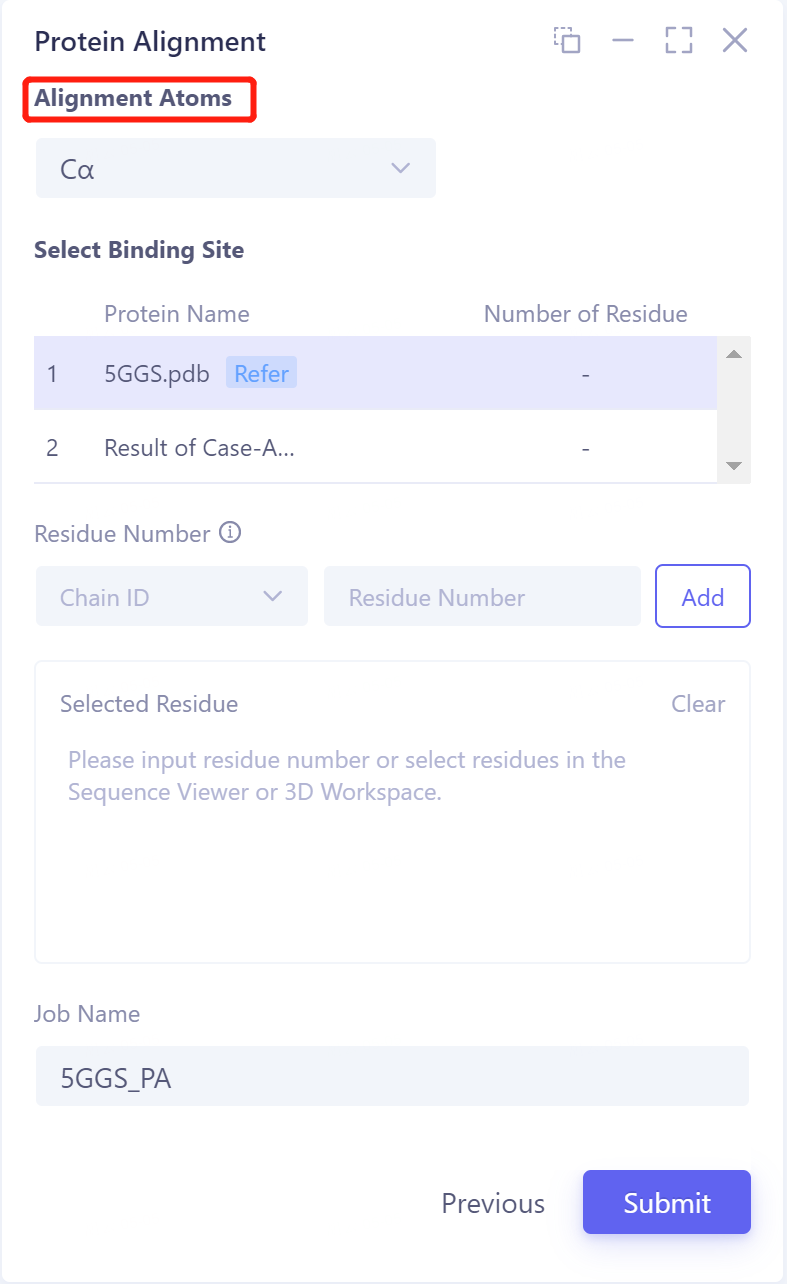
2.3 Analysis of results
2.3.1 Presentation of results
- General menu bar on the left: Menu Job → Find the task '5GGS_PA' in the pop-up Job List window → Click 'Show' under Operation to display the task.
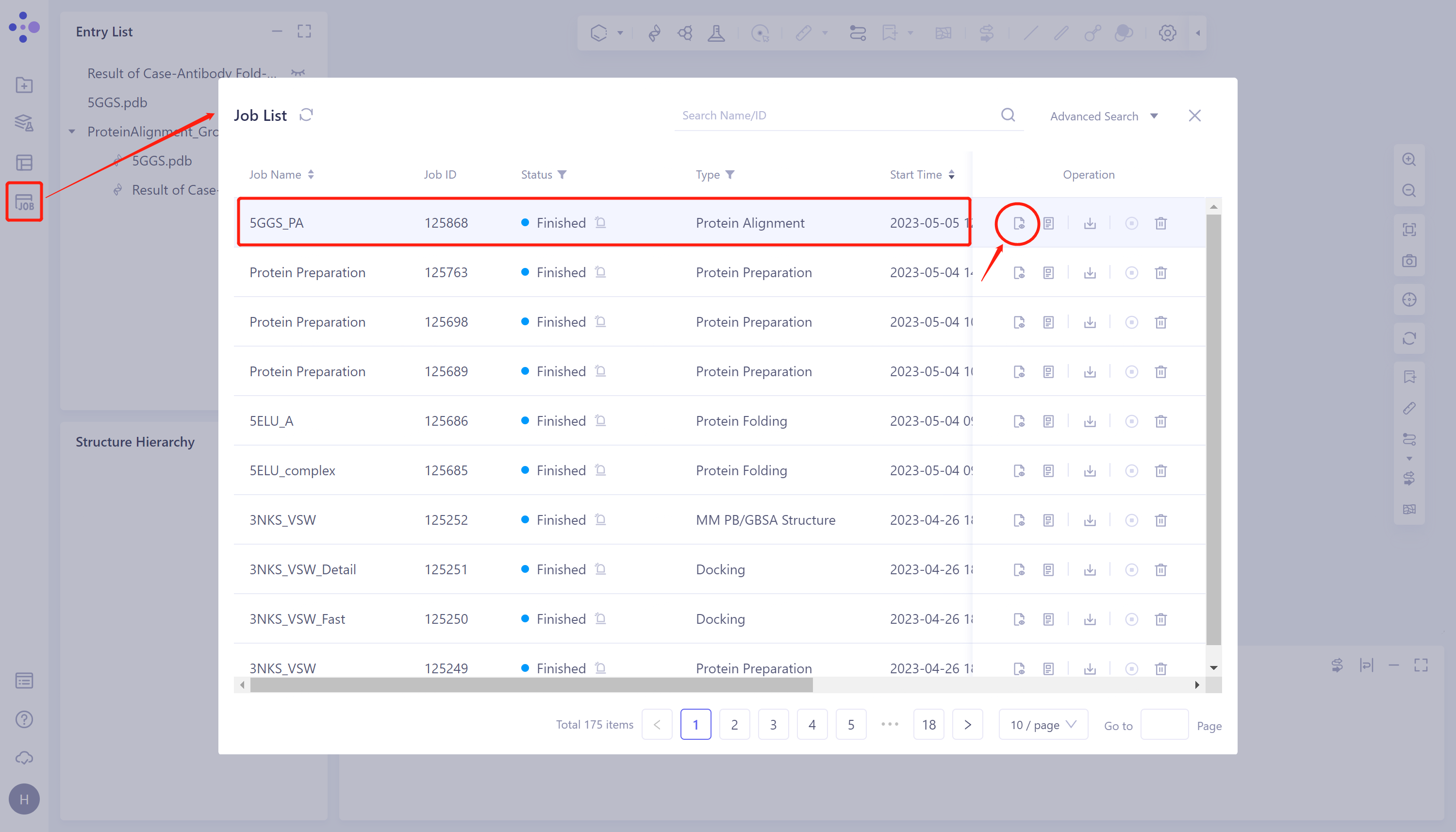
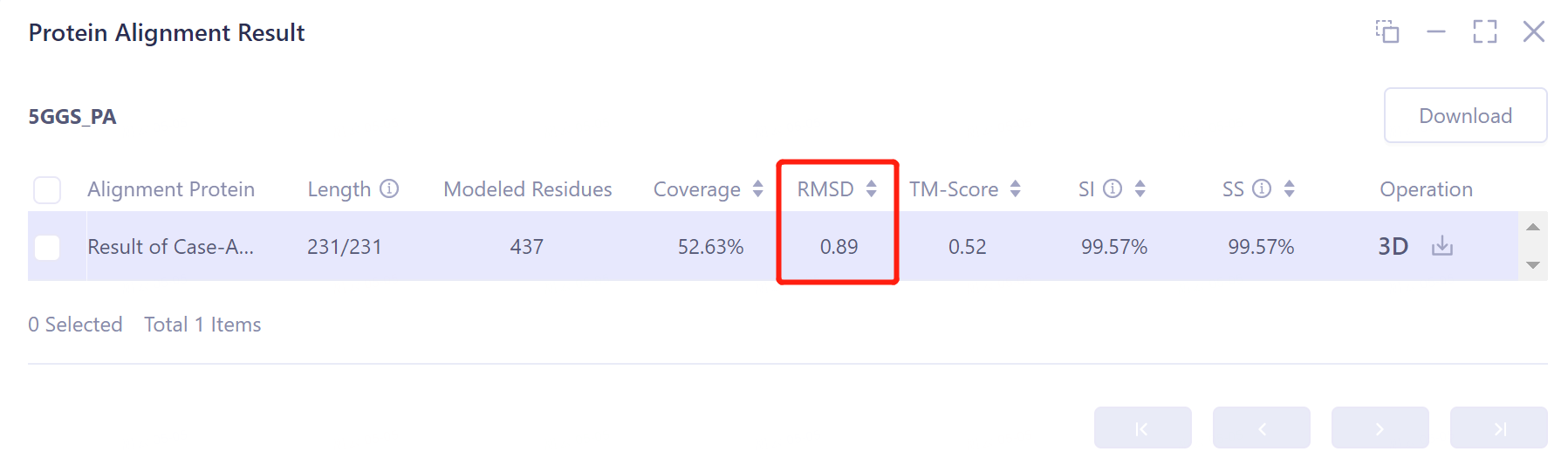
- The overlay image result of protein structure is displayed in the 3D Works pace window, and the Protein Alignment Result window appears on the right side of the interface to display the protein alignment result.
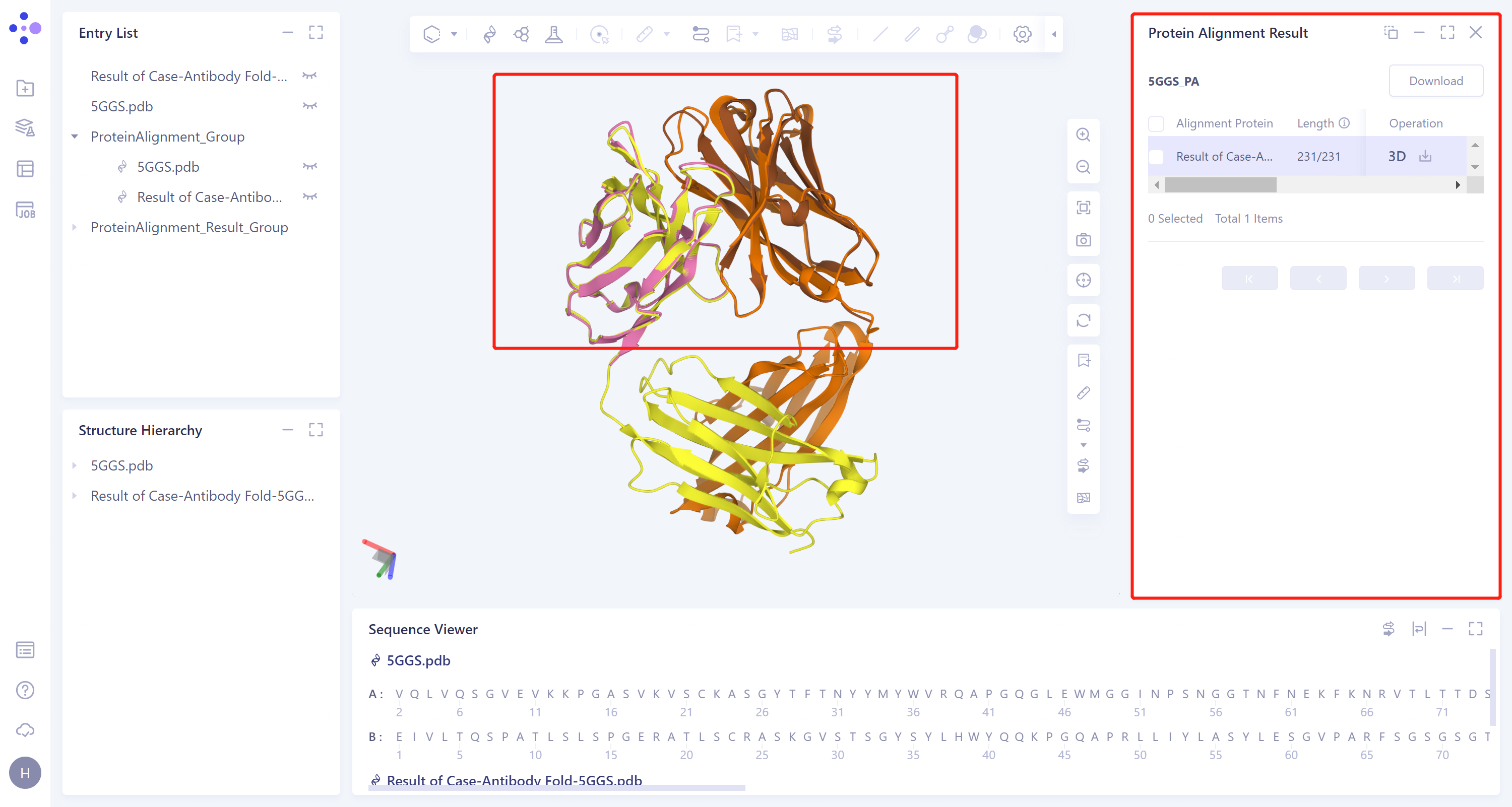
2.3.2 Explanation of results
- After protein structure comparison, it was found that the RMSD value of Antibody Folding predicted the structure of antibody 5GGS and 5GGS structure was 0.89 Å, indicating that Antibody Folding could well predict the protein structure of 5GGS antibody.