FEP Calculation
Introduction
With the development of free energy perturbation (Free Energy Perturbation, FEP) theory and algorithm, as well as the improvement of computer performance, the accuracy and practicability of FEP calculation have been greatly improved. Using FEP calculation to evaluate the difference of Affinity between a series of structurally similar candidate drug molecules (Ligands) and the target Protein can quickly verify the ideas of drug designers, improve the efficiency of research and development, and reduce the cost of experiments, which has been widely concerned and applied in the field of drug research and development.
Uni-FEP is a complete FEP solution developed by Deep Potential Technology. It combines free energy perturbation theory, molecular dynamics, enhanced sampling algorithm and high-performance computing, and can efficiently evaluate the binding affinity between receptor and ligand with chemical accuracy. It is suitable for the optimization stage of lead compounds.
KRAS oncogene is the first human tumor gene to be discovered and is one of the most commonly mutated genes in human tumors. Missense mutations in KRAS reduce GTP hydrolysis and alter the homeostatic balance between GDP and GTP. Mutations at position 12 are the most common mutations in KRAS, and G12D, G12V and G13D are the most common mutations. KRAS protein has a smooth surface, nearly spherical spatial structure, and lacks deep hydrophobic pockets, which hinders the recognition of high-affinity inhibitors, so it has been considered as a typical target of refractory drugs in the industry [1]. Human beings have been devoted to the research of KRAS for more than 40 years. Until recent years, drugs covalently targeting KRAS-G12C mutation subtype have been launched [2]. However, drugs targeting other KRAS mutations, such as KRAS-G12D, are still difficult to develop. Therefore, the development of effective KRAS-G12D inhibitors is expected to become a new strategy for cancer treatment.
In this tutorial, you will learn to use the Uni-FEP module of Hermite ® to quickly verify the binding ability of seven compound molecules with KRAS-G12D mutants that are already active in patents and literature [3]. The binding free energy predicted by Uni-FEP is highly correlated with the experimental affinity.
Protein structure files used in this tutorial:
Ligand structure files used in this tutorial:
1. Create a project and import the structure
1.1 Log in to the system
- Login address: https://hermite.dp.tech

1.2 Create the project
- Once in the system, create a new project KRAS FEP ”.
1.3 Import Protein Structure
- Left general menu bar Menu → File → Import Structure
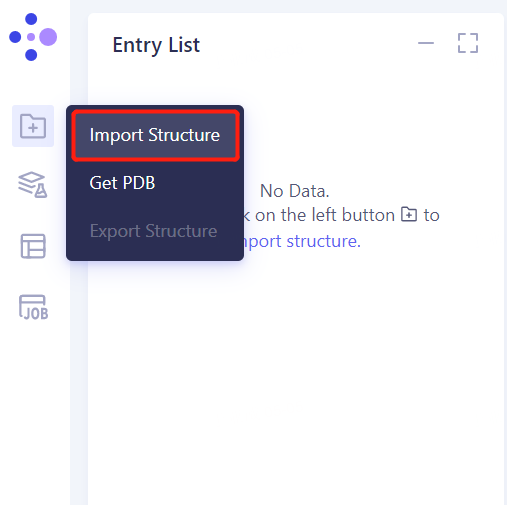
- Click Select Files, select the KRAS. PDB file, and click Import to import the protein structure
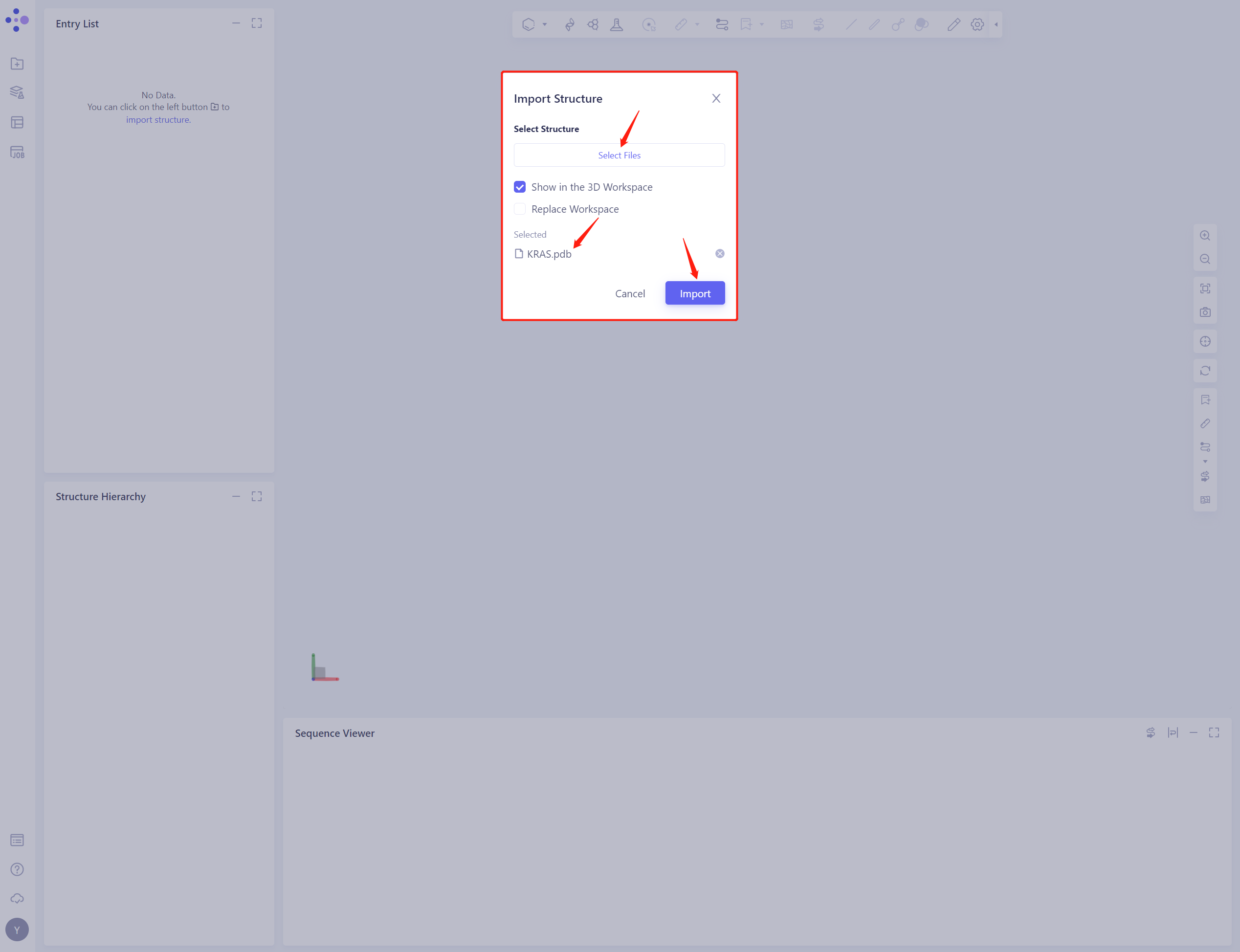
2. System preparation
2.1 Select Structure
- Left general menu bar Function → Structure Modeling → Protein Preparation
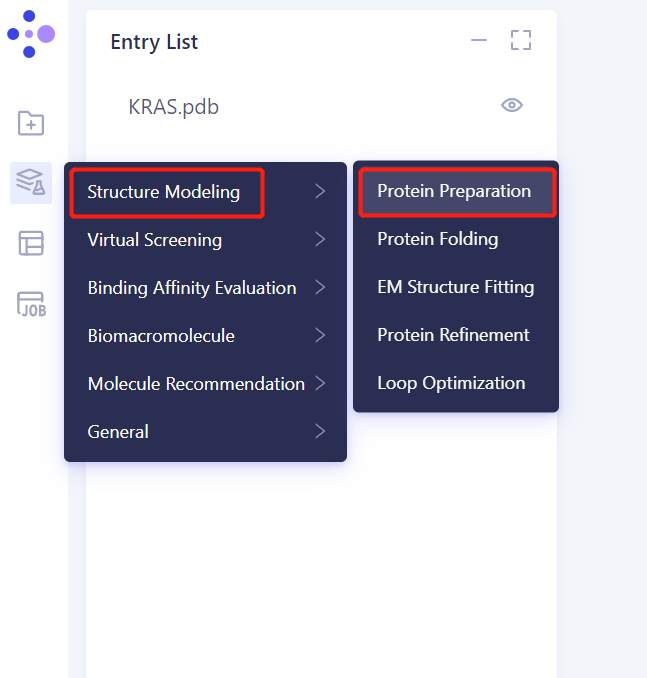
- Select Structure from 3D Works pace: Select Protein in 3D Works pace and click Ok
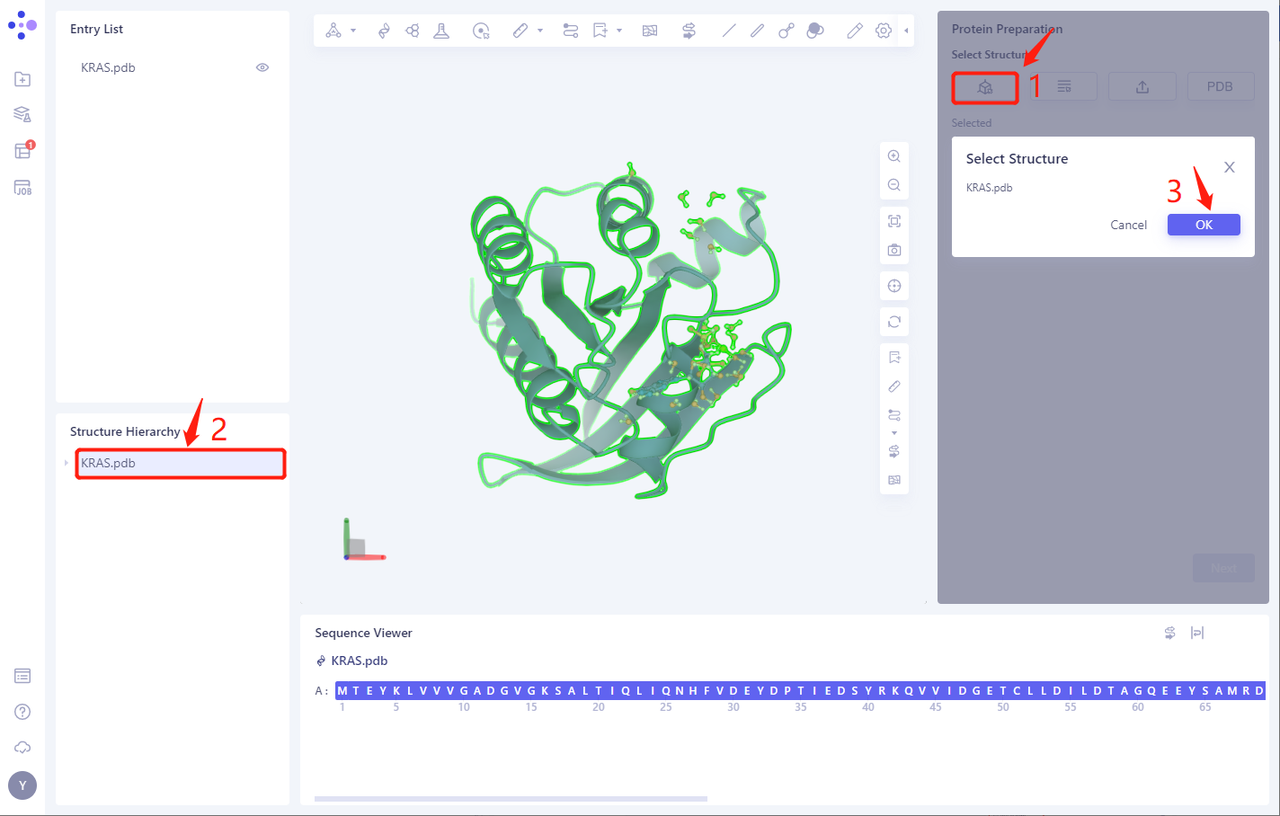
- The KRAS. PDB is loaded into the Protein Preparation parameter setting panel, and click Next
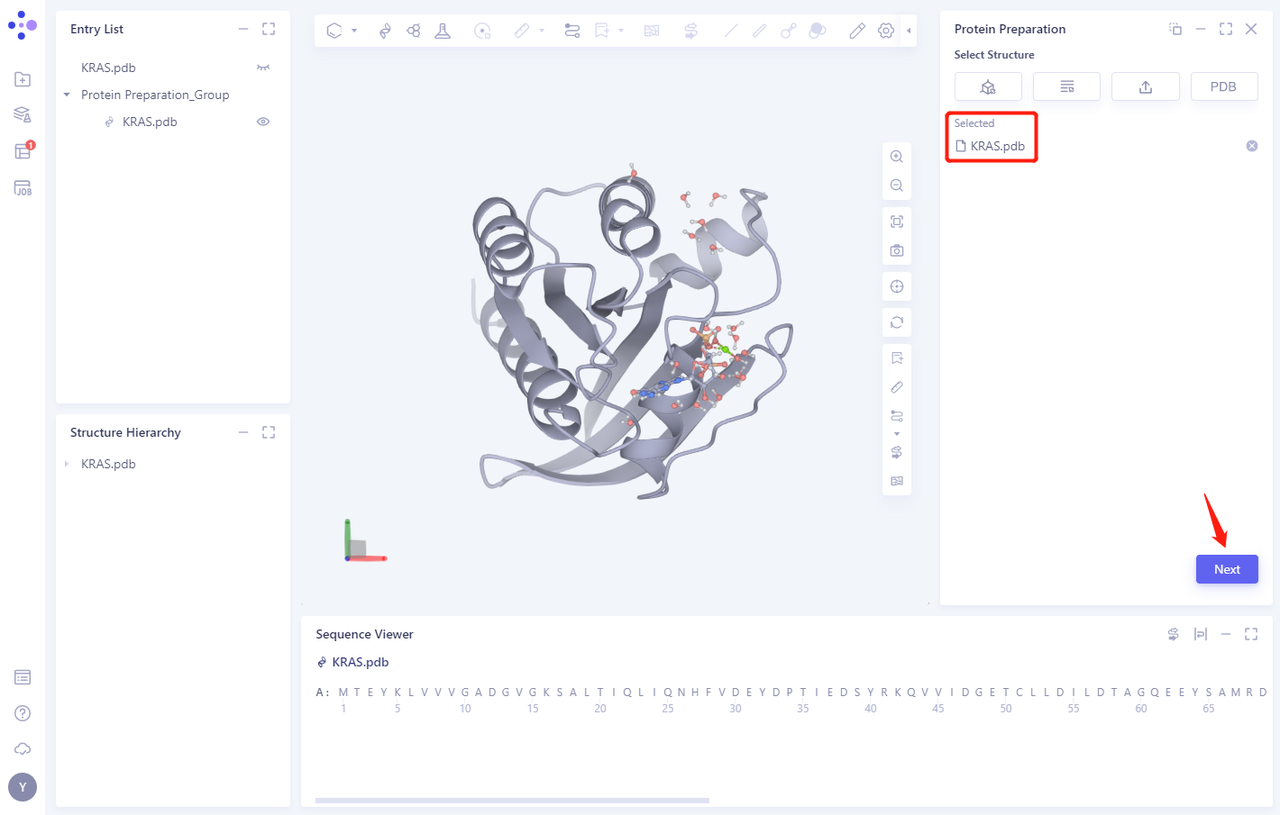
2.2 Select Polymer、Other Groups to Keep:
- Select the A-chain protein and click Next (there are no other groups in this protein structure)
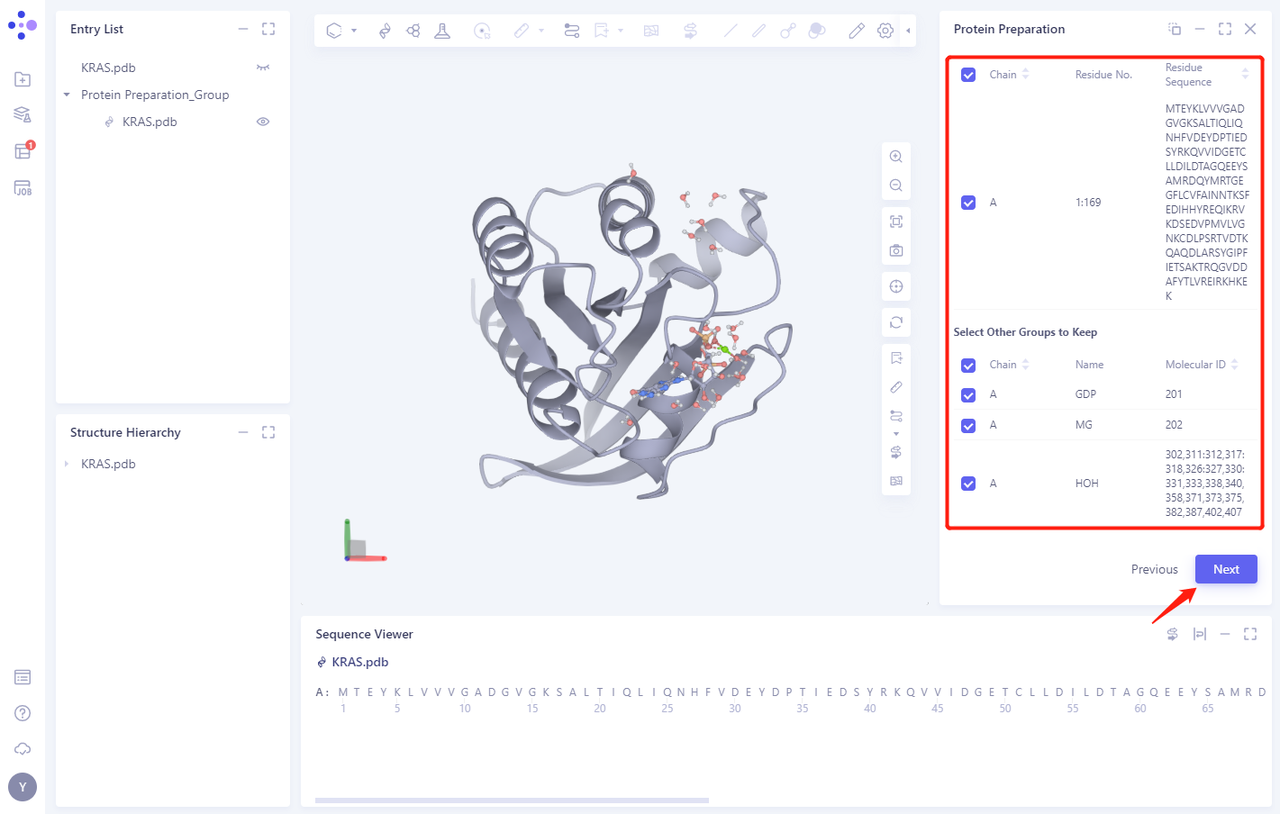
2.3 Select Missing Residues to Repair
- No Missing Residues in this case
2.4 Prepared Settings
- See Figure 2.2.5 for setting parameters. Uncheck the Energy Minimization option.
2.5 Name the Job and submit it
- Name the Job as “KRAS Protein Prepare” “and click” Submmit “to submit the job
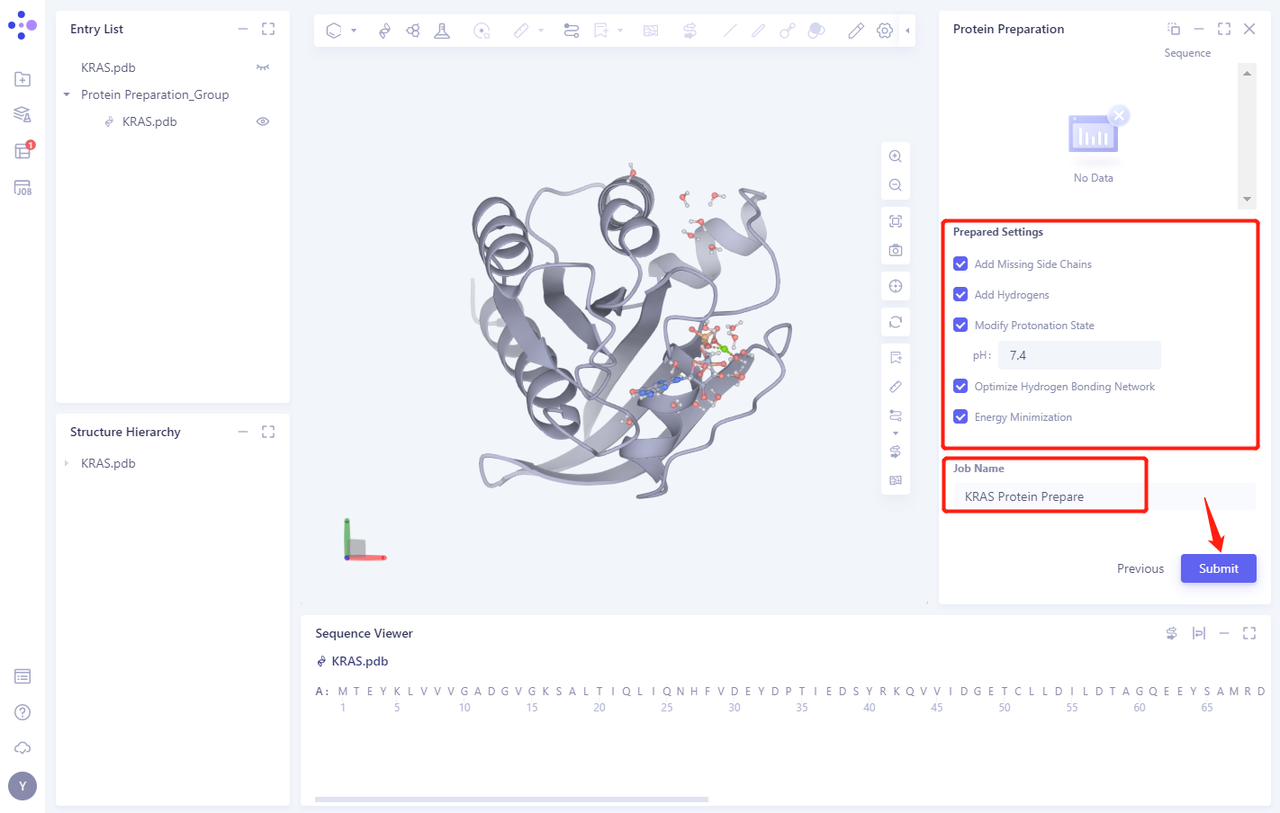
2.6 View protein preparation results
- Protein Preparation computing tasks are generally completed in a dozen seconds to a few minutes. After the task is completed, click Jobs to view the corresponding task;
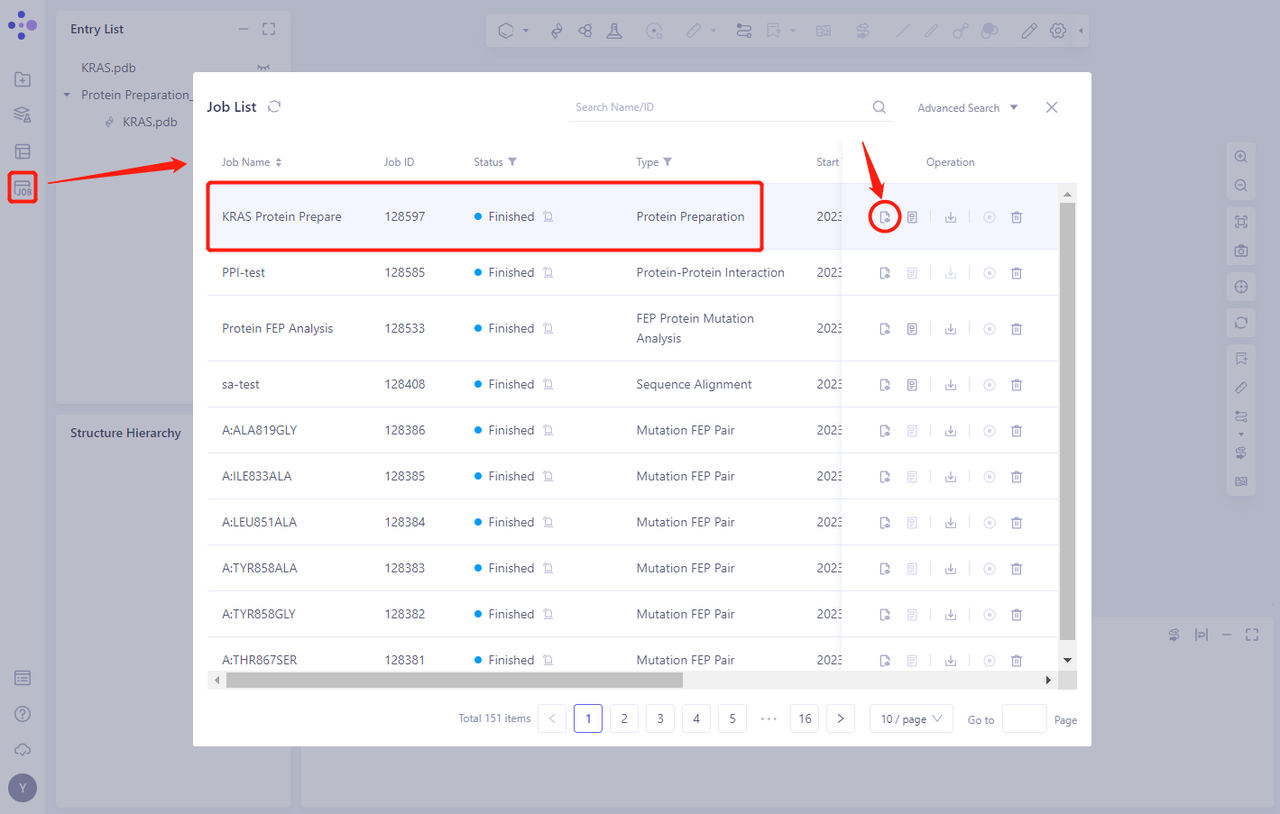
- Click the “show” button to display the prepared protein structure in 3D Workspace
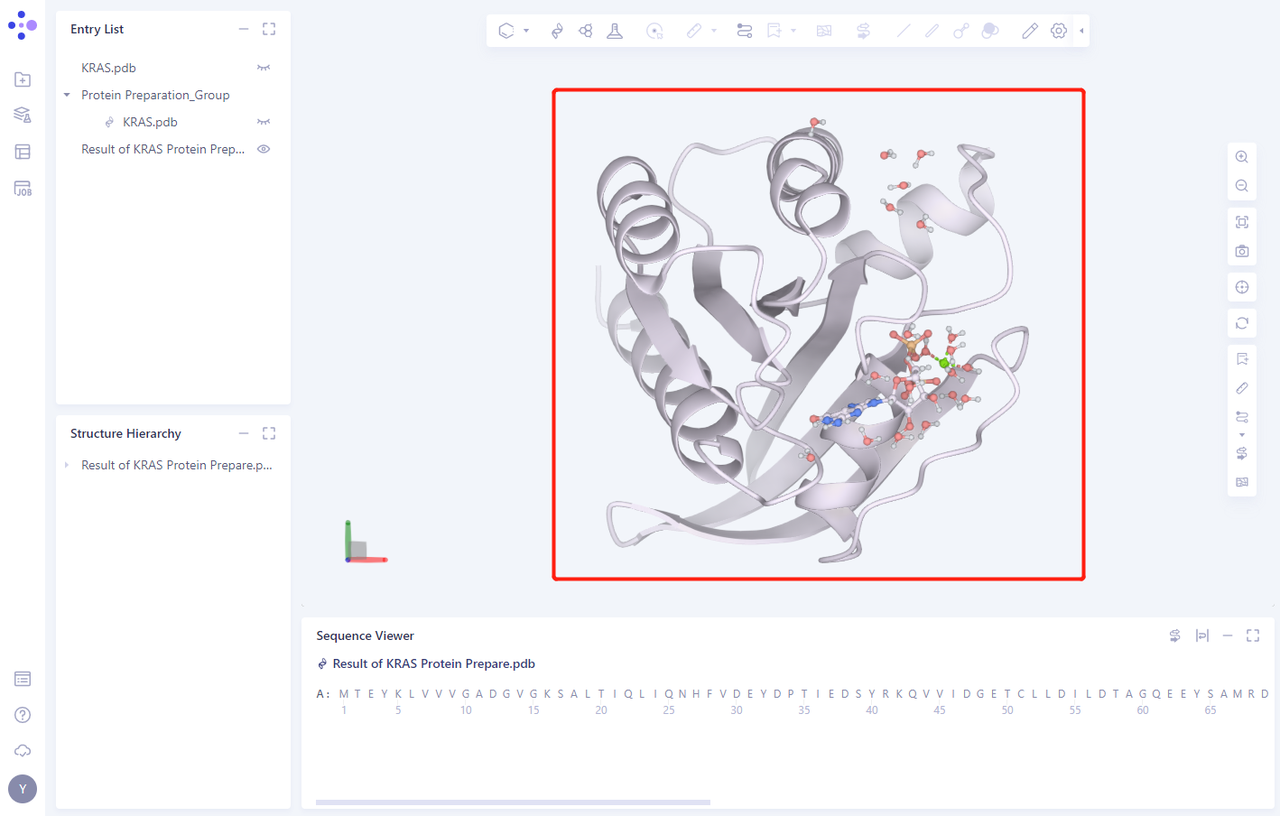
3. Create FEP Calculation Task
3.Click the general menu bar on the left: Menu → Function → Binding Affinity Evaluation → FEP Calculation
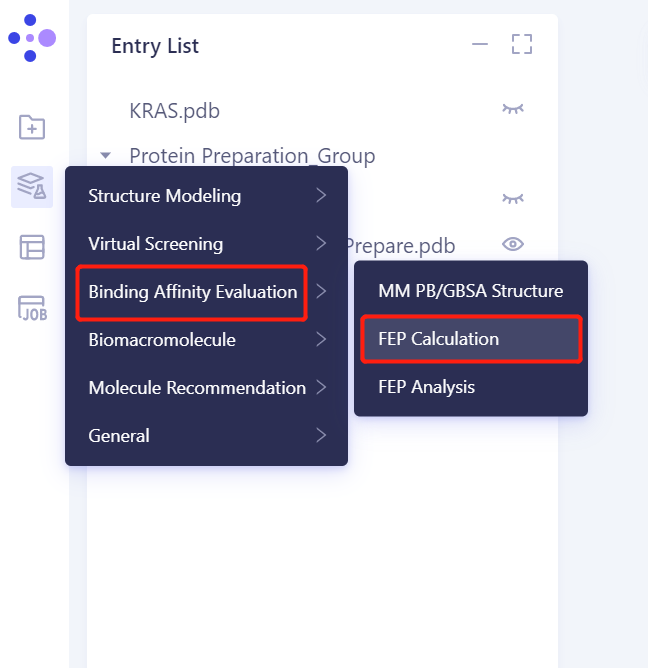
3.2 Select a Prepared Protein From Project
- Select “KRAS _ PDB _ prepared” “as the prepared protein structure and click OK
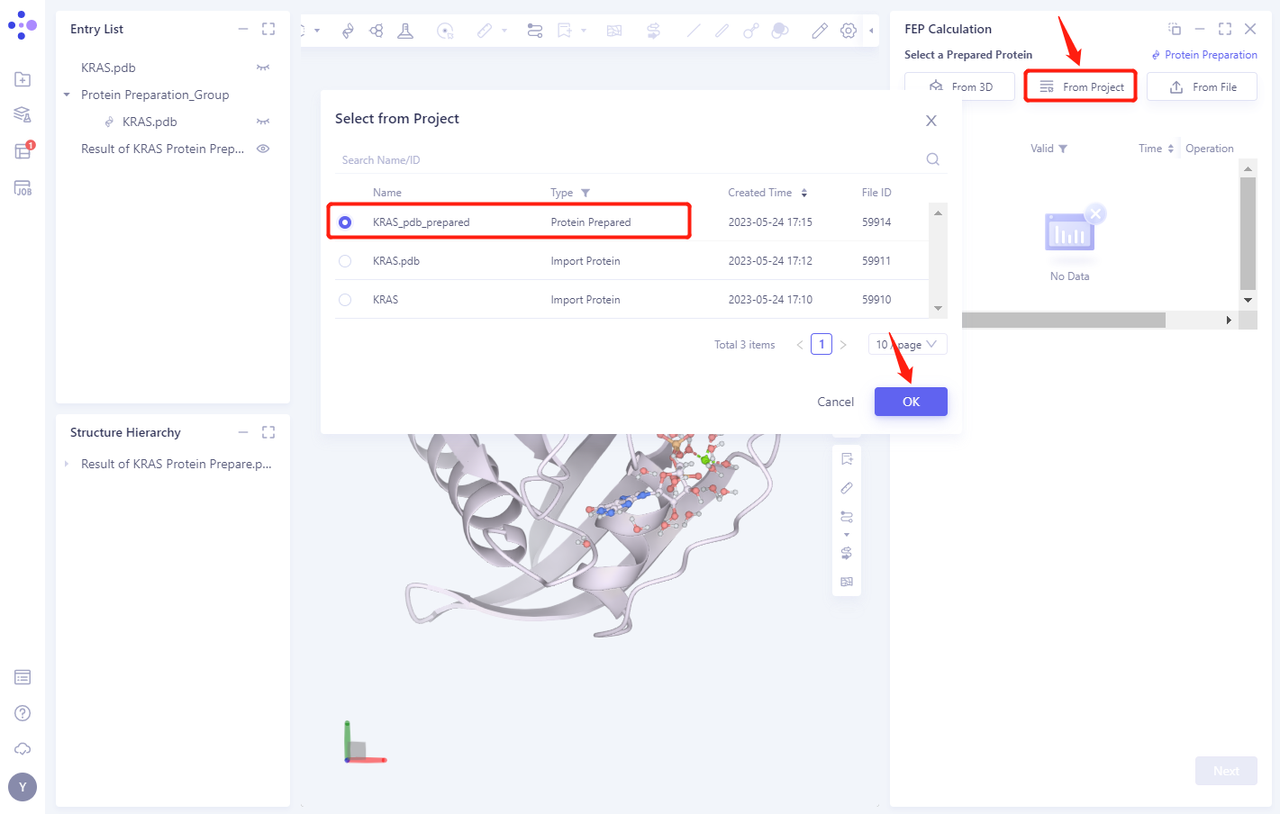
- After clicking OK, the system will automatically check whether the input protein meets the calculation requirements, and the status is “Processing”.
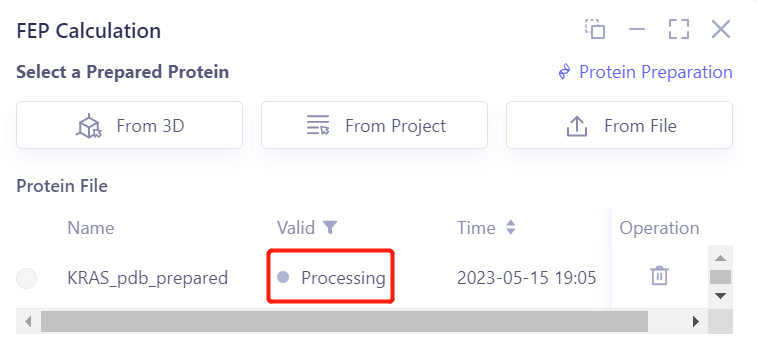
- In less than 1 minute, the system will judge that the protein is “Valid” and click “Next”.
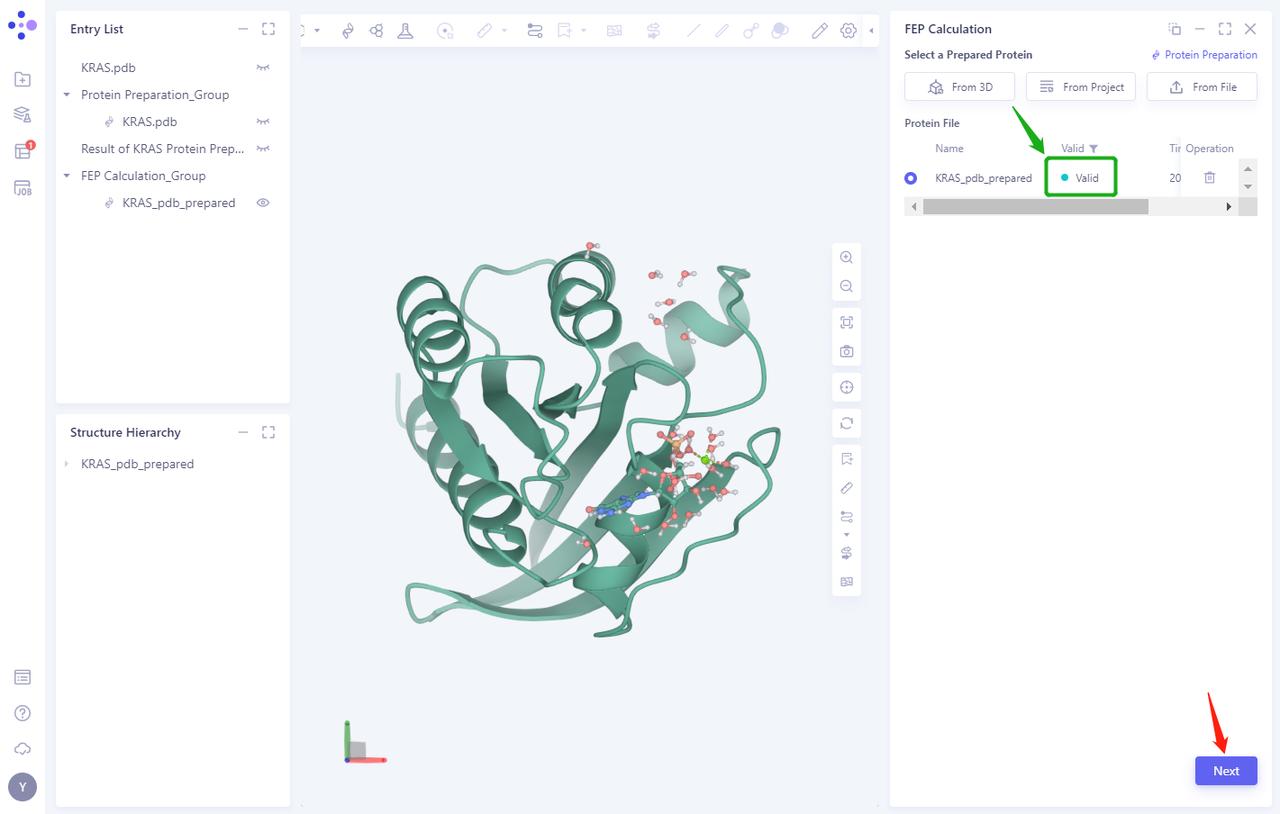
3.3 Select Prepared Ligands From File
-
Click “From File”;
-
Select the 7 compound molecules that have been downloaded in the local folder;
-
Click “Open” to import the ligand structure.
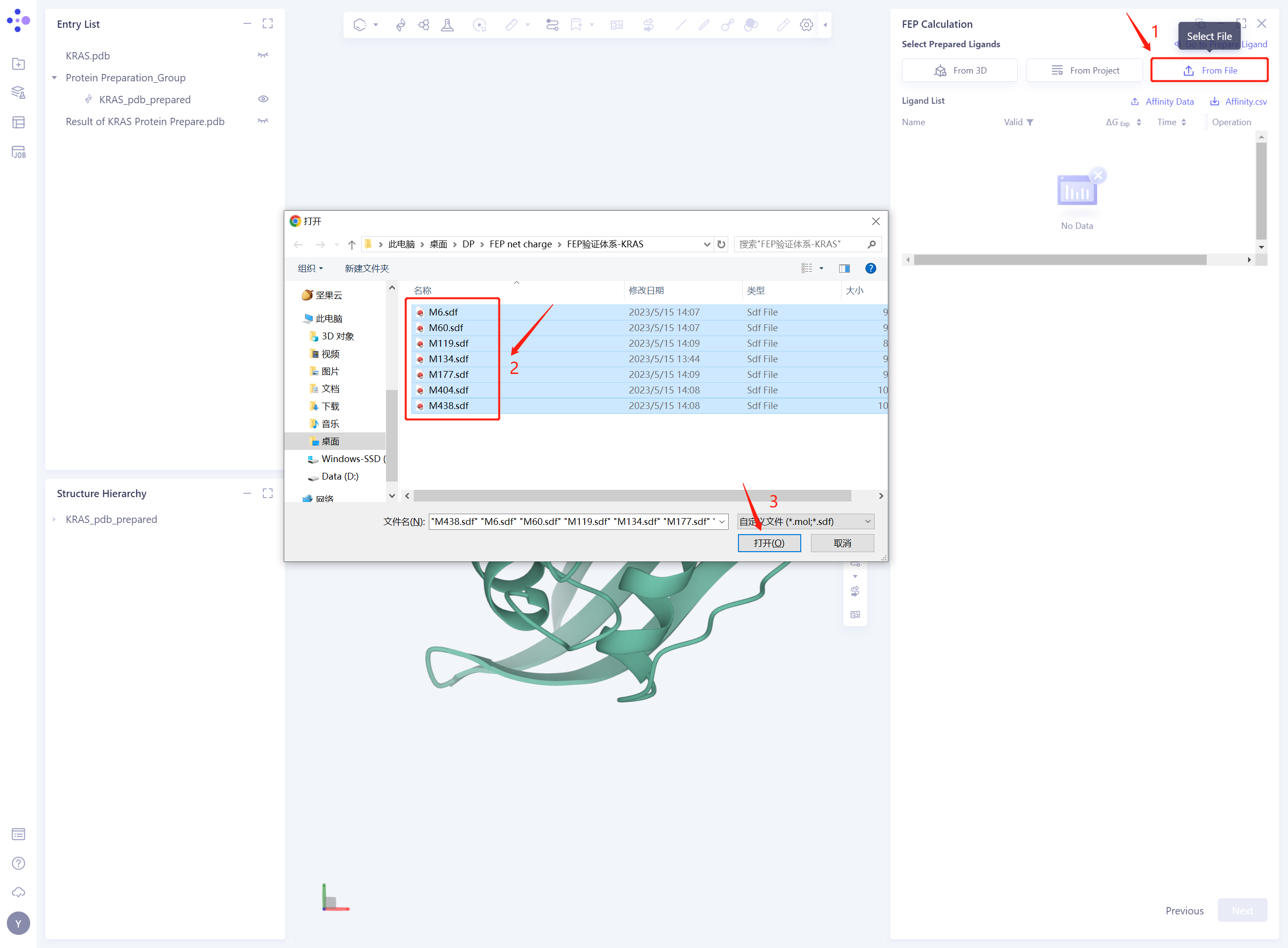
-
Seven ligand structures will be loaded into the Selected File ”. The system automatically checks whether the Ligand entered meets the calculation requirements and the status is “Processing”, and then the system judges that the Ligand is “Valid”.
-
Click Next
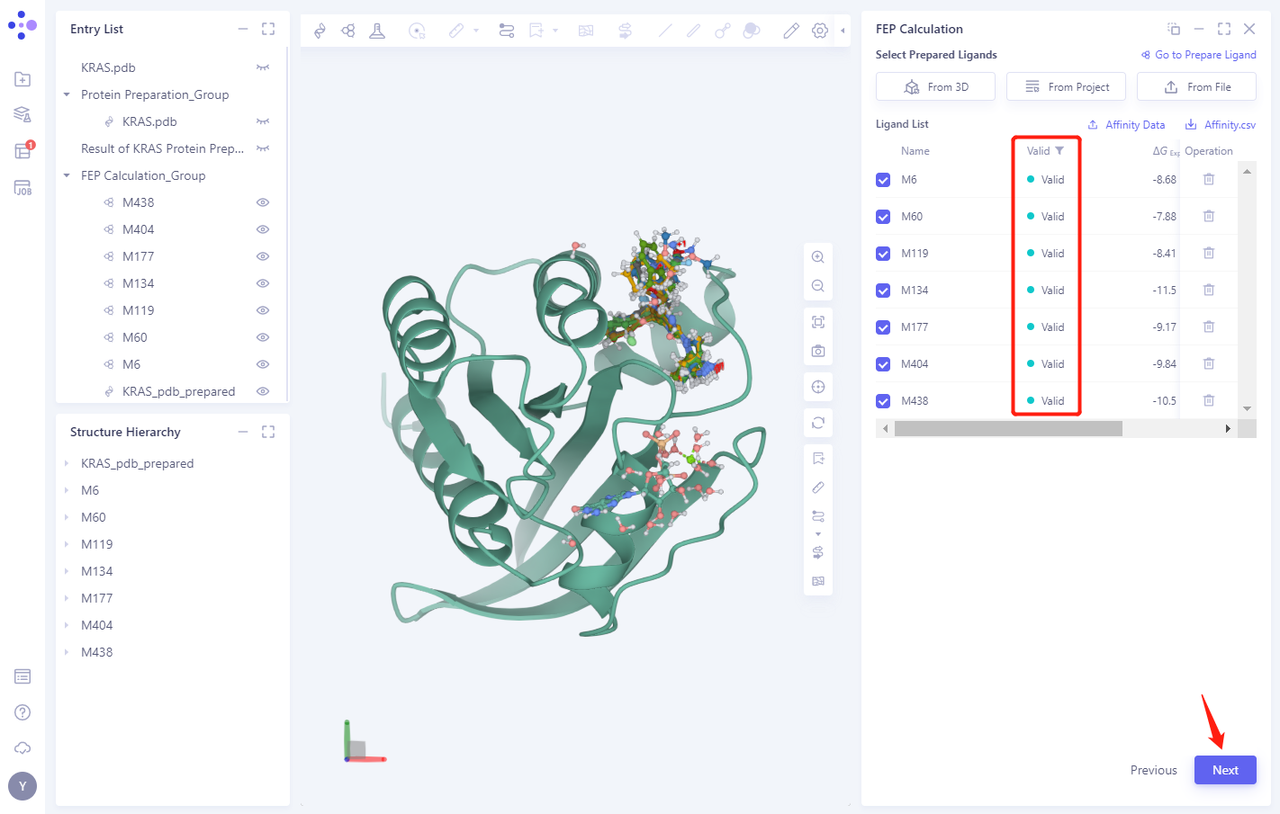
3.4 Alignment
- When prompted whether to perform Ligands Alignment, click Yes
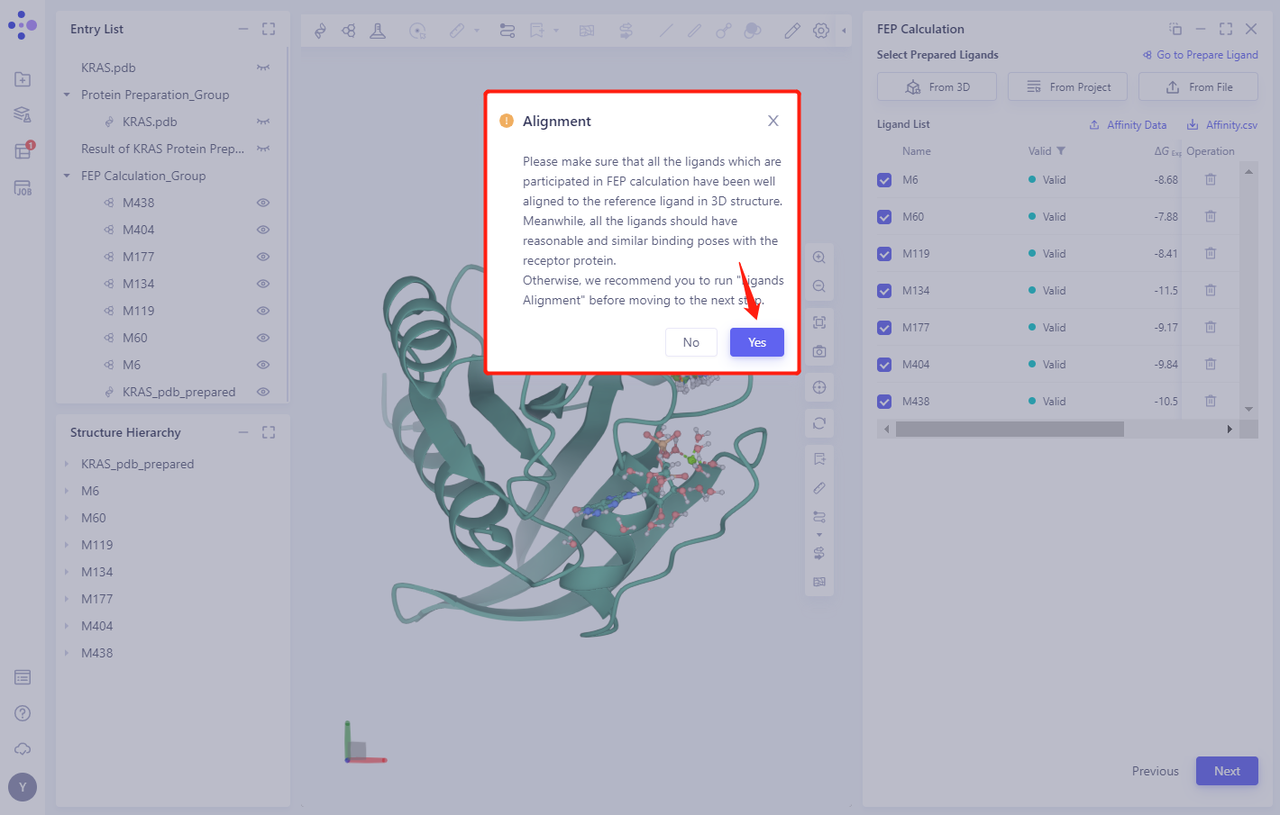
- Select the reference molecule for Alignment. In this case, compound M6 is selected as the reference molecule.
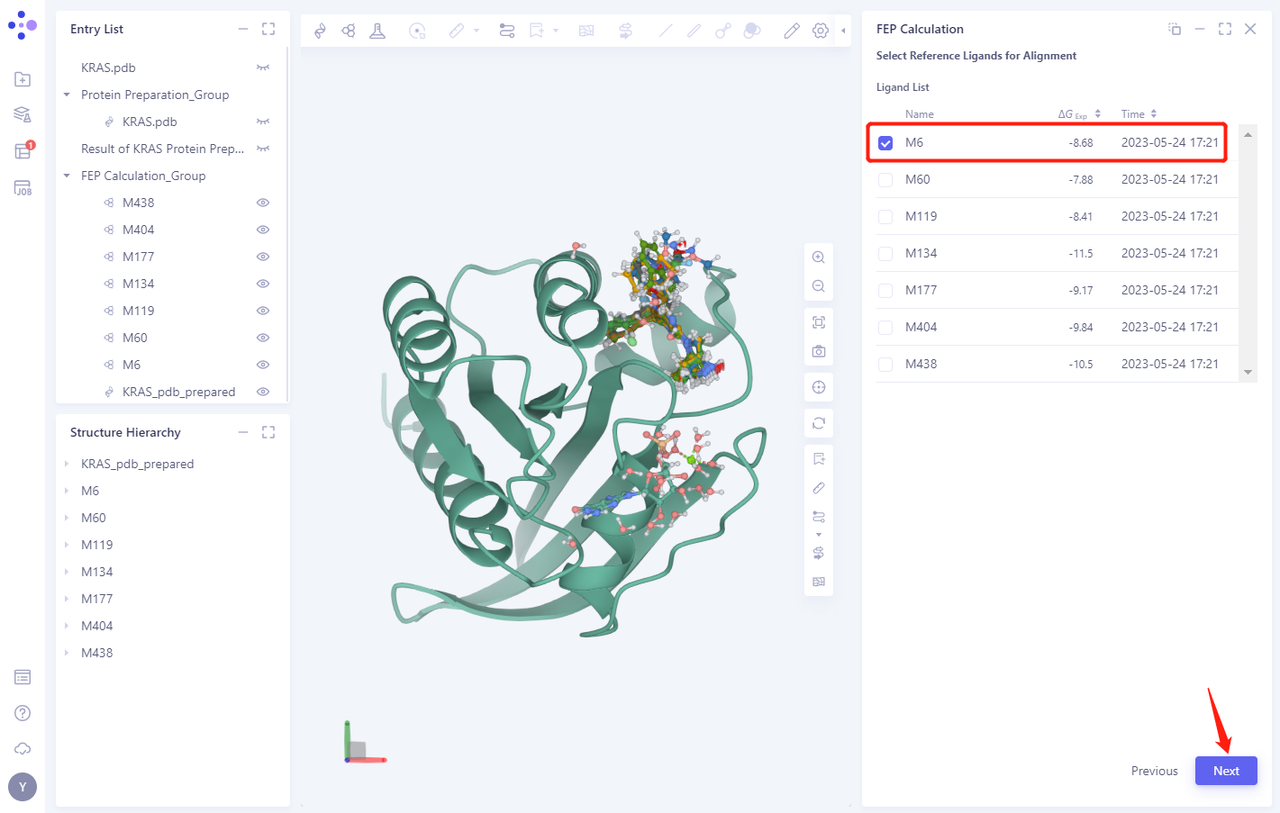
-
Click Next, and the system will automatically select the remaining Ligands for Alignment
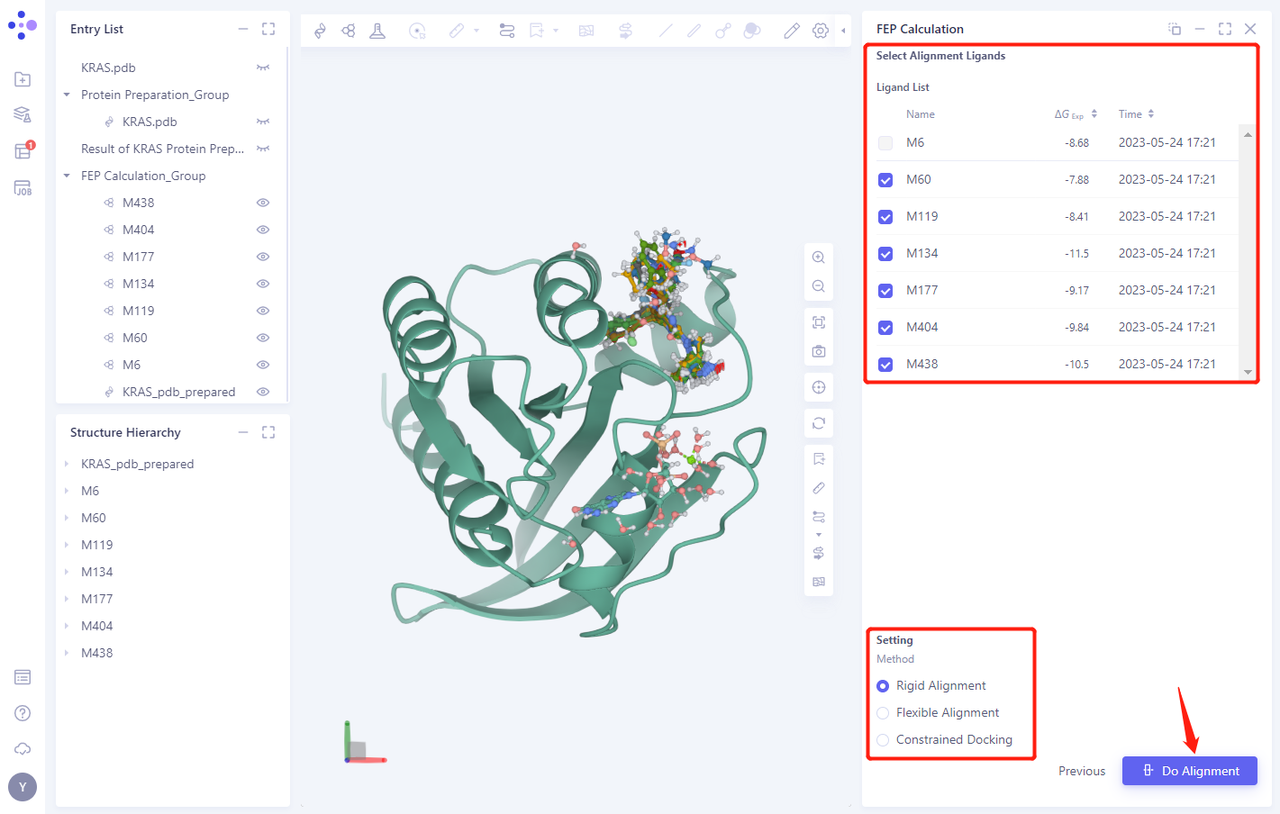
-
For this tutorial, select the Rigid Alignment “Method Alignment
-
Click Do Alignment to submit the task
- After Alignment calculation, the system automatically displays the Alignment Ligand Checklist “list to check whether there is any error in the Alignment task
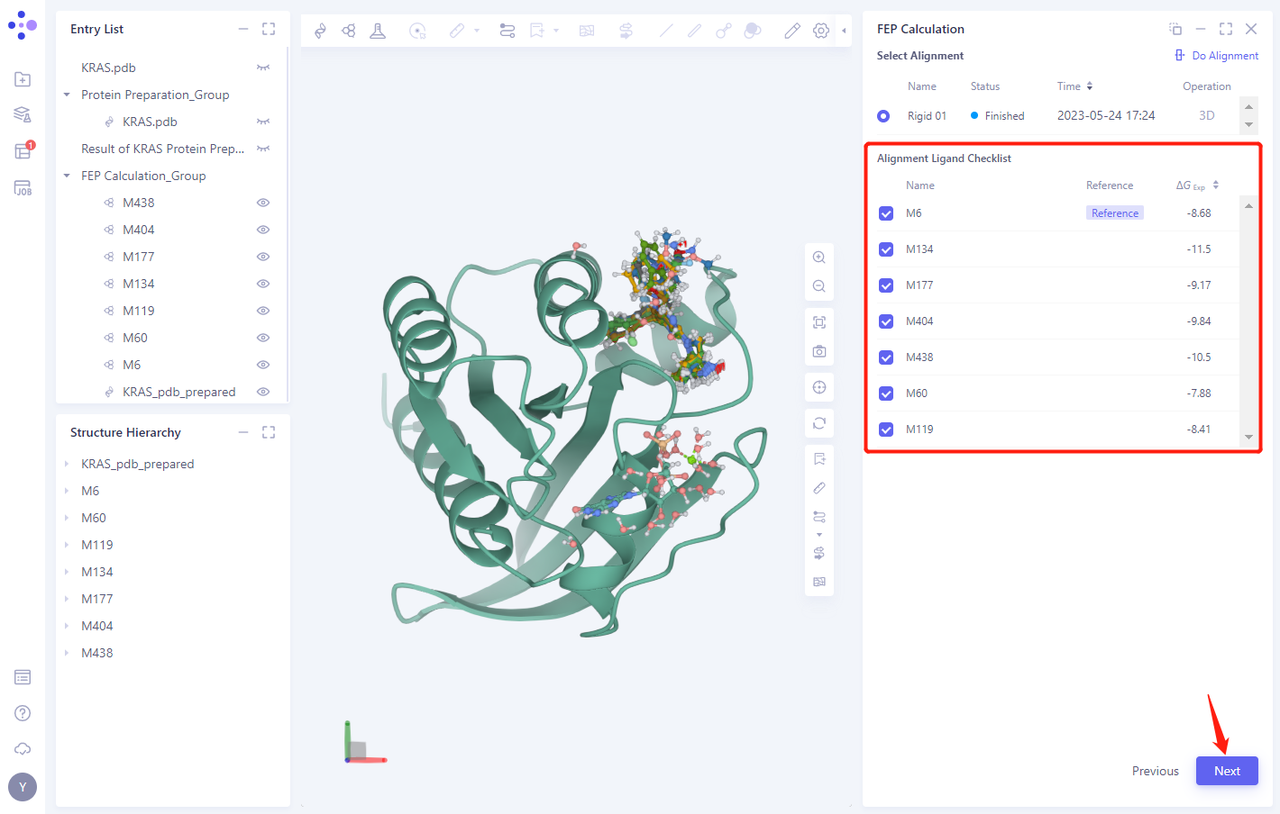
- Click Next to proceed to the next step
3.5 Build Network
-
Build Network Seeting:
-
In this case, A-optimal is chosen as the strategy to optimize the perturbation graph
-
Based on the number of Ligands, the Number of Cycles is automatically set to 4
-
Leave the rest of the parameters as default and click Build Network
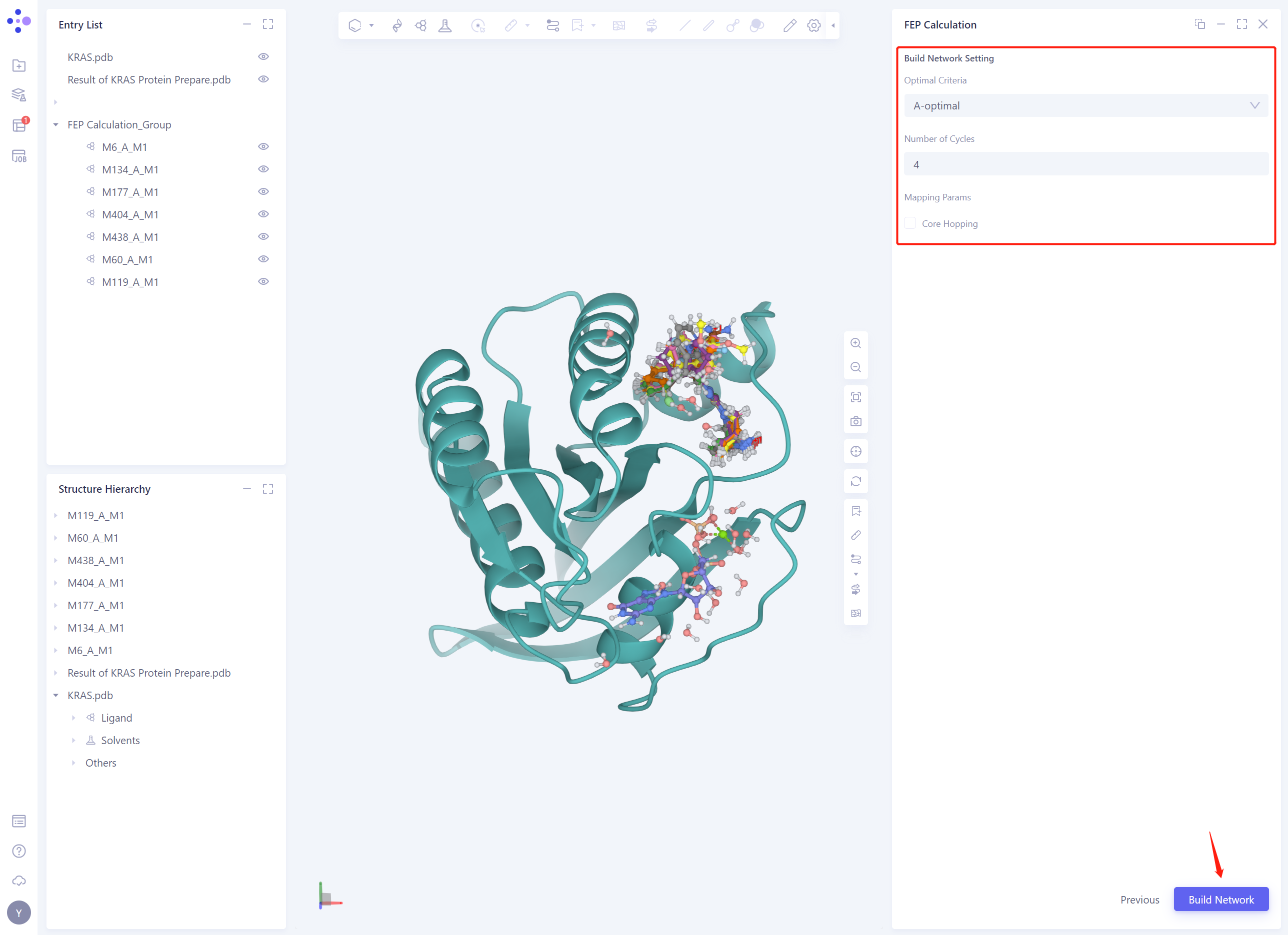
- After the Build Network task is submitted, the interface displays the Perturbation is running. ”.
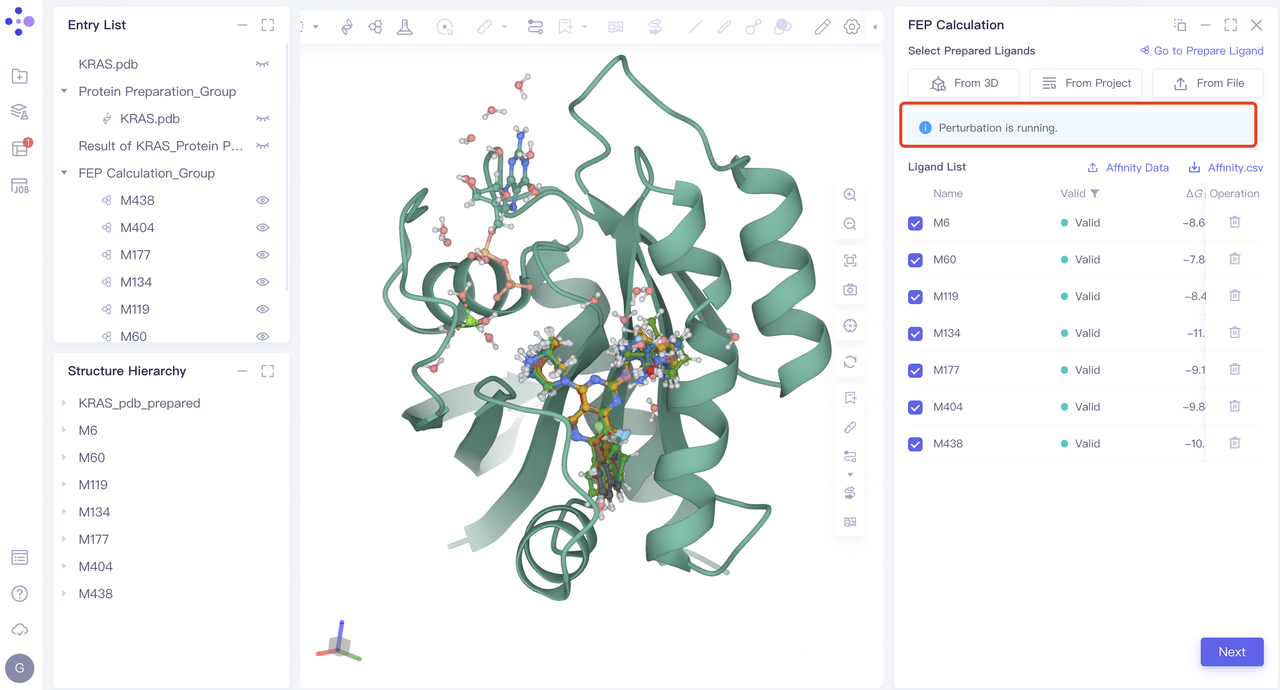
-
After the Perturbation task calculation is completed, the Perturbation Map “interface will pop up automatically
-
Note: If there is no popup of the perturbation graph page, you can check whether the page is blocked.
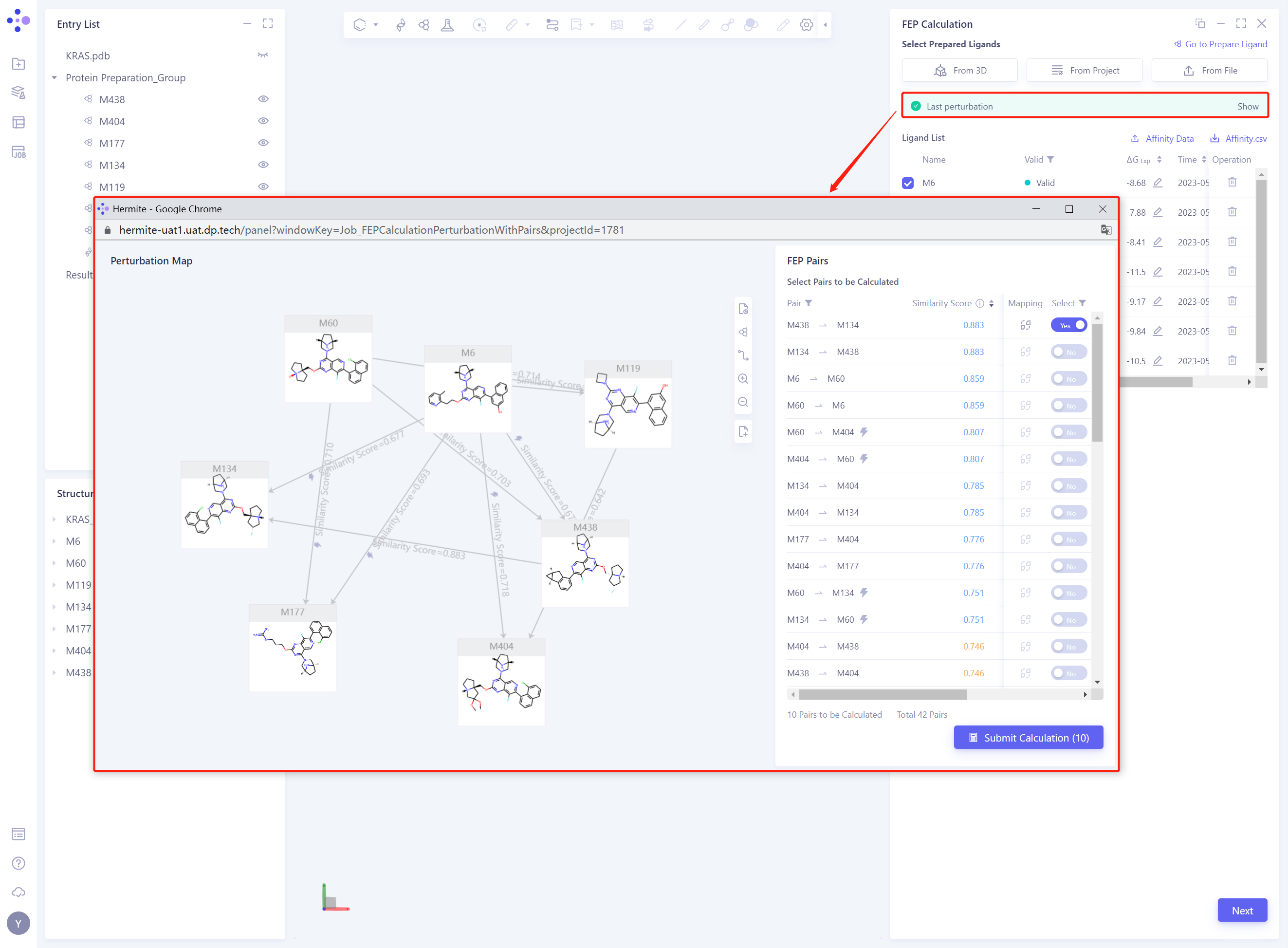
3.6 Modify Perturbation Map
- Select status of Pairs is changed to Yes, only connected Pairs are displayed
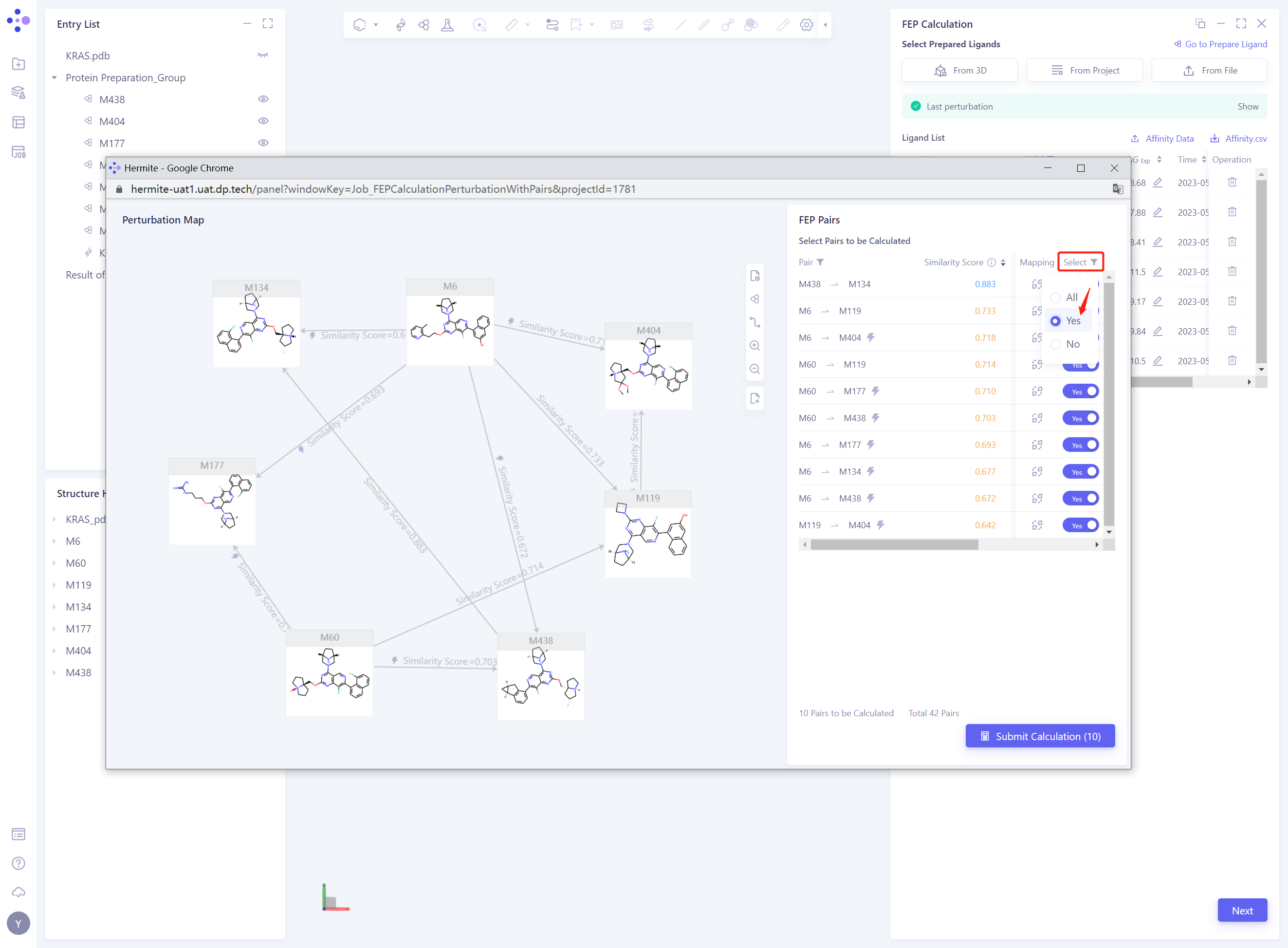
- Delete the Pairs with Similarity Score “lower than 0.7, that is, change the” Select “status of the corresponding Pairs from” Yes “to” No ”
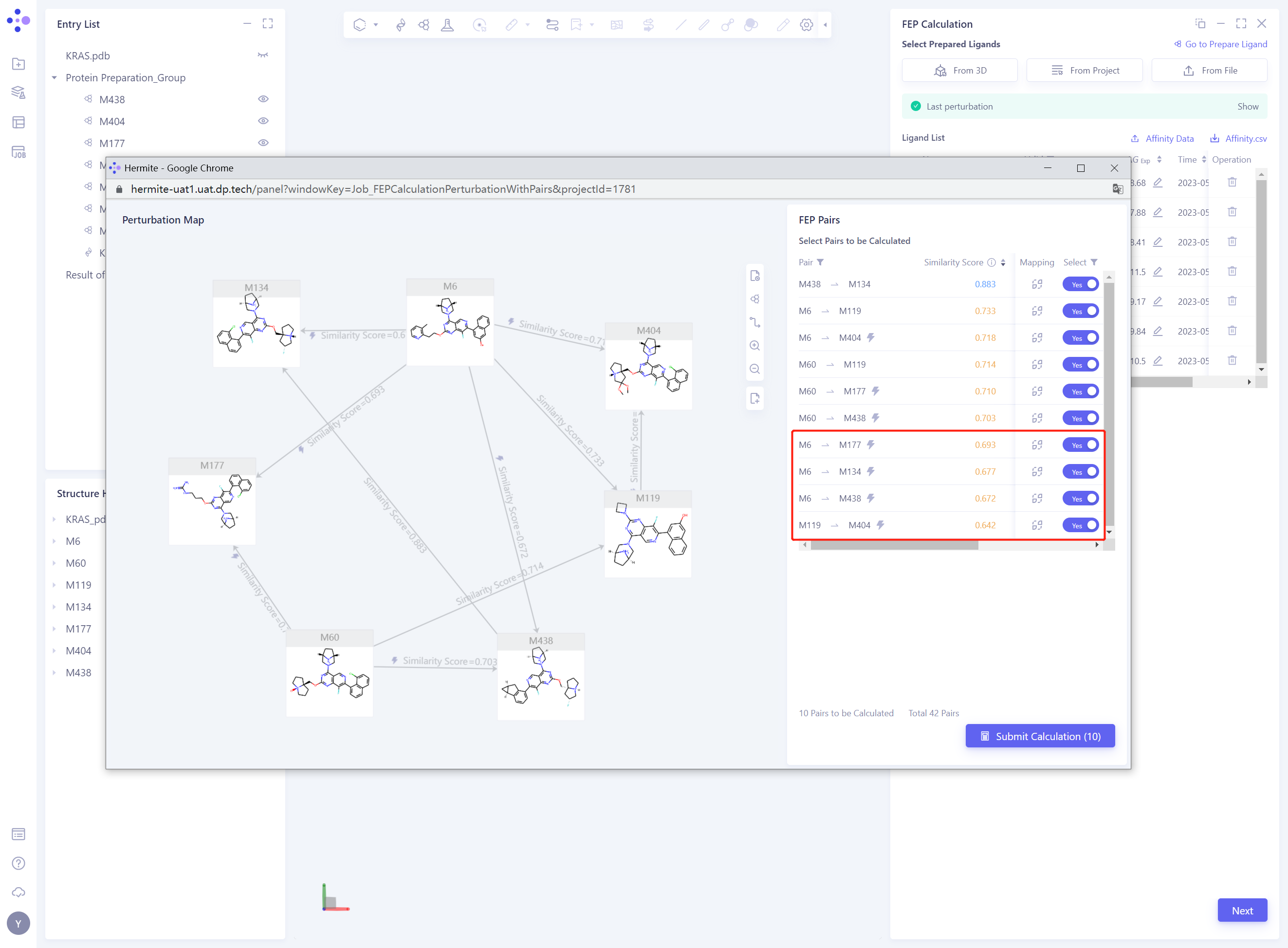
- Modify the Perturbation Map by changing the Select status of the Pairs to All to display all the Pairs
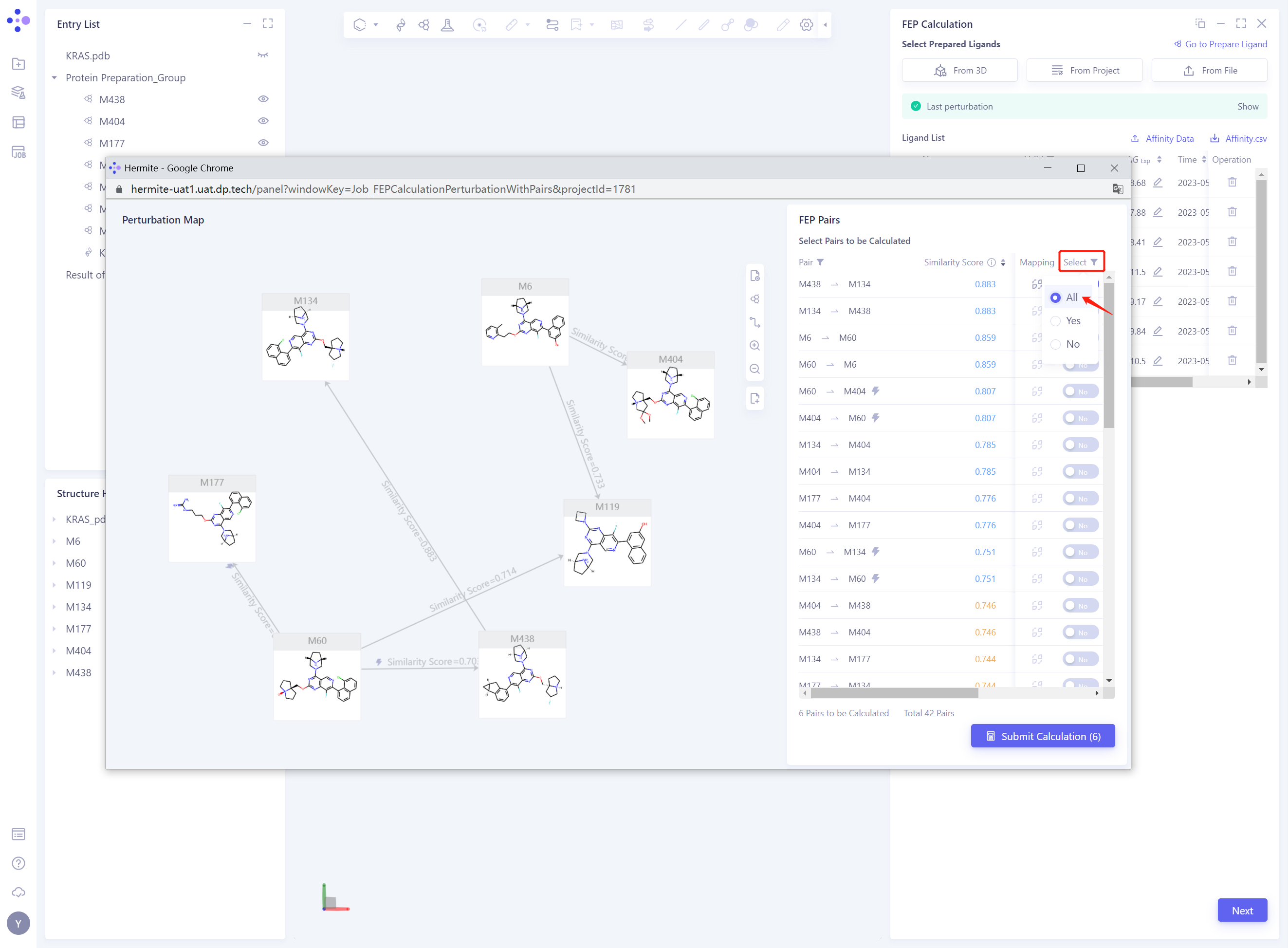
-
Modify the Perturbation Map based on the recommendations in the FEP calculation:
-
To reduce the error in FEP calculation, it is recommended that the calculated intermolecular Similarity Score should be greater than 0.7
-
In order to reduce the error of FEP calculation, it is suggested that the Pairs involved in FEP calculation should form a thermodynamic cycle
-
-
We modify the Perturbation map to the following form:
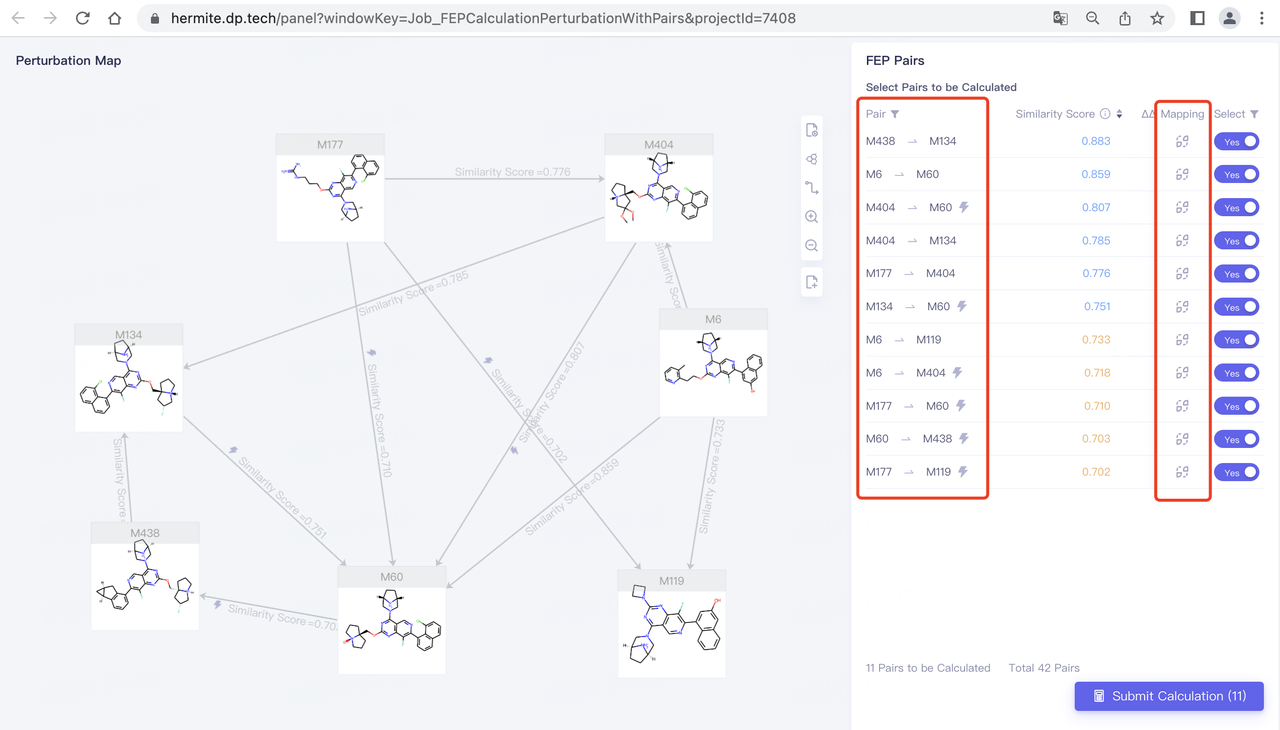
- Click Submit Calculation to set FEP task submission
3.7 FEP parameter setting and submission task
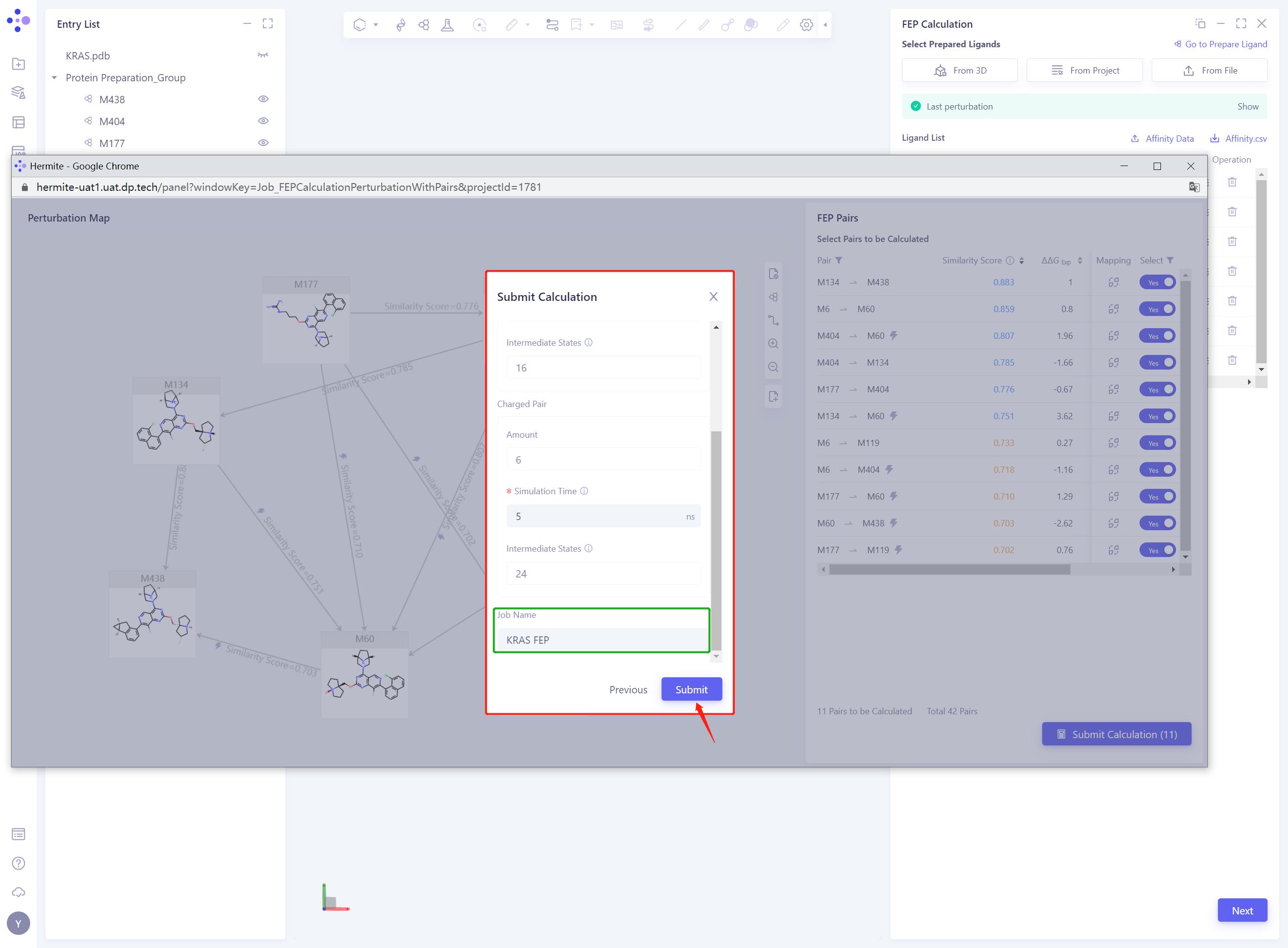
- Check the Pairs used for the calculation and click Next without error
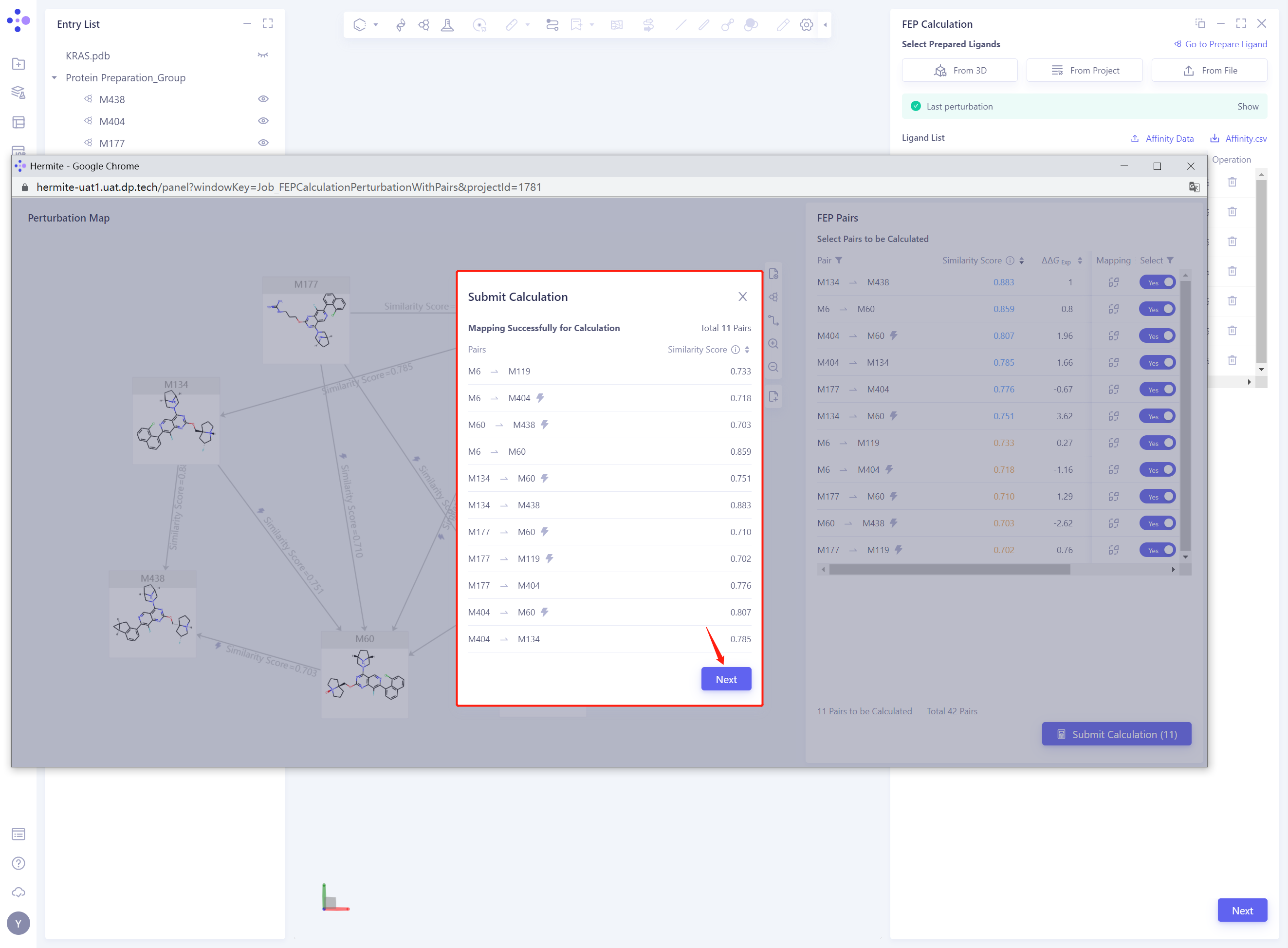
- Keep the default parameters for FEP calculation, set the Job Name to “KRAS FEP” ”, and click Submit to submit the task
4. Analysis of results
- Note: After FEP Calculation is completed, the program will automatically complete FEP Analysis based on the pairs submitted by FEP Calculation, so you only need to click FEP Analysis to view the Uni-FEP calculation results.
4.1 Entrance
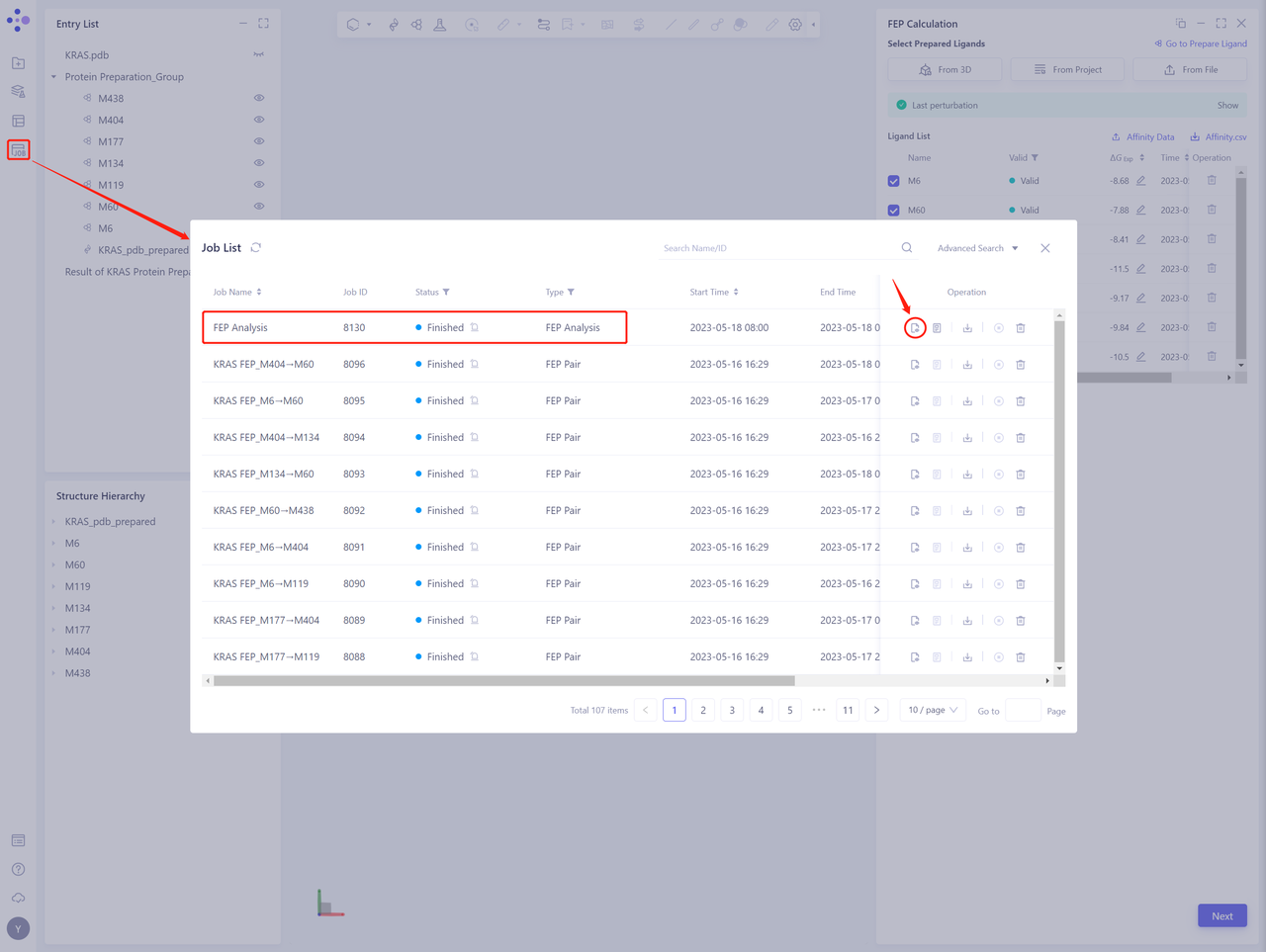
- In the left general menu bar Job → Job List, find the “FEP Analysis” Calculation Task “and click” Show ”.
4.2 Results presentation
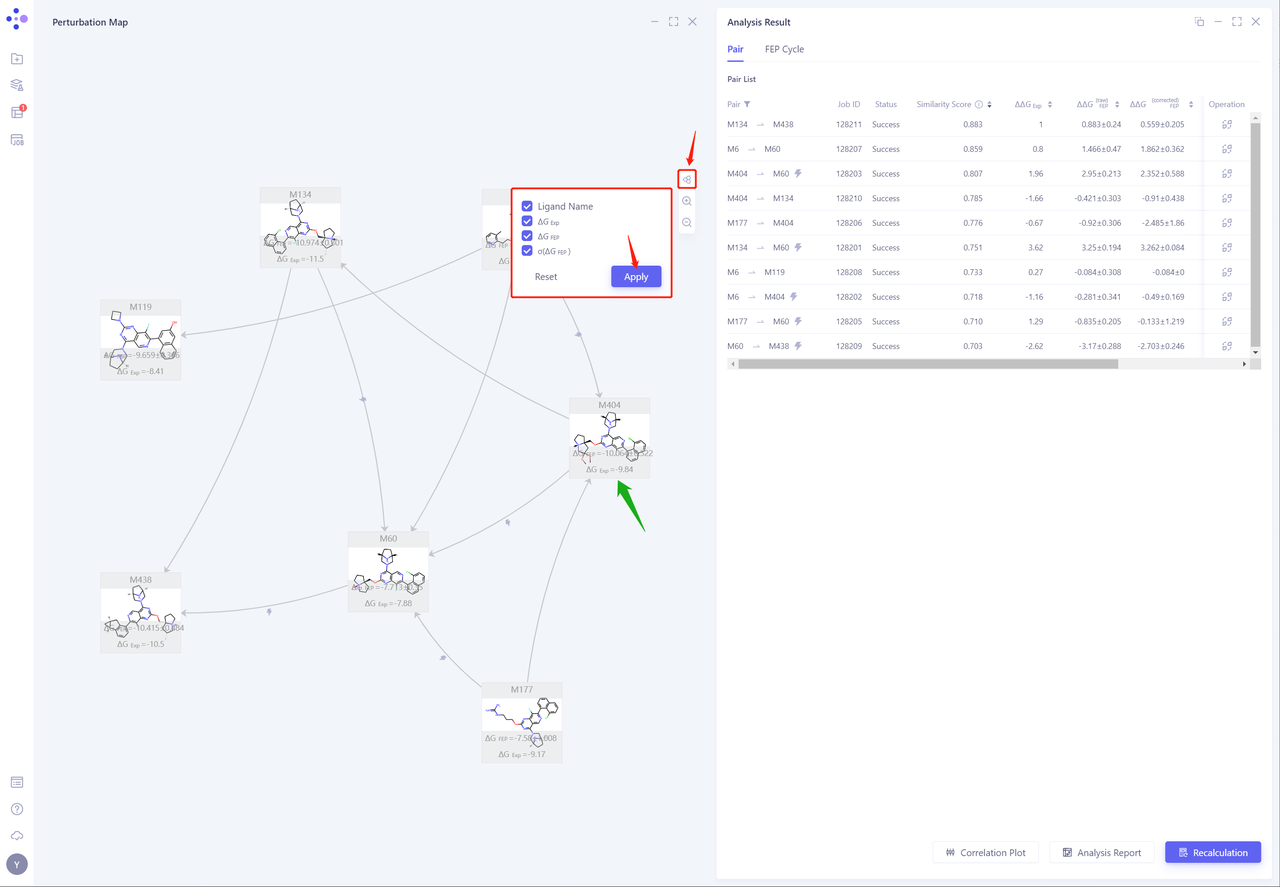
- Click the “Node” button, check “Ligand Name, ΔG EXP, ΔG FEP and σ (ΔG FEP)”, and click “Apply” to display the experimental affinity and calculated affinity values
-
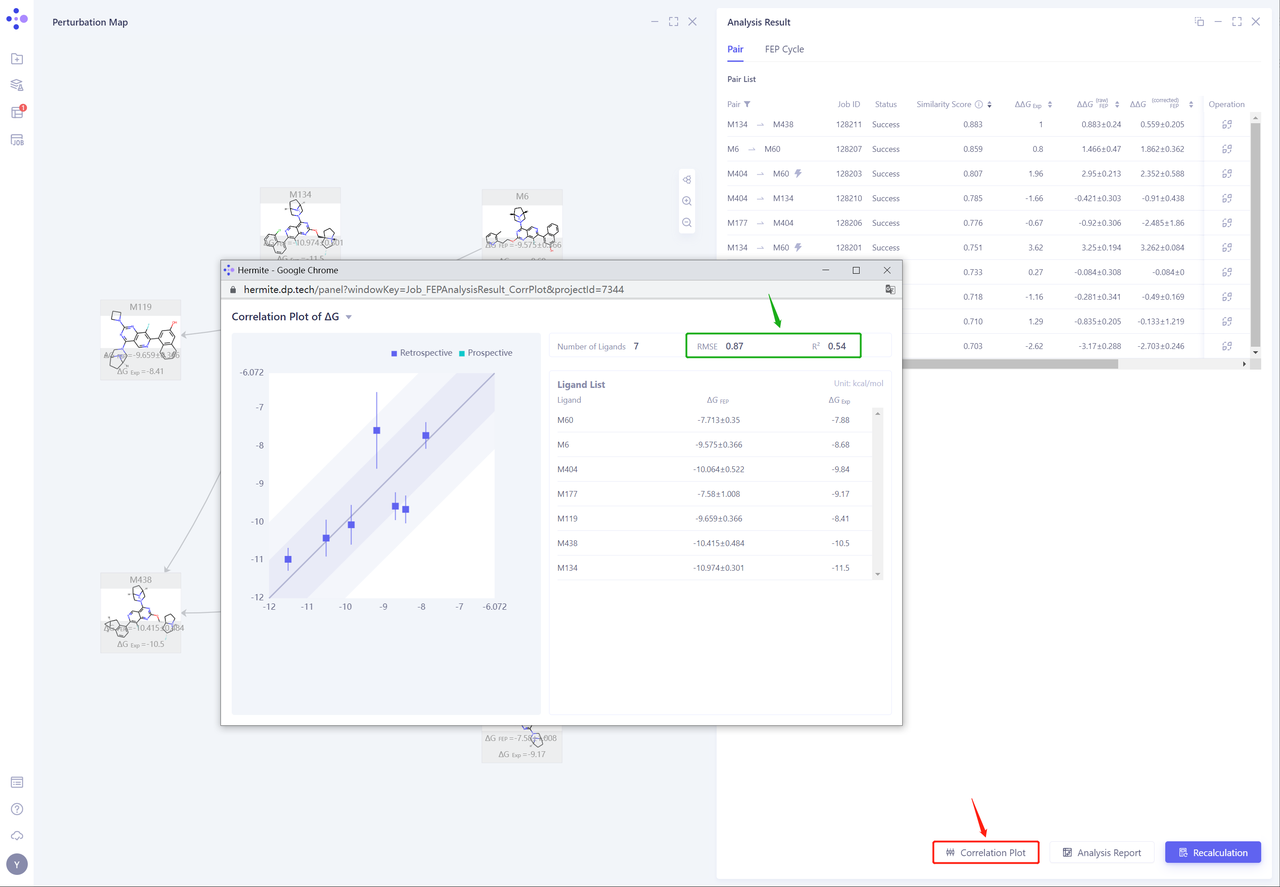
- Click “Correlation Plot” to analyze the correlation between the experimental ΔG and the calculated ΔG. The RMSE and R ² are 0.87 kcal/mol and 0.54, respectively, showing a good correlation.
- Click “Analysis Report” to analyze the reliability of calculation results of each Pairs.
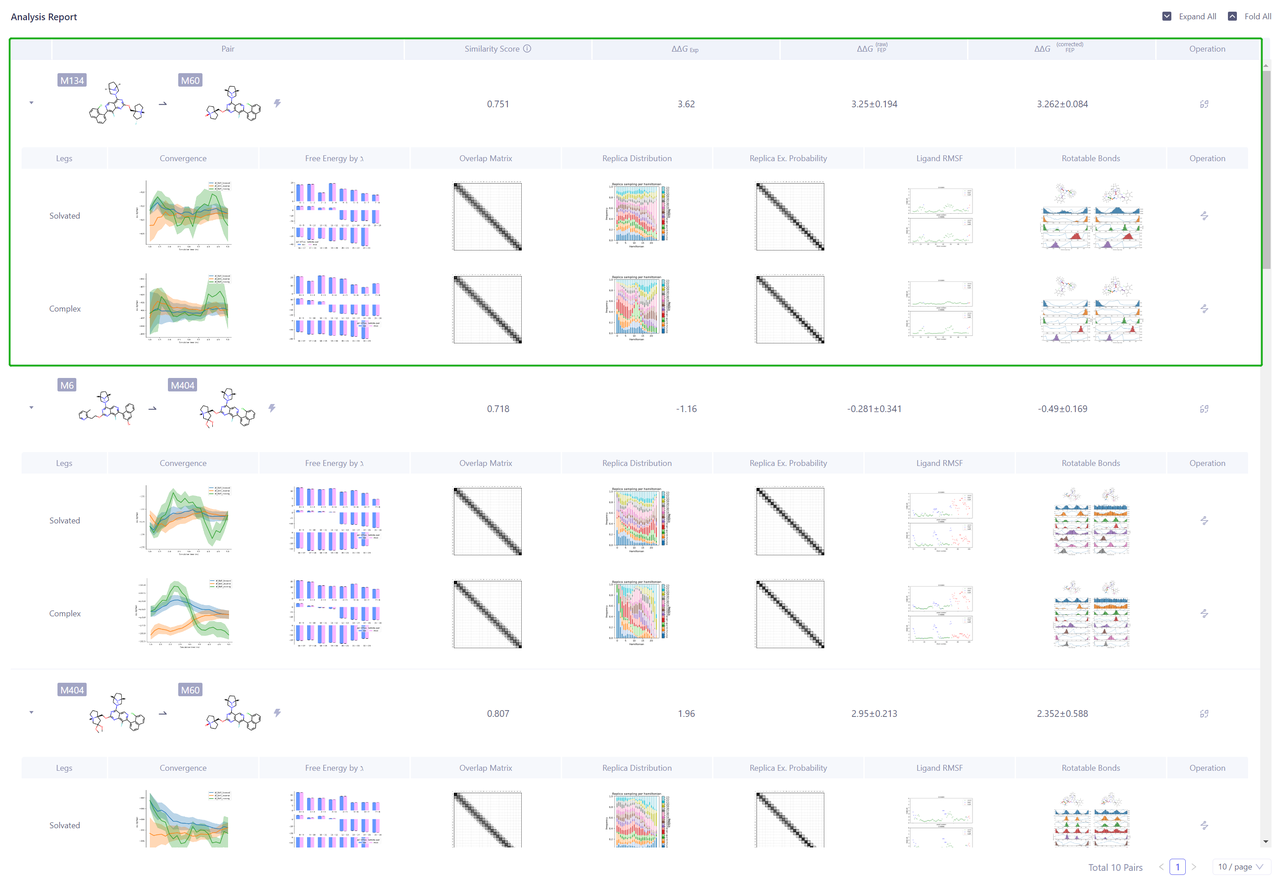
5. Sum up
In this case, KRAS-G12D mutant was used as an example to calculate the FEP between protein and small molecule for seven compounds with known activity. The results show that Uni-FEP can accurately evaluate the relative binding free energy between the compound and the target for the KRAS-G12D mutant, thus guiding the optimization of the lead compound.
6. References
[1] KRAS as a Therapeutic Target. Clin Cancer Res. (2015) 21 (8): 1797–1801.
[2] The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature. volume 575, pages 217–223 (2019)
[3] Identification of MRTX1133, a Noncovalent, Potent, and Selective KRAS(G12D) Inhibitor. J. Med. Chem. 2022, 65, 4, 3123–3133