Protein Alignment
Introduction
Protein structure determines its function and contains a large amount of biological information. Small differences in structure may lead to great differences in function. Comparing the differences between two protein structures is one of the most commonly used research methods in structural biology and computational biology.
Iron-sulfur cluster protein is an important mitochondrial functional protein, which plays a key role in cell energy metabolism, electron transfer, iron/sulfur storage and many other processes. IscA is one of the highly conserved members of the hesB iron-sulfur protein subfamily, which is involved in the synthesis of iron-sulfur cluster proteins, so it plays an important role in the iron-sulfur cluster assembly protein and cascade reaction system. IscA (PDB ID: 1R94) of Escherichia coli is a homotetrameric structure. SufA (PDB ID: 2D2A), a homologous protein of IscA, acts as a scaffold in the biosynthesis of iron-sulfur clusters and has a homodimeric structure. These two different proteins function in the same biological pathway. This tutorial aligns the 1R94 protein and the 2D2A protein based on the Protein Alignment module of the Hermite ® platform.
1. Import Protein Structure File
-
In the left general menu bar 'File' → Get PDB → In the pop-up window, enter the PDB ID: 1R94 → Import.
-
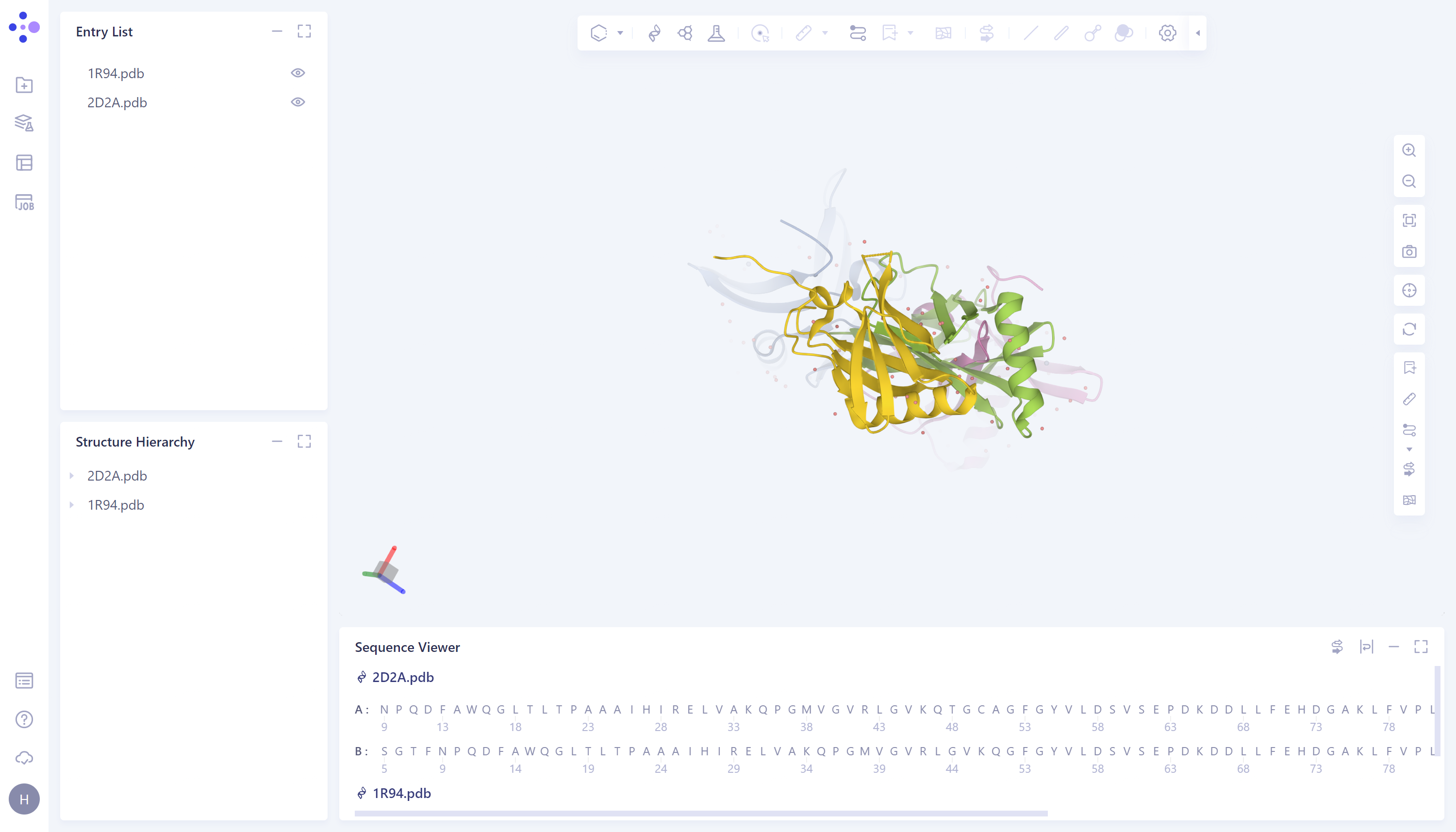
The import procedure of '2D2A' is the same as above.
2. Protein Alignment
2.1 Entrance
- Left general menu bar Menu → Function → General → Protein Alignment.
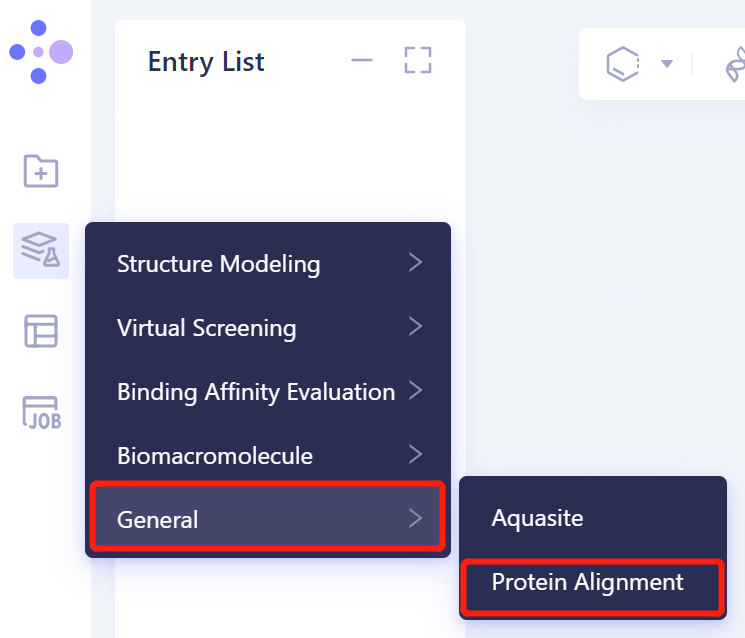
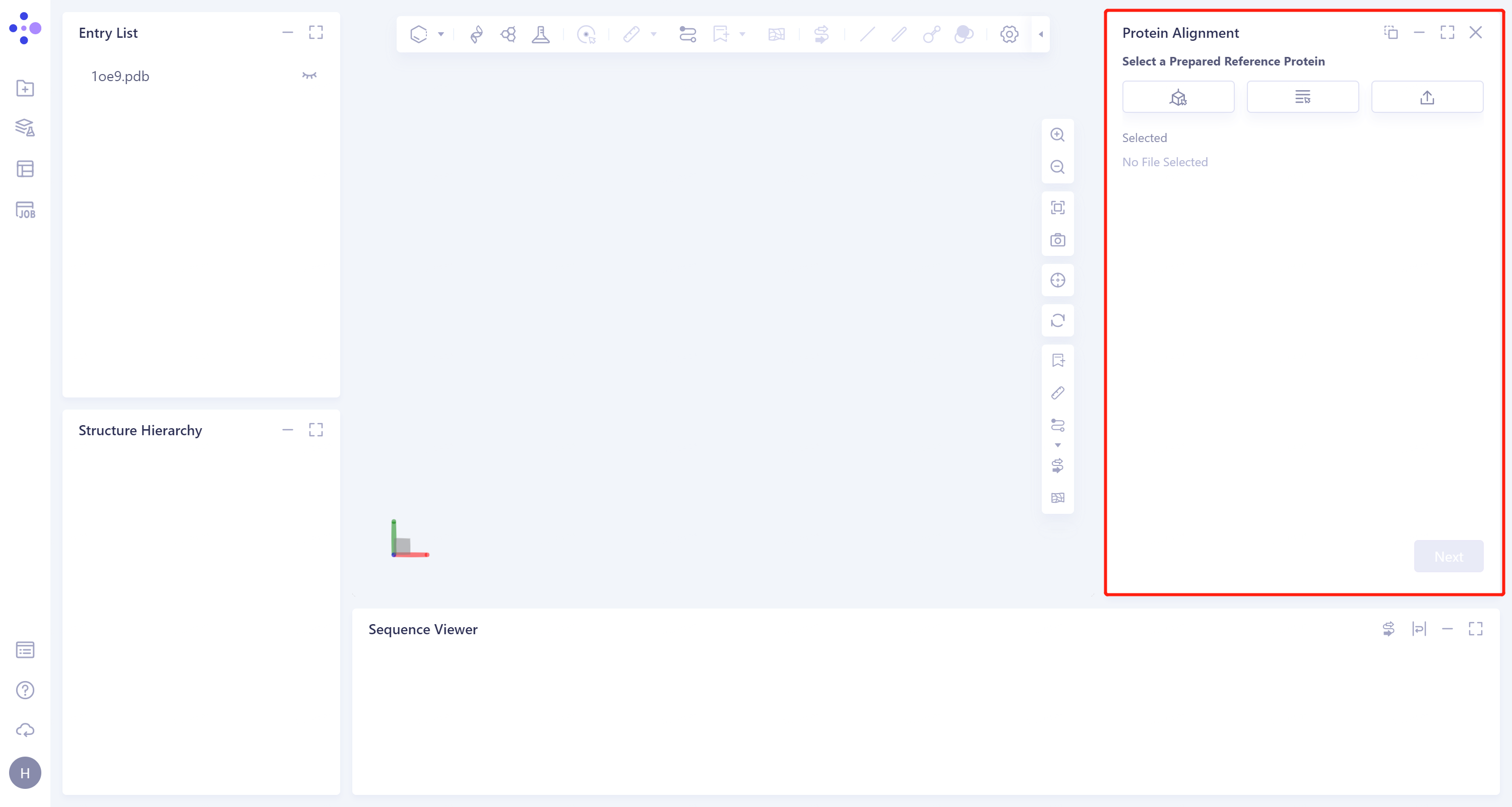
- The interface of 'Protein Alignment' (shown in the red box) appears on the right side, and the overall interface is as follows:
2.1 Introduction of reference protein
- In the 'Protein Alignment' operation window on the right side of the interface, first import the reference protein structure: click 'Select from 3D Workspace', the 'Select Structure' window pops up → Select the protein structure of '1R94' in the Structure Hierarchy → Click 'OK' in the 'Select Structure' window.
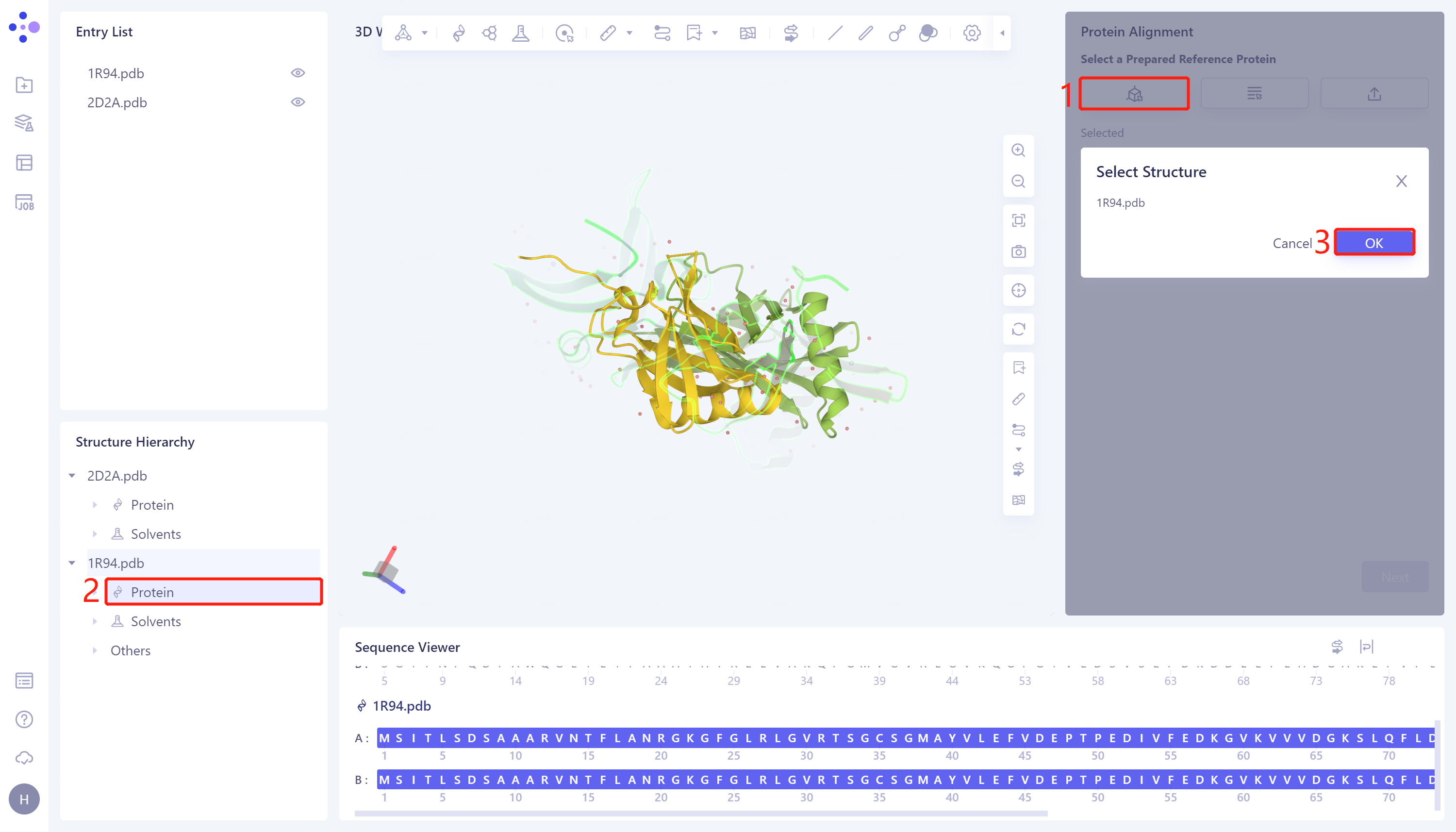
2.2. Introduction of proteins to be compared
- Click 'Next' to enter the selection window of protein to be compared, and import the protein structure to be compared: click 'Select from 3D Workspace', the 'Select Structure' window pops up → Select the '2D2A' protein structure in the 'Structure Hierarchy' → Click 'OK' in the 'Select Structure' window.
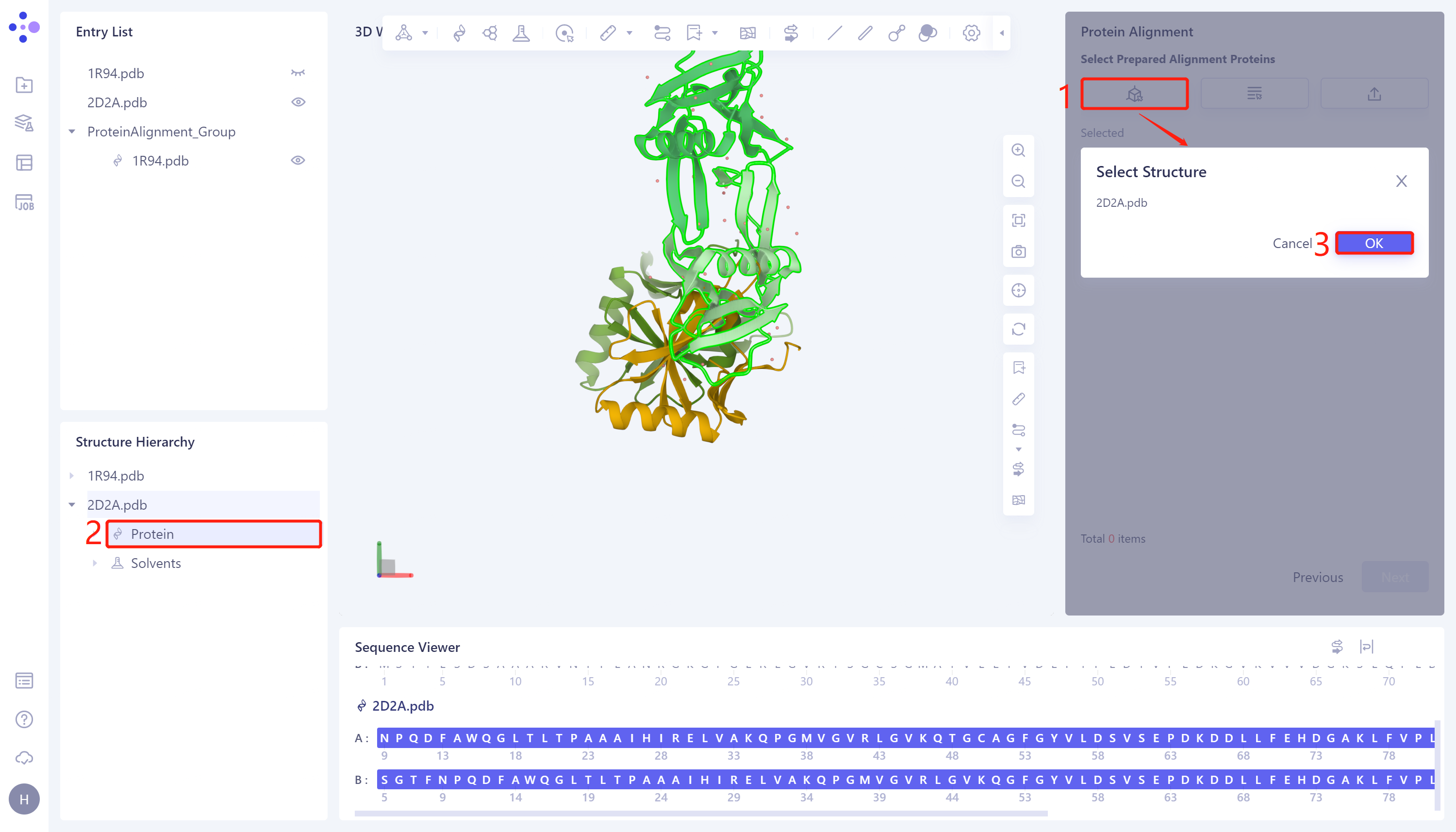
2.3 Parameter setting
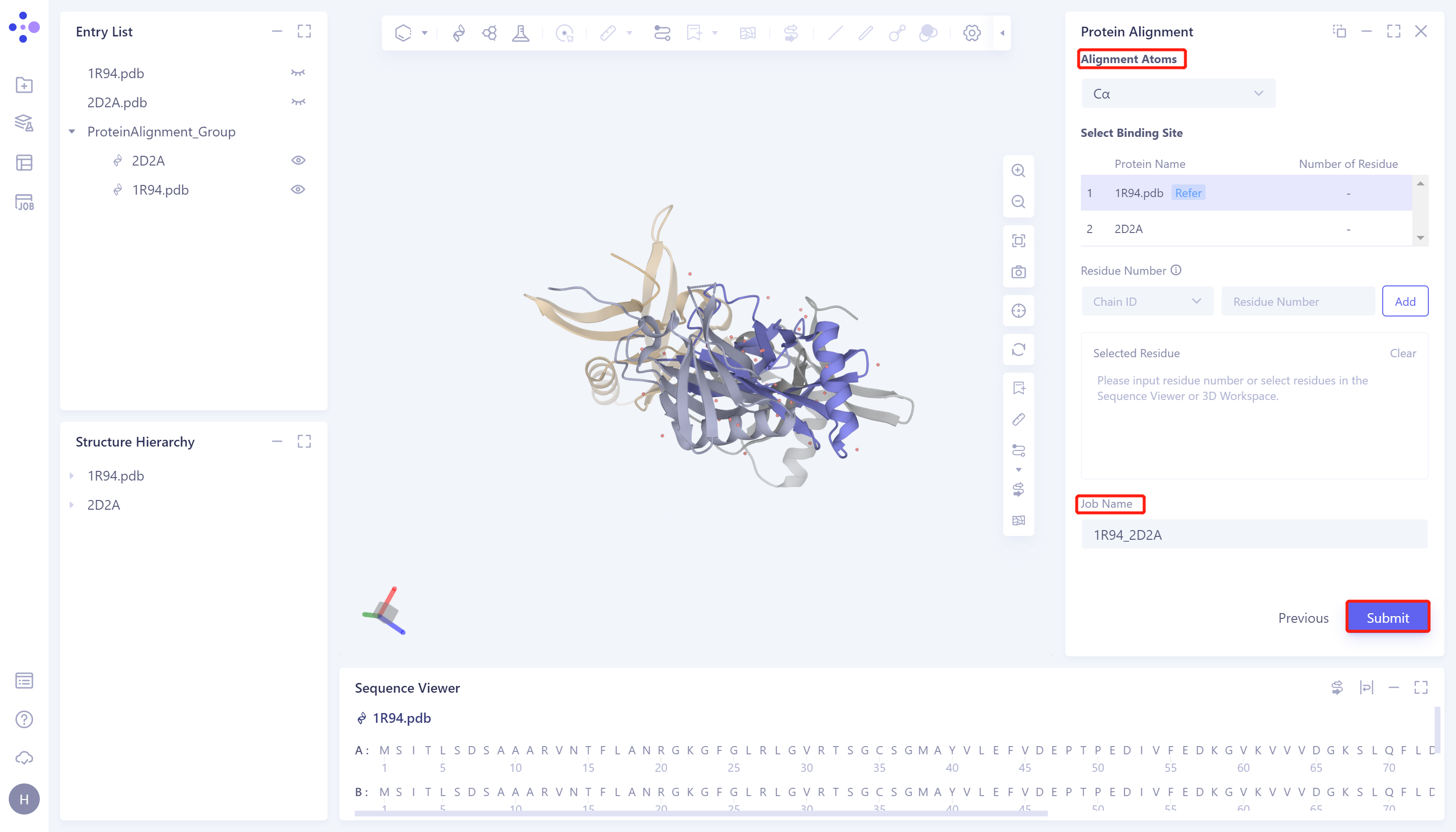
- Click 'Next' to enter the parameter setting page. 'Alignment Atoms' selects 'Cα' to perform alignment based on the α-carbon atoms between proteins. Since there is no special comparison requirement, the parameter setting of 'Select Binding Site' is skipped. Name the task as '1R94_2D2A'. Click 'Submit' to submit the task.
3. Analysis of results
3.1 Entrance
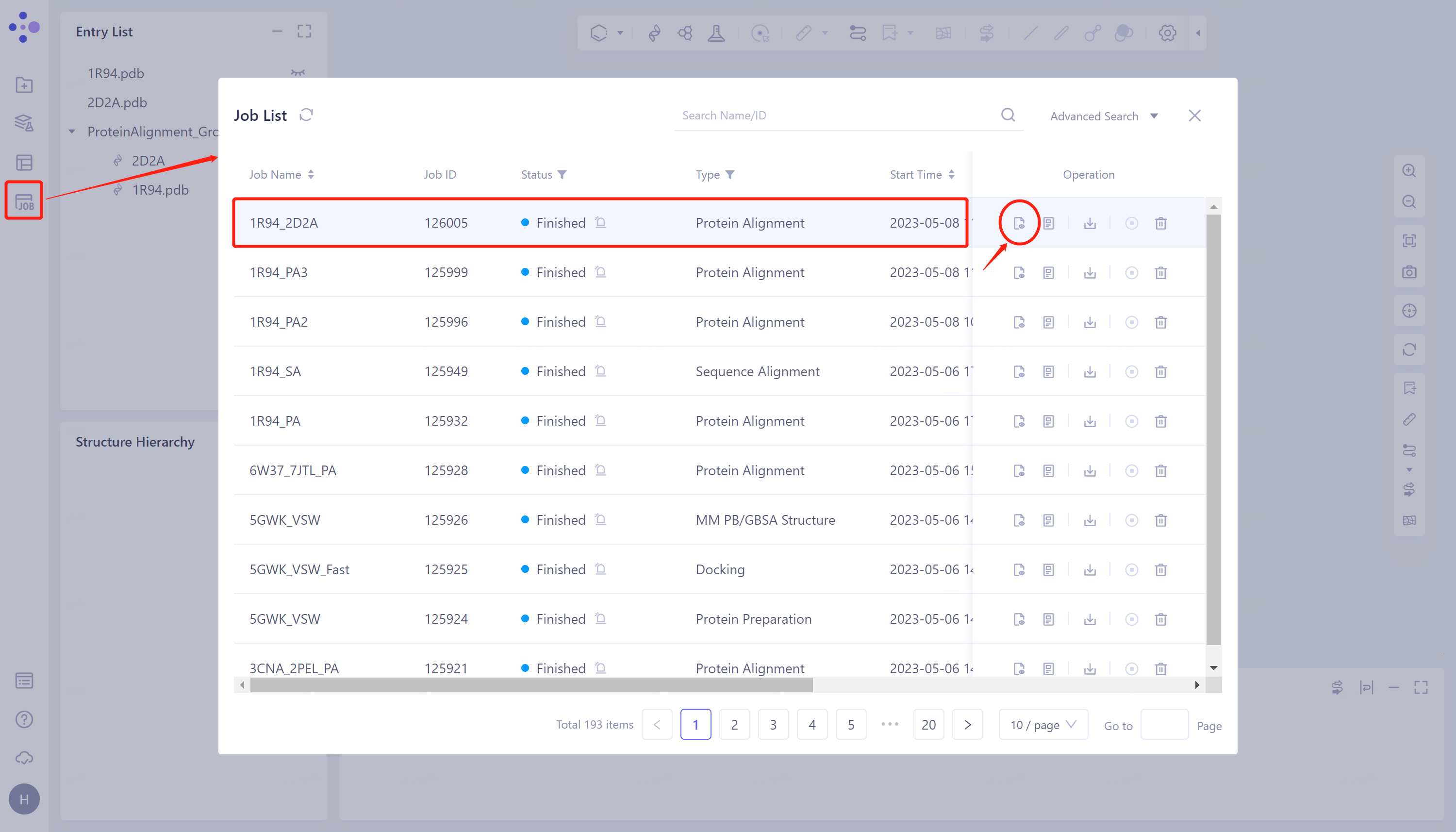
- General menu bar on the left: Job → Find the task '1R94_2D2A' → Click 'Show' in the Operation column to display the results of the task, as shown in the figure below.
 |  |
3.2 Description of results
-
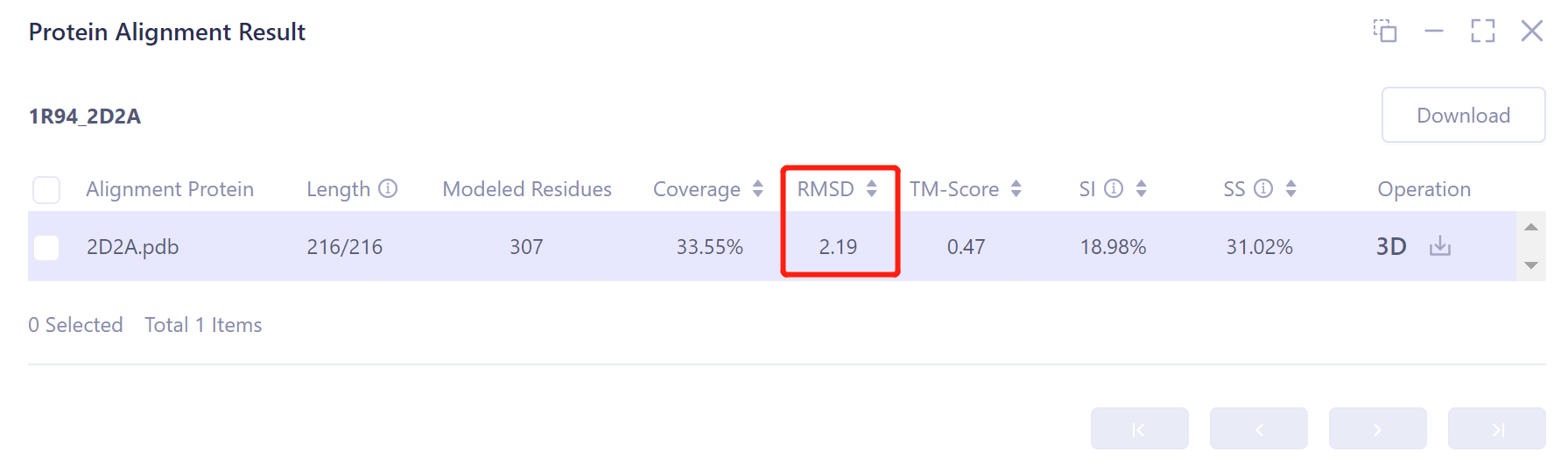
The result information of protein structure alignment is listed in the list box on the right, including:
-
Length: the number of amino acids in the alignment of the two proteins is 216;
-
Modeled Residues: the maximum length of the input structure is 307 amino acids;
-
Coverage: the coverage rate of comparison is 33.55%;
-
RMSD: The structural deviation of the aligned two protein conformations is 2.19 Å, indicating that the two protein structures are highly similar;
-
TM-Score: An index to evaluate the topological similarity of protein structures. The value of TM-Score is (0,1), where 1 represents a perfect match between two structures. The TM-Score value of the aligned two proteins is 0.47;
-
SI: The sequence identity of the two proteins was 18.98%, which was low;
-
SS: The sequence similarity of the two proteins was 31.02%.
-
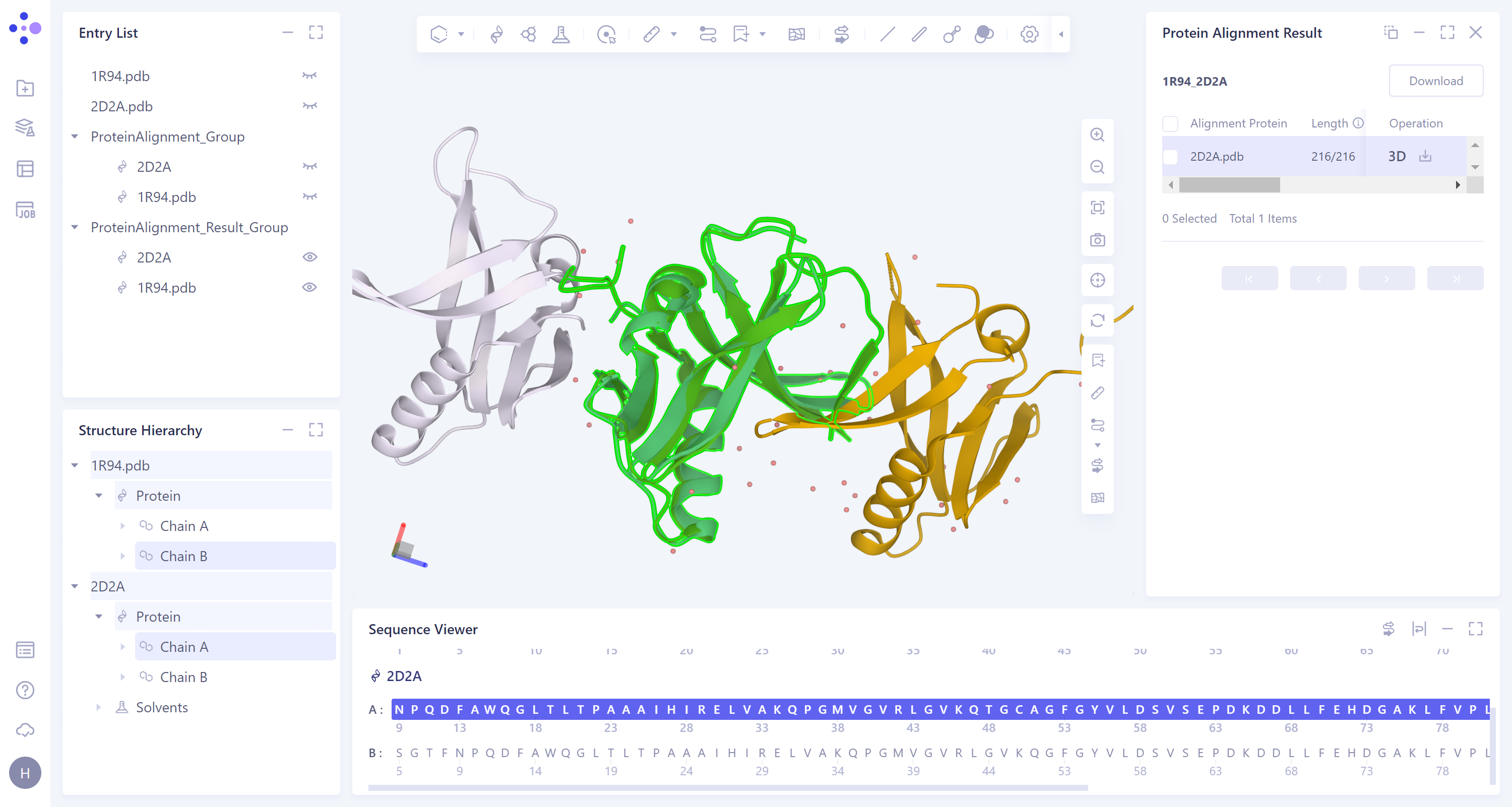
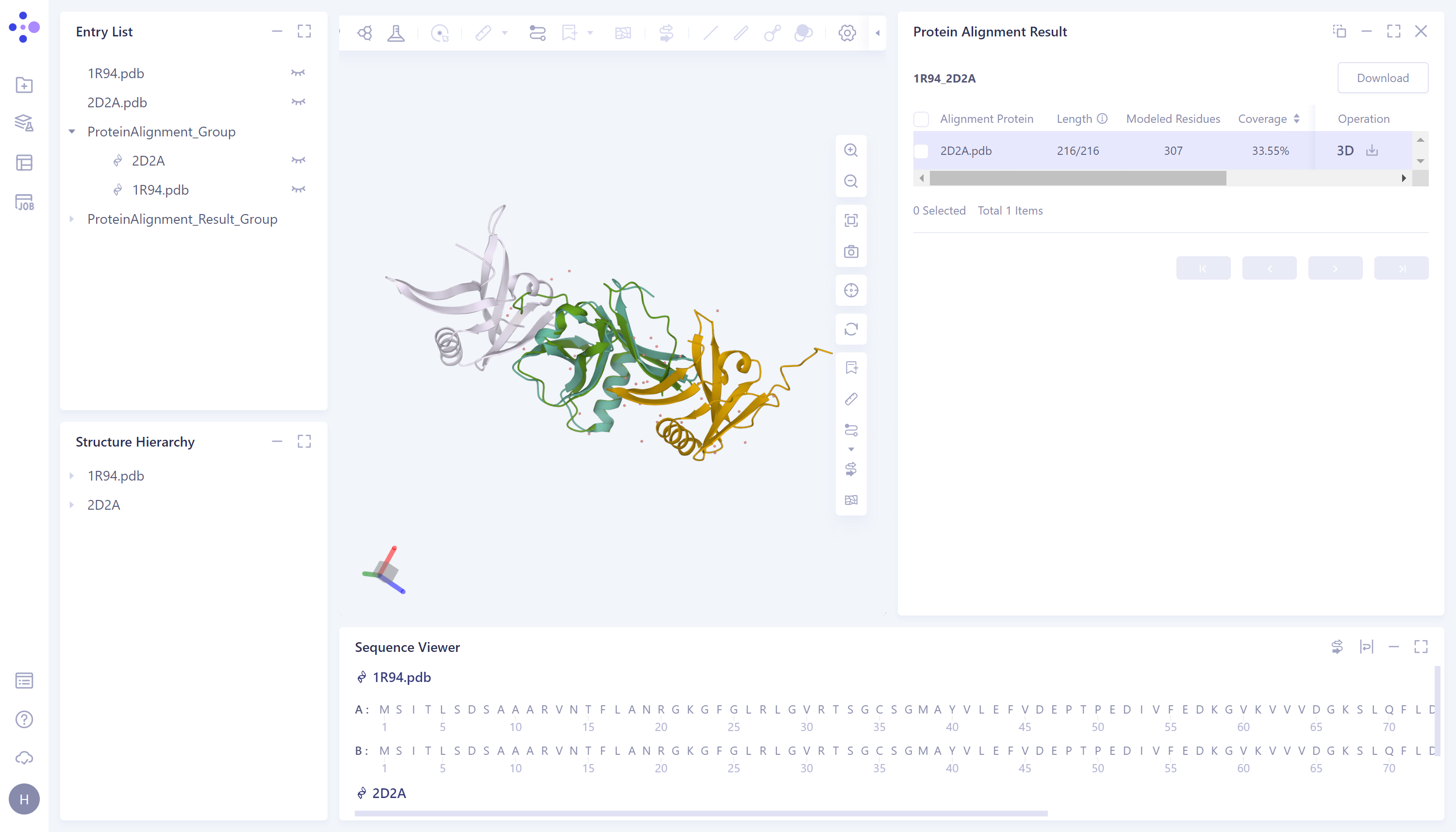
- The 3D workspce window displays the visual results of protein alignment. The chain B of '1R94' and the chain A of '2D2A' are aligned. It can be seen from the figure that the two structures can be well superimposed and have high structural similarity.