Prediction and Evaluation of Monomer Protein Structure Based on Uni-Fold
Introduction
For the first time, the Uni-Fold module in the Hermite ® platform successfully reproduces the full scale training of AlphaFold2, which can realize the structure prediction of protein monomers/polymers. In this tutorial, you will learn to use the Uni-Fold module to realize the structure prediction of protein multimers, and evaluate the accuracy of protein structure prediction by using the Protein Alignment module in Hermite to compare the predicted structure with the experimental structure.
4PYP is the crystal structure of the human glucose transporter GLUT1 at 3.2 Å resolution. The glucose transporter GLUT1 catalyzes the diffusion of glucose into red blood cells and is responsible for supplying glucose to the brain and other organs. Dysfunctional mutations may contribute to GLUT1 deficiency syndrome, and GLUT1 overexpression is a prognostic indicator of cancer. This tutorial takes 4PYP as an example to predict its structure based on the protein sequence and evaluate the accuracy of the results.
The data required for this tutorial is as follows:
> 4PYP_1|Chain A|Solute carrier family 2, facilitated glucose transporter member 1|Homo sapiens(9606)MEPSSKKLTGRLMLAVGGAVLGSLQFGYNTGVINAPQKVIEEFYTQTWVHRYGESILPTTLTTLWSLSVAIFSVGGMIGSFSVGLFVNRFGRRNSMLMMNLLAFVSAVLMGFSKLGKSFEMLILGRFIIGVYCGLTTGFVPMYVGEVSPTALRGALGTLHQLGIVVGILIAQVFGLDSIMGNKDLWPLLLSIIFIPALLQCIVLPFCPESPRFLLINRNEENRAKSVLKKLRGTADVTHDLQEMKEESRQMMREKKVTILELFRSPAYRQPILIAVVLQLSQQLSGINAVFYYSTSIFEKAGVQQPVYATIGSGIVNTAFTVVSLFVVQRAGRRTLHLIGLAGMAGCAILMTIALALLEQLPWMSYLSIVAIFGFVAFFEVGPGPIPWFIVAELFSQGPRPAAIAVAGFSNWTSNFIVGMCFQYVEQLCGPYVFIIFTVLLVLFFIFTYFKVPETKGRTFDEIASGFRQGGASQSDKTPEELFHPLGADSQVLEHHHHHHHHHH
1. Protein Folding
1.1 Entrance
- The left general menu bar 'Function' → 'Structure Modeling' → 'Protein Folding'.
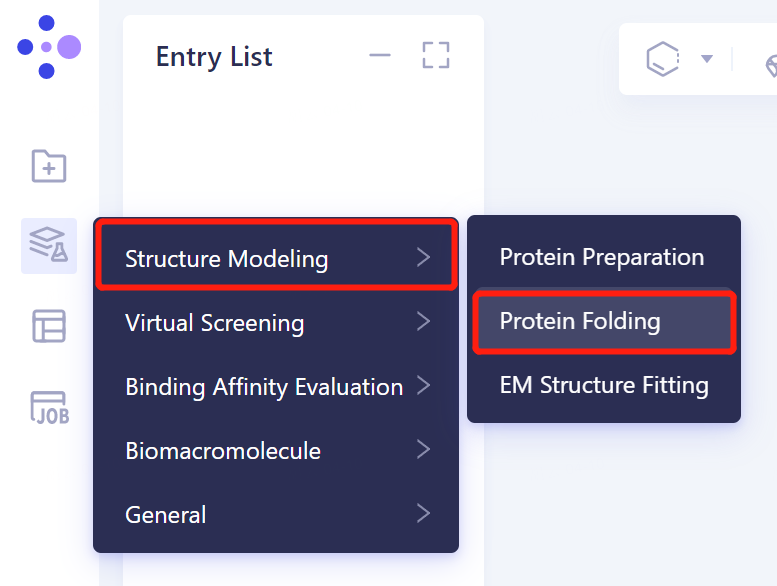
1.2 protein sequence introduction
-
Enter the protein sequence in the Protein Folding interface (shown in the red box) that appears on the right. Paste the sequence of 4PYP into the input box of Chain A.
-
After naming the task as 4PYP_Uni-Fold at the 'Job Name', click 'Submit' to submit the task.
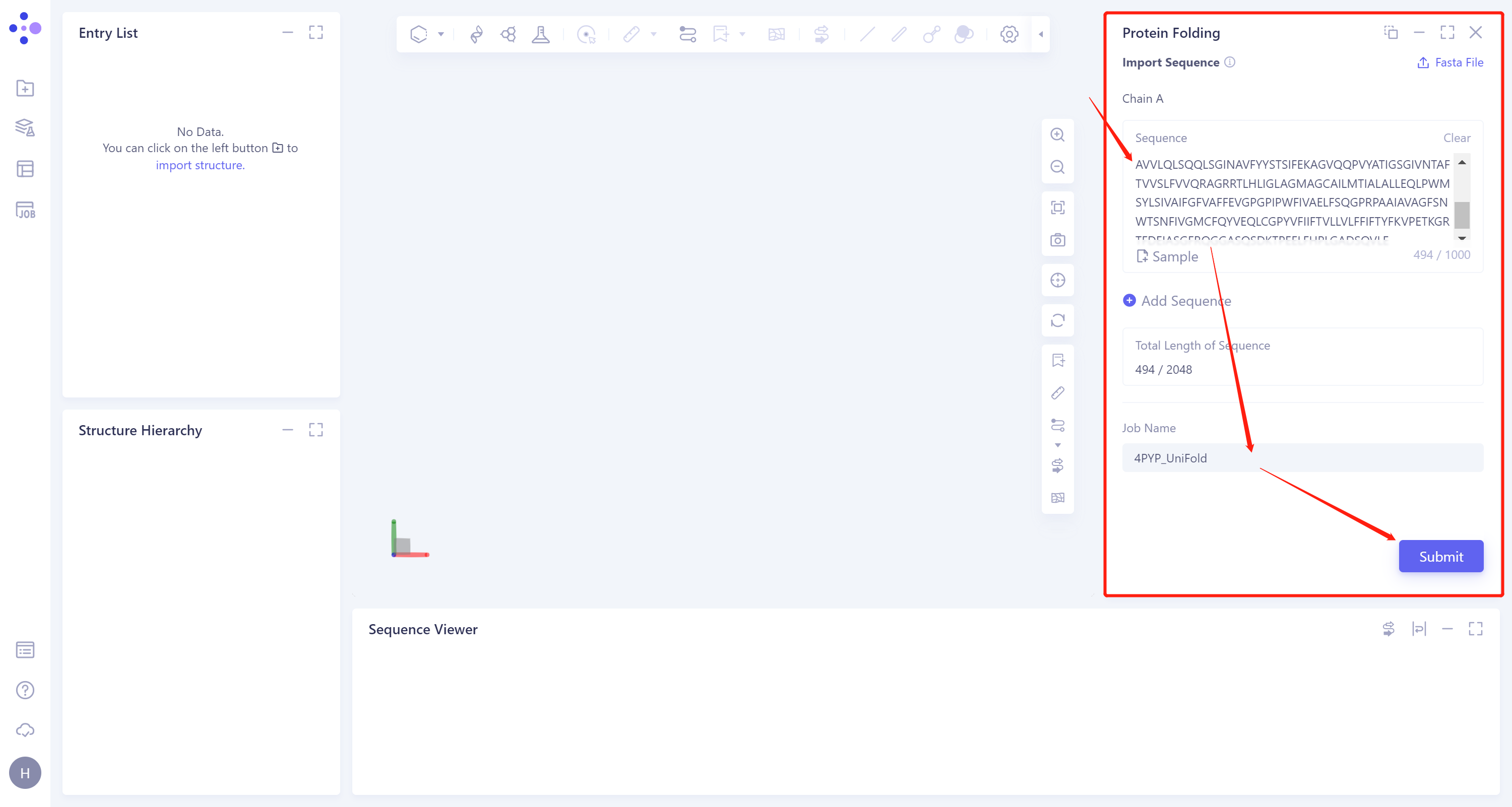
1.3 Results View
- Click 'Job' in the left general menu bar → find the 4PYP_Uni-Fold task in the pop-up 'Job List' interface → click 'Show'.
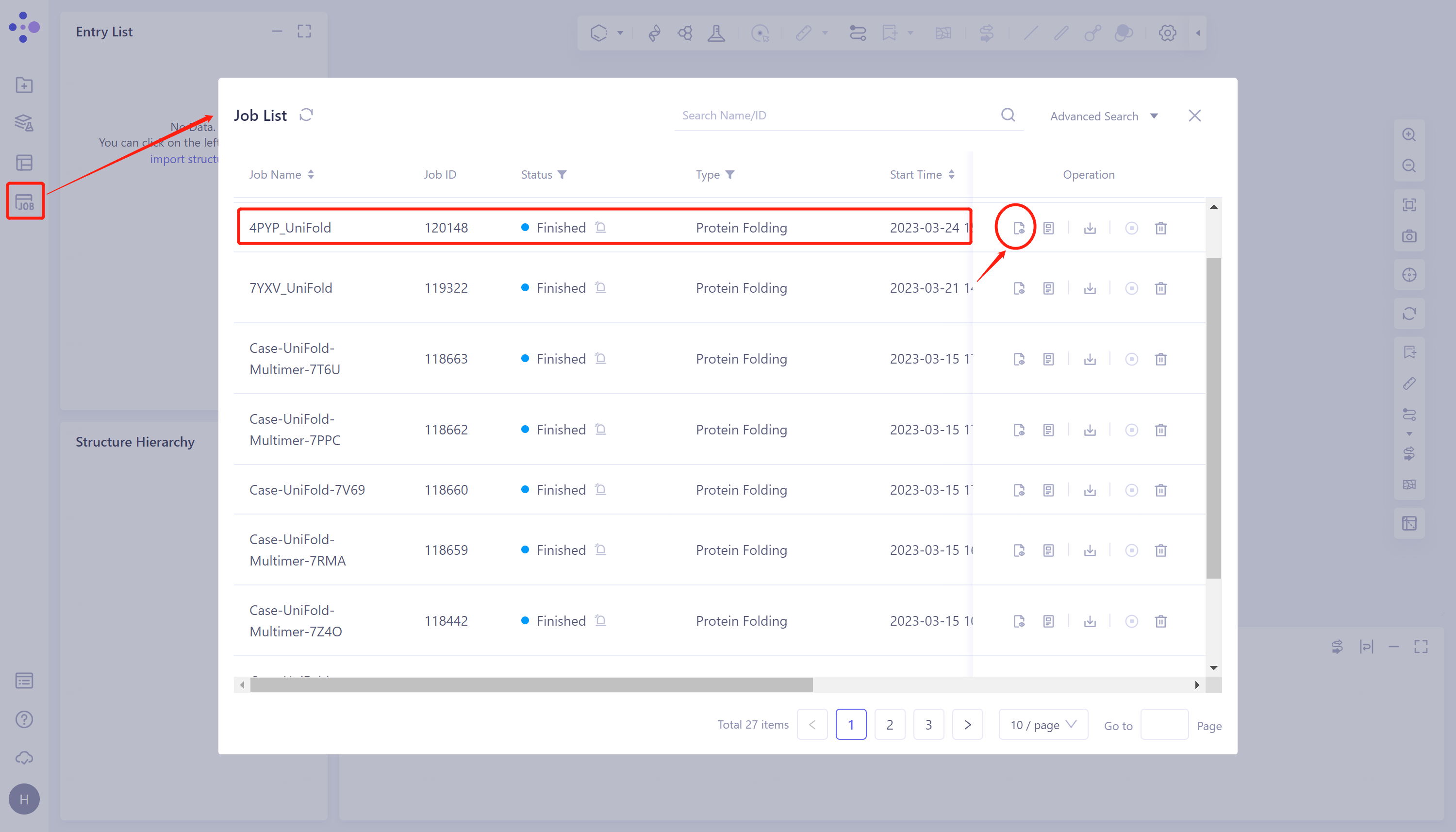
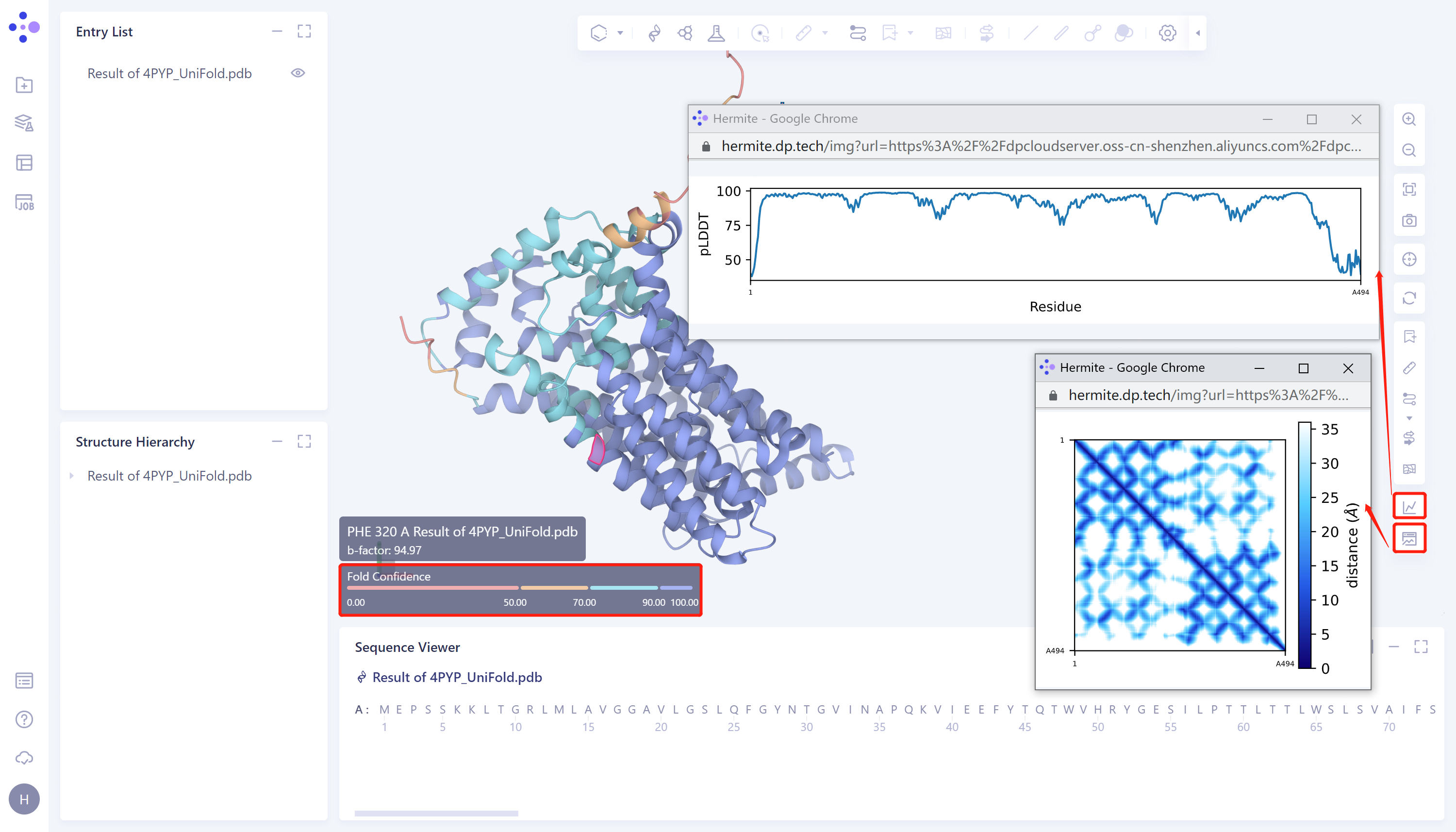
- Protein prediction results were displayed in the 3D Workspace interface, and the pLDDT score and distance map of the predicted Uni-Fold_4PYP structure showed that the structure had high confidence. The higher the pLDDT score, the more accurate the structural prediction.
2. Protein Alignment
2.1 Import protein structures into the 3D Workspace window
- In the same Project, click 'File' → 'Get PDB' on the left general menu bar.
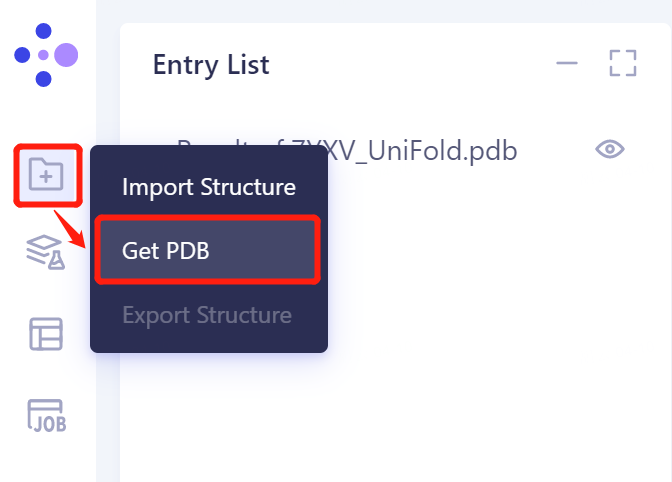
- Enter 4PYP in the pop-up window and click 'Import' to import the protein structure.

2.2 Invoke Protein Alignment module
2.2.1 Inlet
- Invoke the 'Protein Alignment' module to align the protein structure: 'Function' → 'General' → 'Protein Alignment' in the left general menu bar.
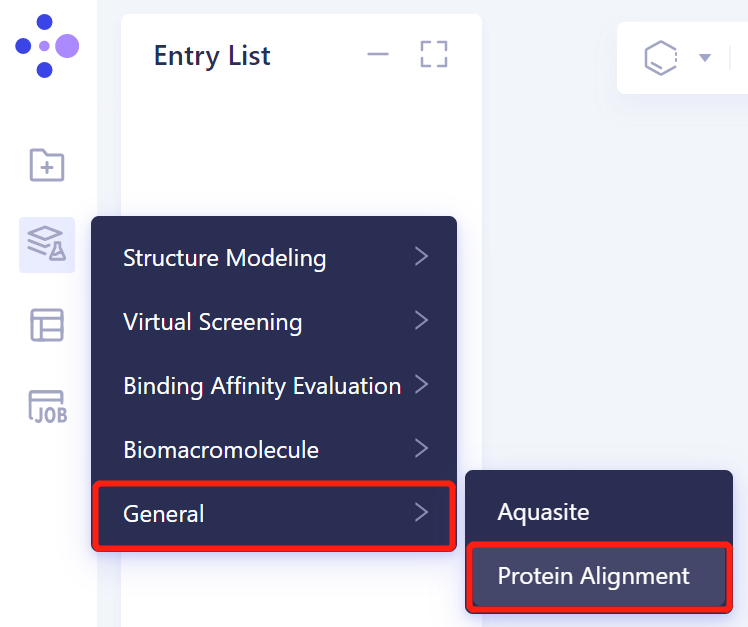
2.2.2 Introduction of reference protein structures
- The Protein Alignment operation window appears on the right side of the interface. First, import the reference protein structure: click Select Structure, The Select Structure window pops up → Select the 4PYP protein structure in the Structure Hierarchy → Click 'OK' in the Select Structure window.
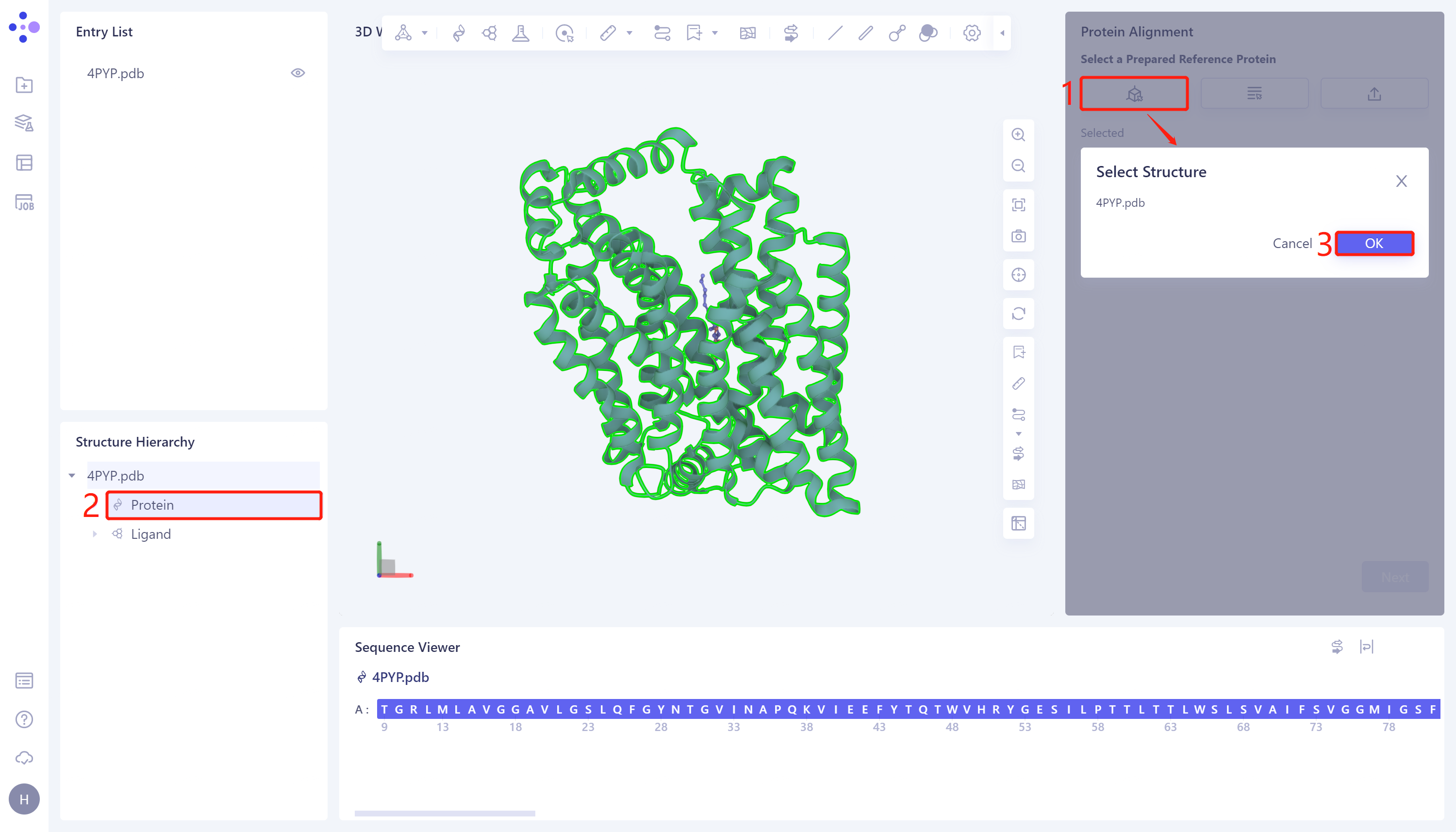
2.2.3 introduction of protein structure to be compare
- Click 'Next' to enter the selection window of protein to be compared and import the protein structure to be compared: click 'Select from Project', The 'Select from Project' window pops up → Select 4PYP_Uni-Fold in the 'Select from Project' window → Click 'OK'.
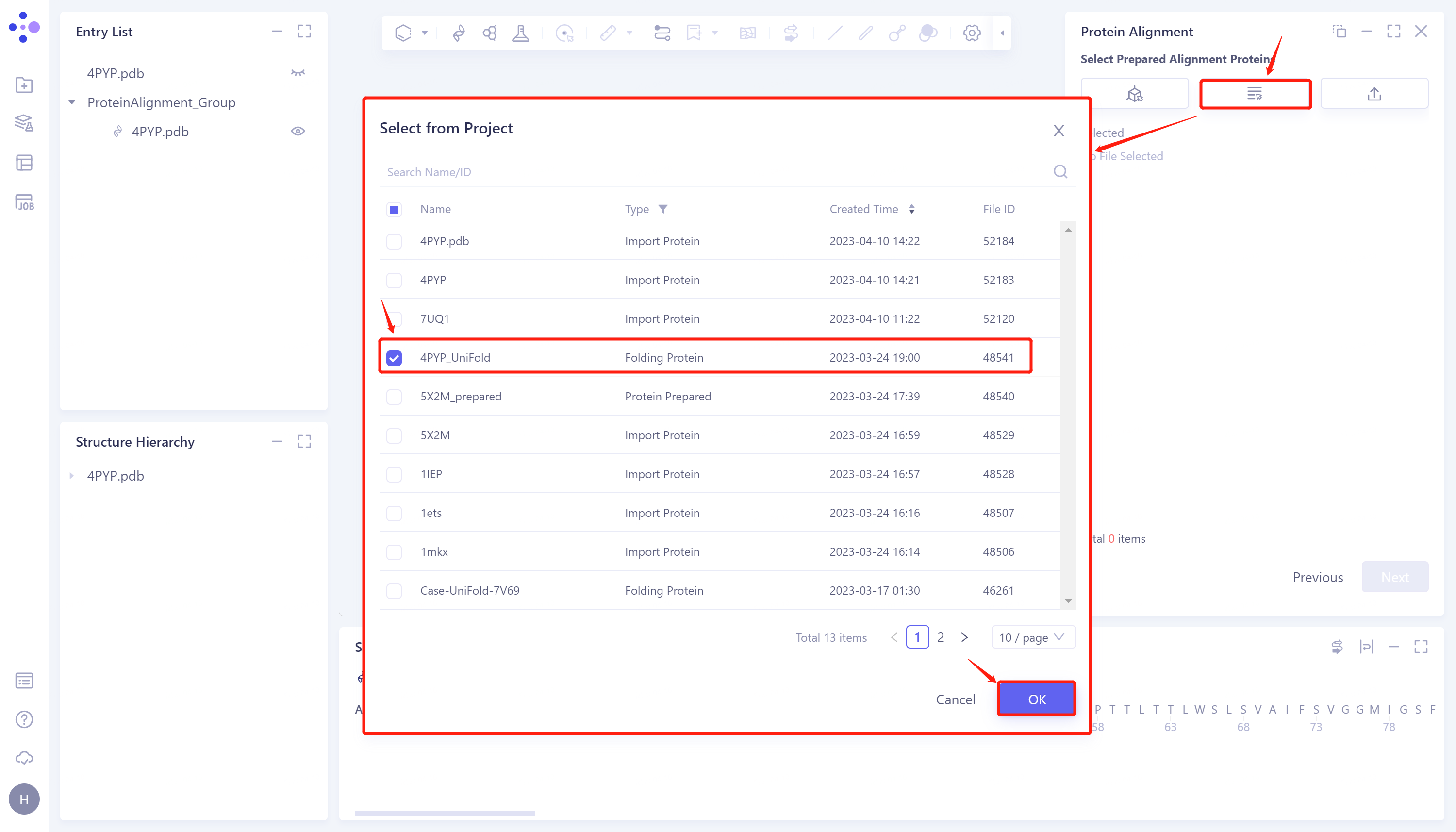
2.2.4 Parameter setting
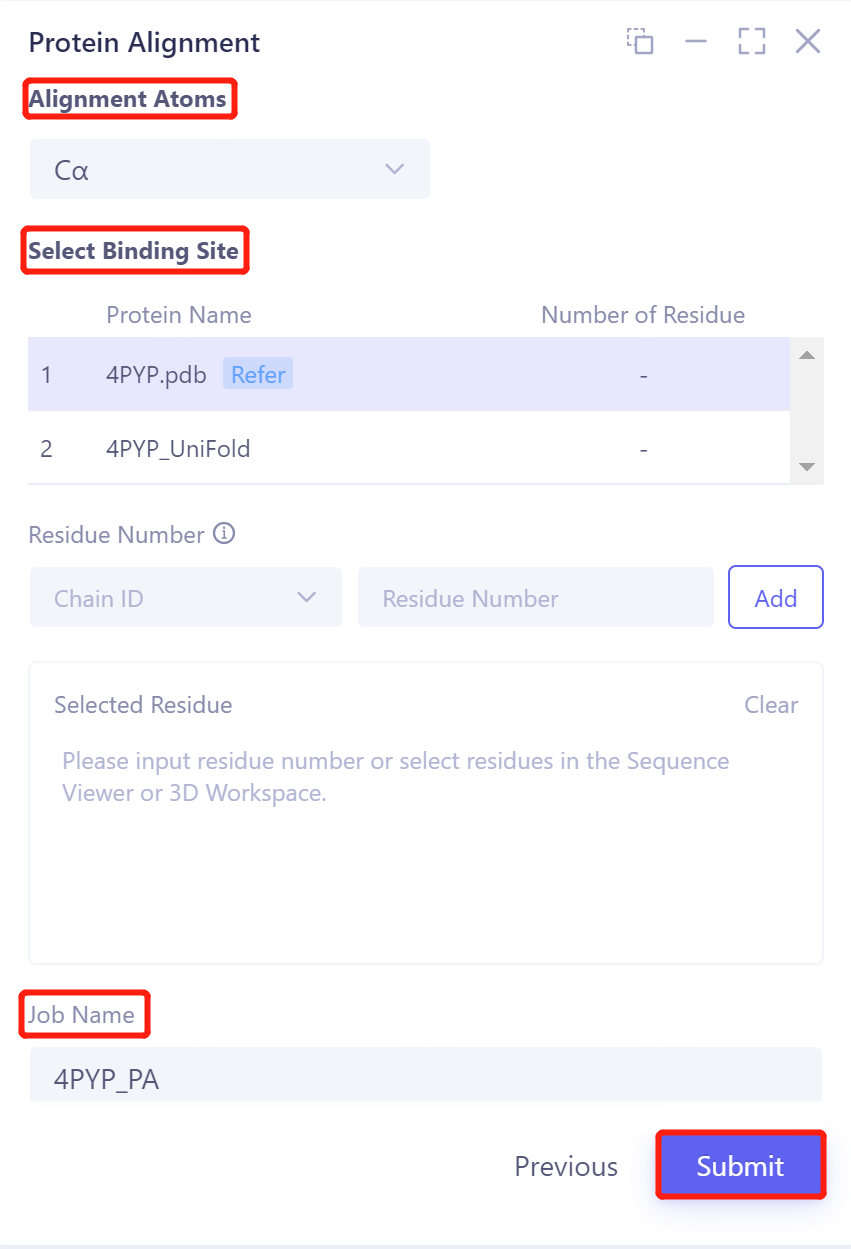
- Click 'Next' to enter the parameter setting page. Alignment Atoms selects Cα to perform alignment based on the α-carbon atoms between proteins. Since there is no special comparison requirement, the parameter setting of Select Binding Site is skipped. The Job Name field names the task as 4PYP _ PA, and click Submit to submit the task.
2.3 Analysis of results
2.3.1 Inlet
- General menu bar on the left Job → Find the task 4PYP _ PA in the pop-up Job List window → Click Operation Show under Operation to display the task.
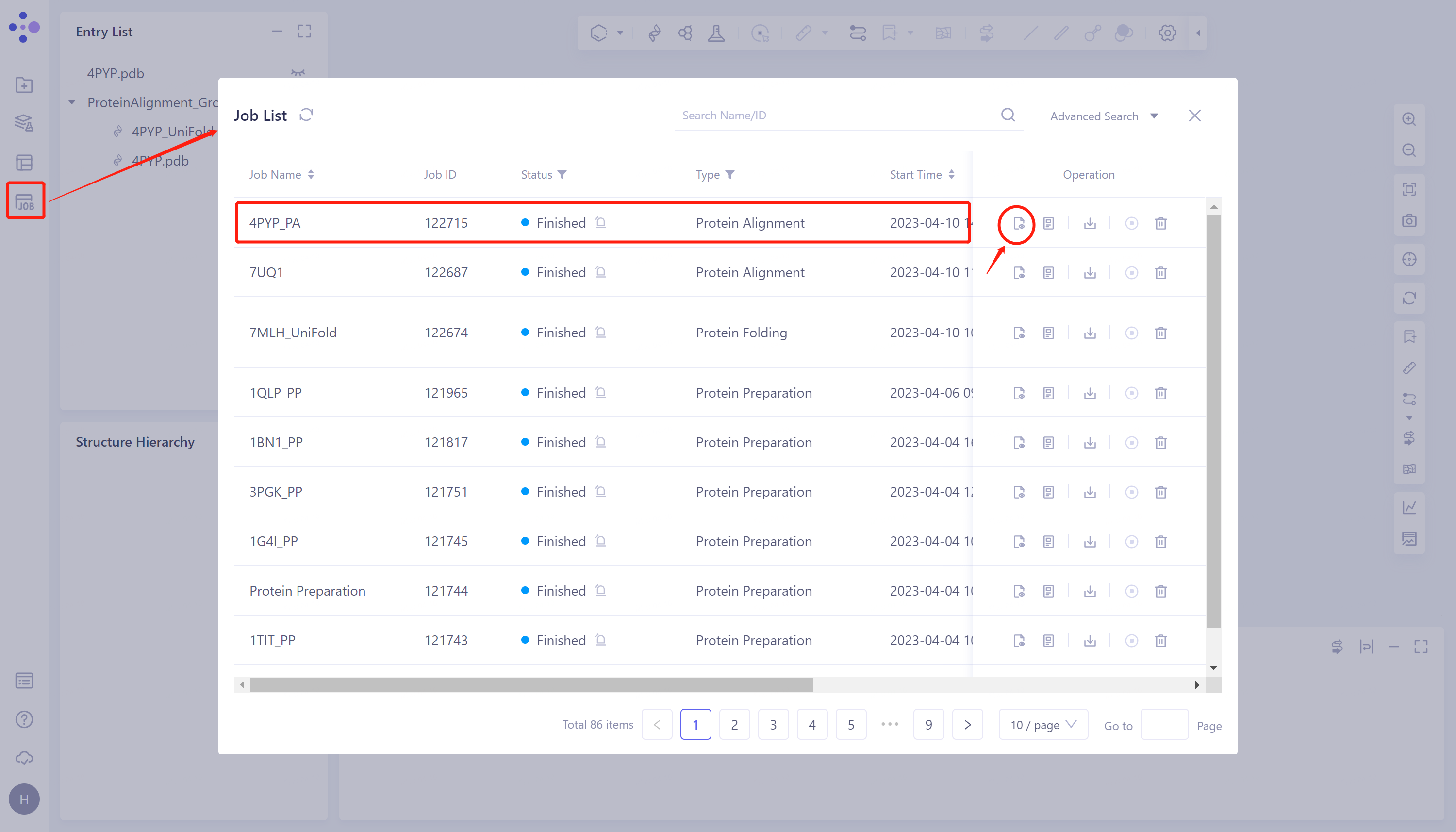
2.3.2 Presentation and explanation of results
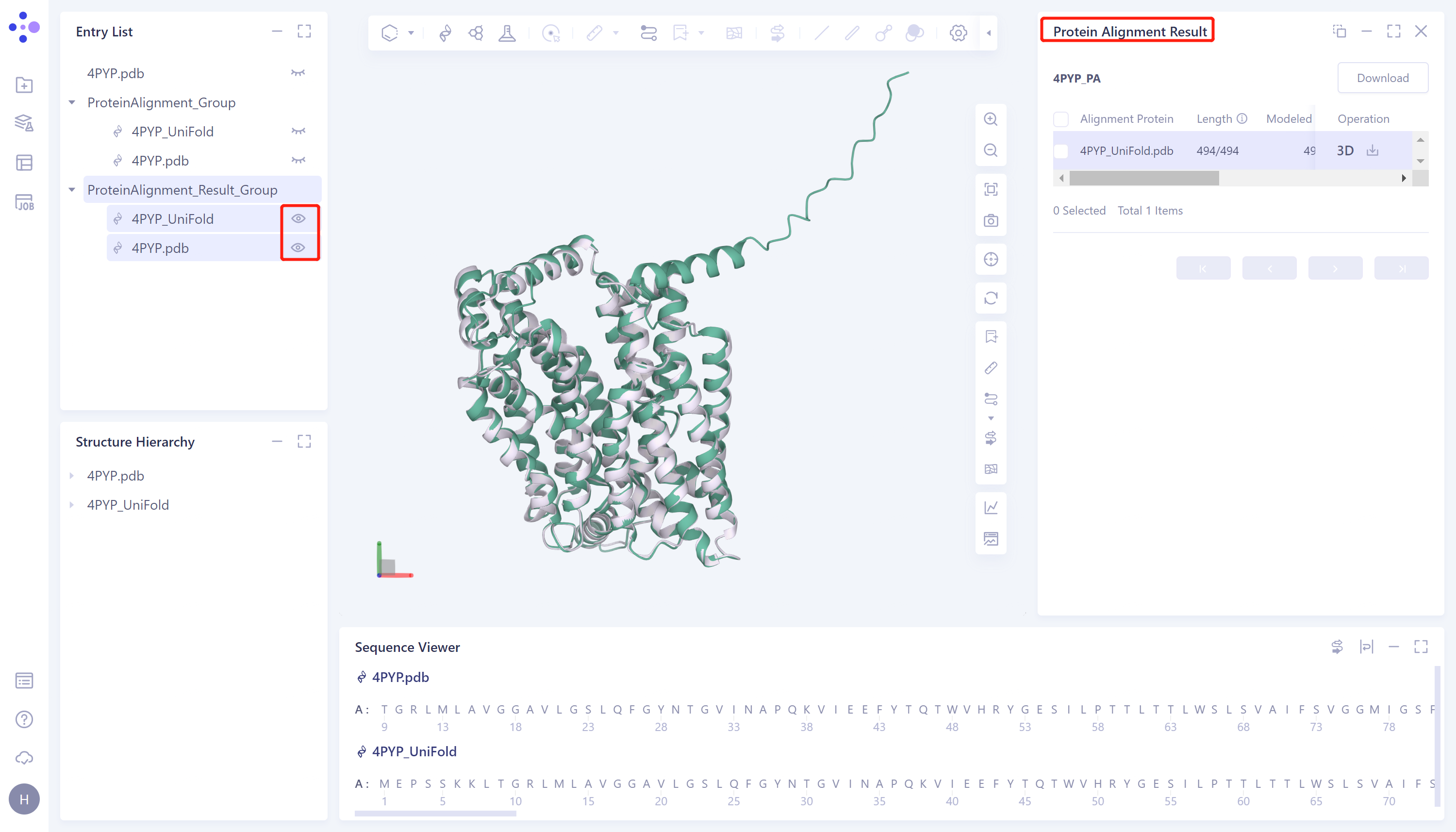
- The overlay image result of protein structure is displayed in the 3D Works pace window, and the Protein Alignment Result window appears on the right side of the interface to display the protein alignment result.
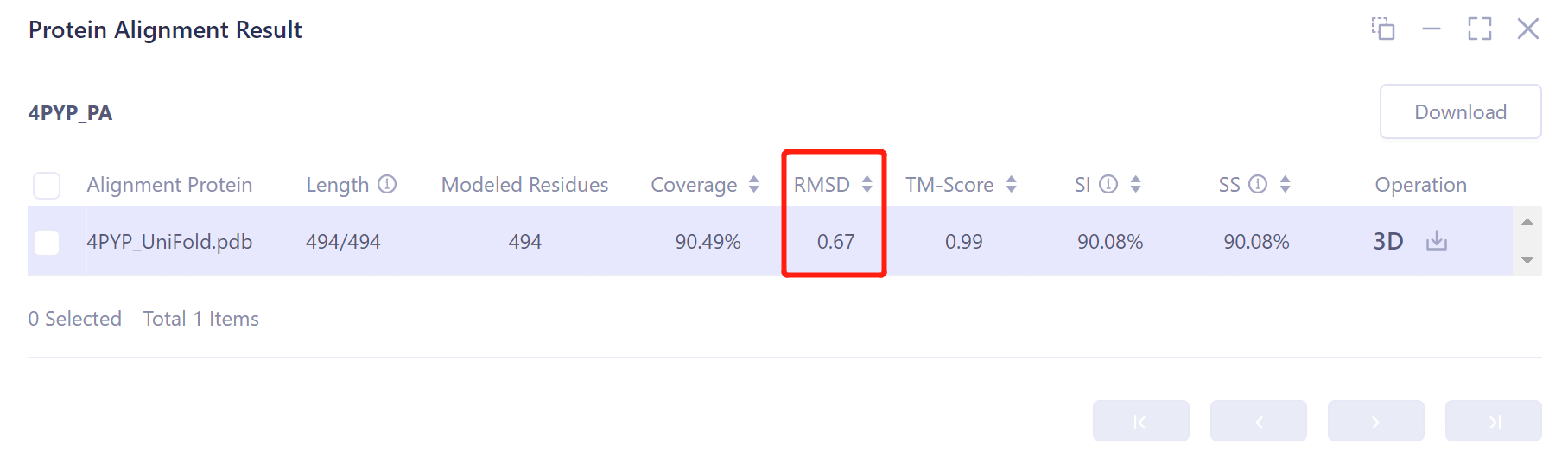
- Through protein structure comparison, it was found that the RMSD value of Uni-Fold predicted the structure of monomer protein 4PYP was 0. 67 Å, and the similarity was high, indicating that Uni-Fold could well predict the protein structure of 4PYP.
 P
P