Binding Stability
Introduction
The process of binding small molecule drugs to proteins is called Binding. Small molecules stably bind to proteins with a certain Binding Pose. Understanding the Binding Pose of protein- ligand complexes at the atomic level can provide us with drug design ideas in the optimization stage of drug development. Therefore, it is particularly important to use molecular dynamics (MD) to verify the stability of small molecule Binding Pose. The Binding Stability module of the Hermite platform provides a method based on molecular dynamics simulation to measure the stability of small molecule Binding Pose in small molecule-protein complexes.
The meaningless fusion of BCR gene and abl gene leads to the formation of active tyrosine kinase (Bcr-Abl), which transforms cells and leads to chronic myeloid leukemia. Bcr-Abl small molecule inhibitors bound to the kinase domain can be used to treat chronic myeloid leukemia. In this case, we used the eutectic structure (PDB: 1IEP) of the Bcr-Abl small molecule inhibitor imatinib (STI-571, hereinafter referred to as STI) and the c-Abl kinase domain [1] to study the stability of the small molecule Binding Pose.
1. Binding Stability
1.1 Entrance
Left Common Menu Bar Function → Virtual Screening → Binding Stability.
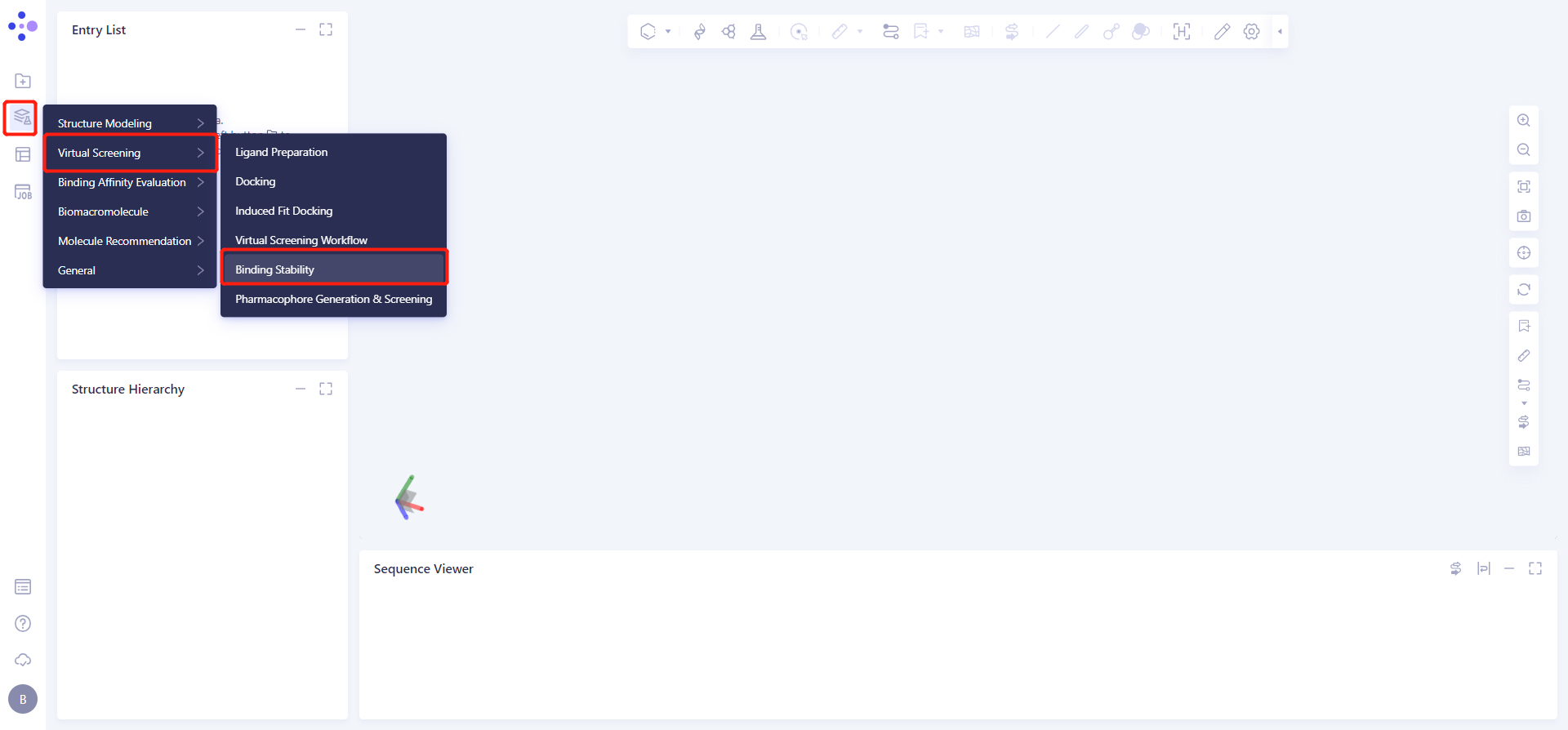
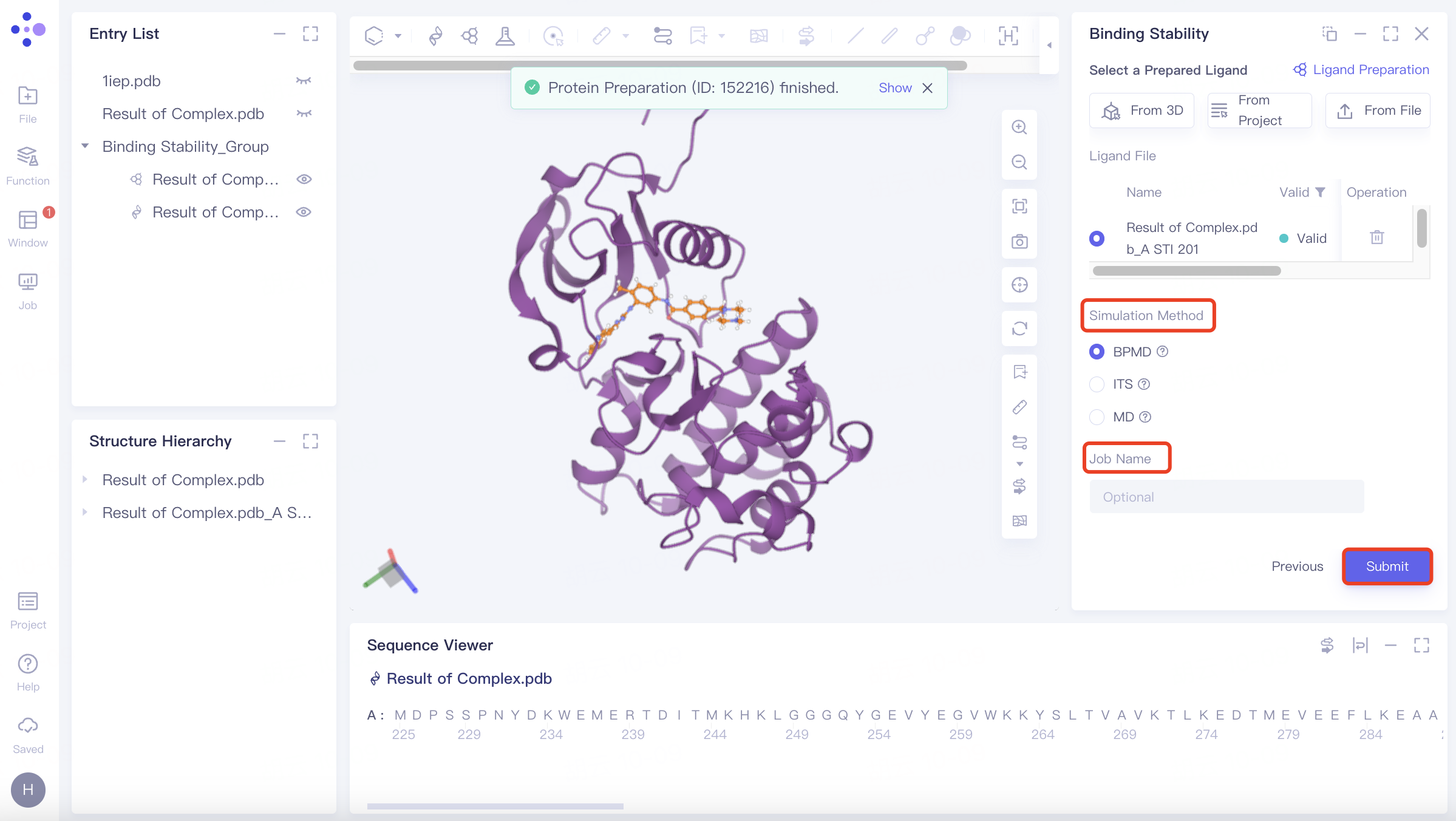
The Binding Stability operation box (shown in red) appears on the right side, and the overall interface is as follows:
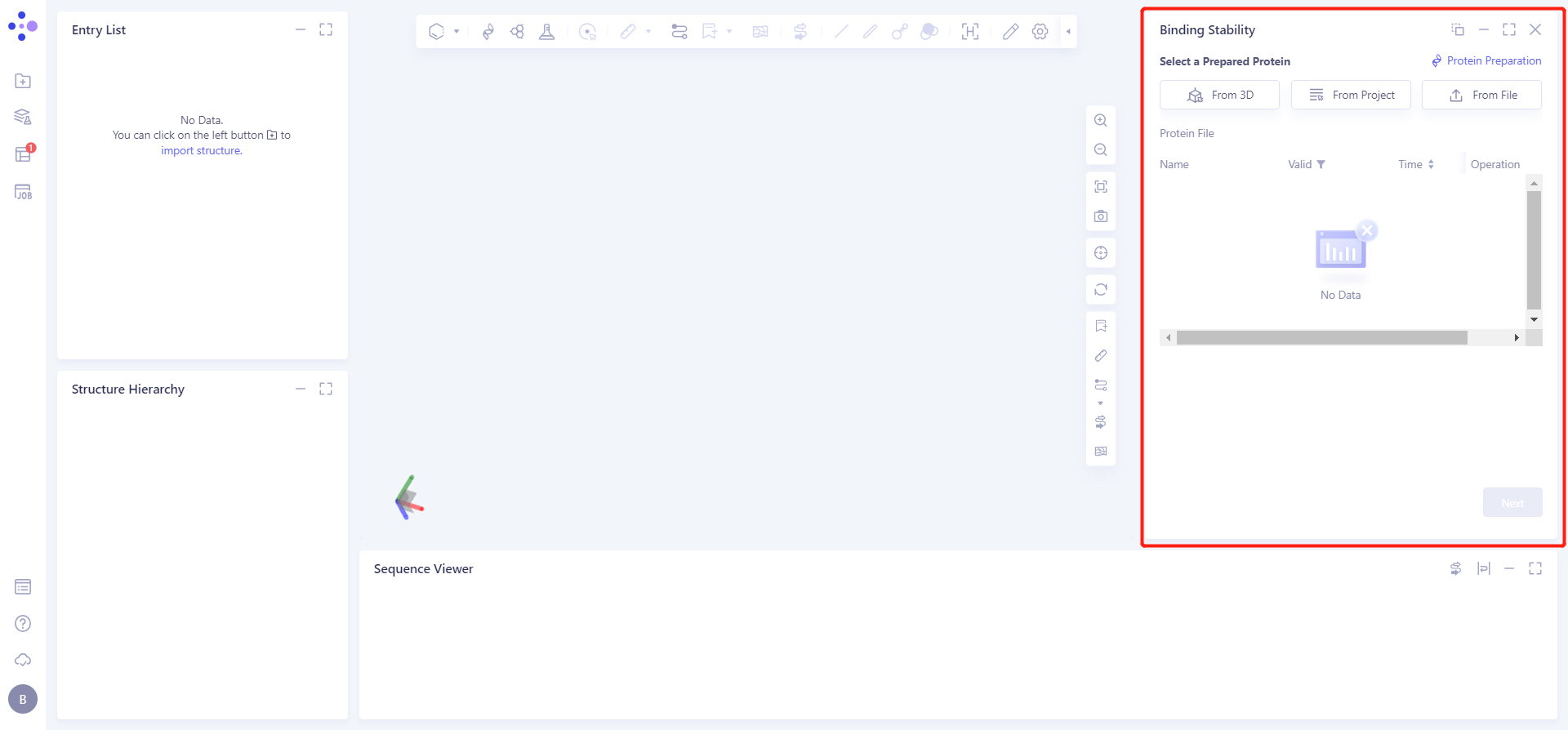
1.2 Upload protein structure
Click the Select File checkbox → Select the desired protein file 1iep_pdb_prepared from the local folder and upload it.
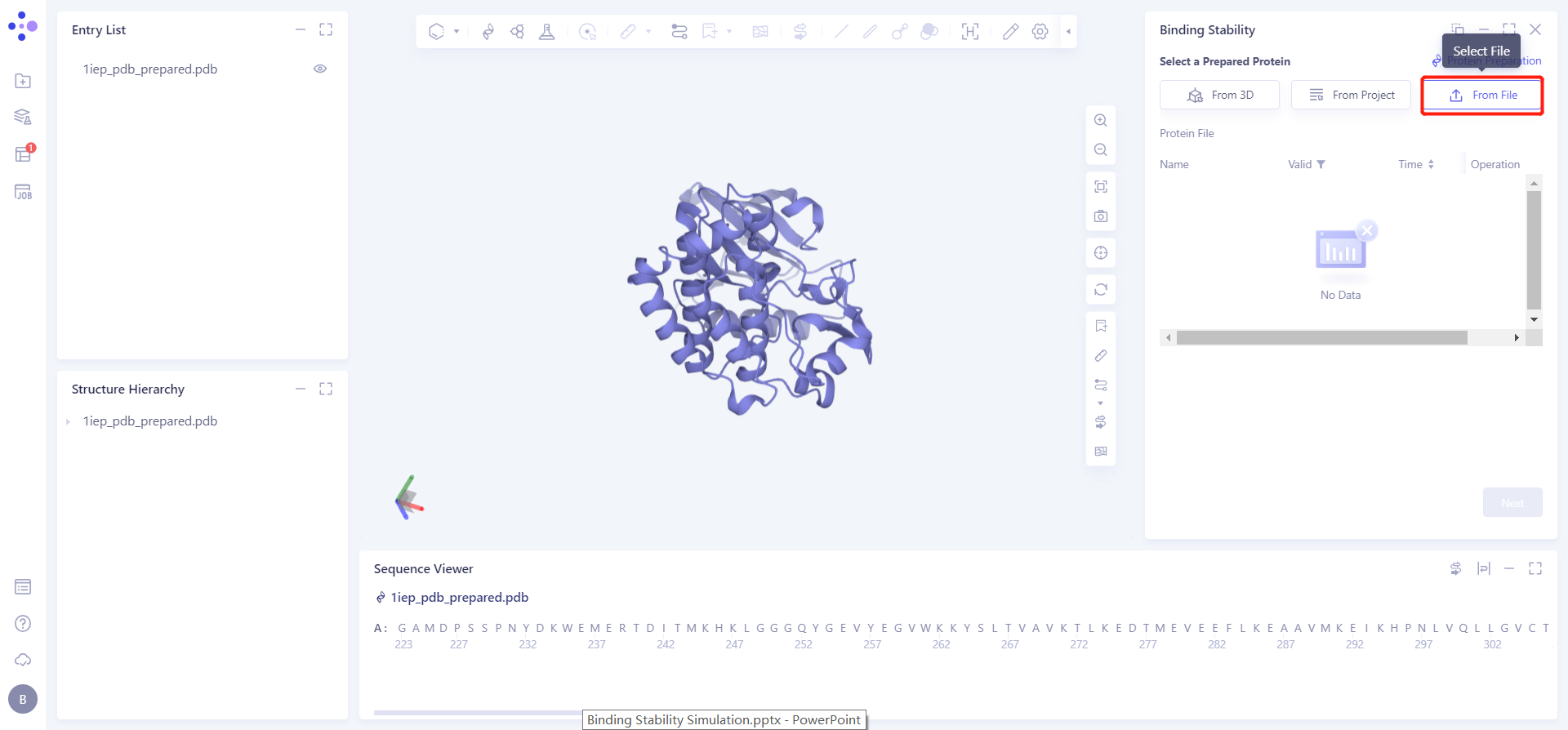
1.3 Upload ligand structure
Click the Select File checkbox → Select the desired ligand file 1iep. pdb_A STI 571.sdf from the local folder and upload it.
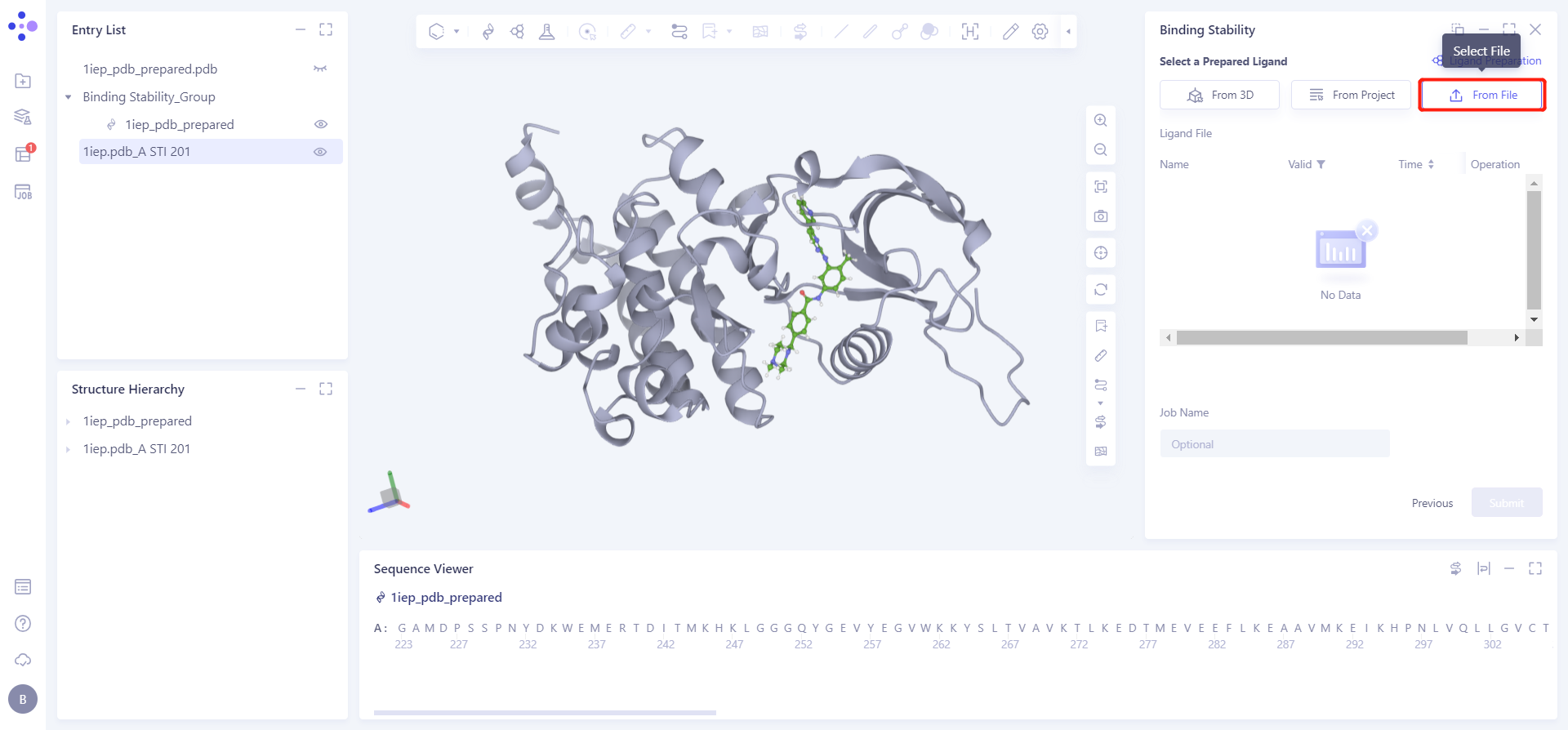
1.3.3 Parameter setting and task submission
Simulation Method Select BPMD;
BPMD: Binding pose meta dynamics, an improved sampling method that can effectively evaluate the stability of protein-bound ligands in solution, was selected to evaluate binding stability.
Click "Submit" to submit the task.
2. Result analysis
2.1 Entrance
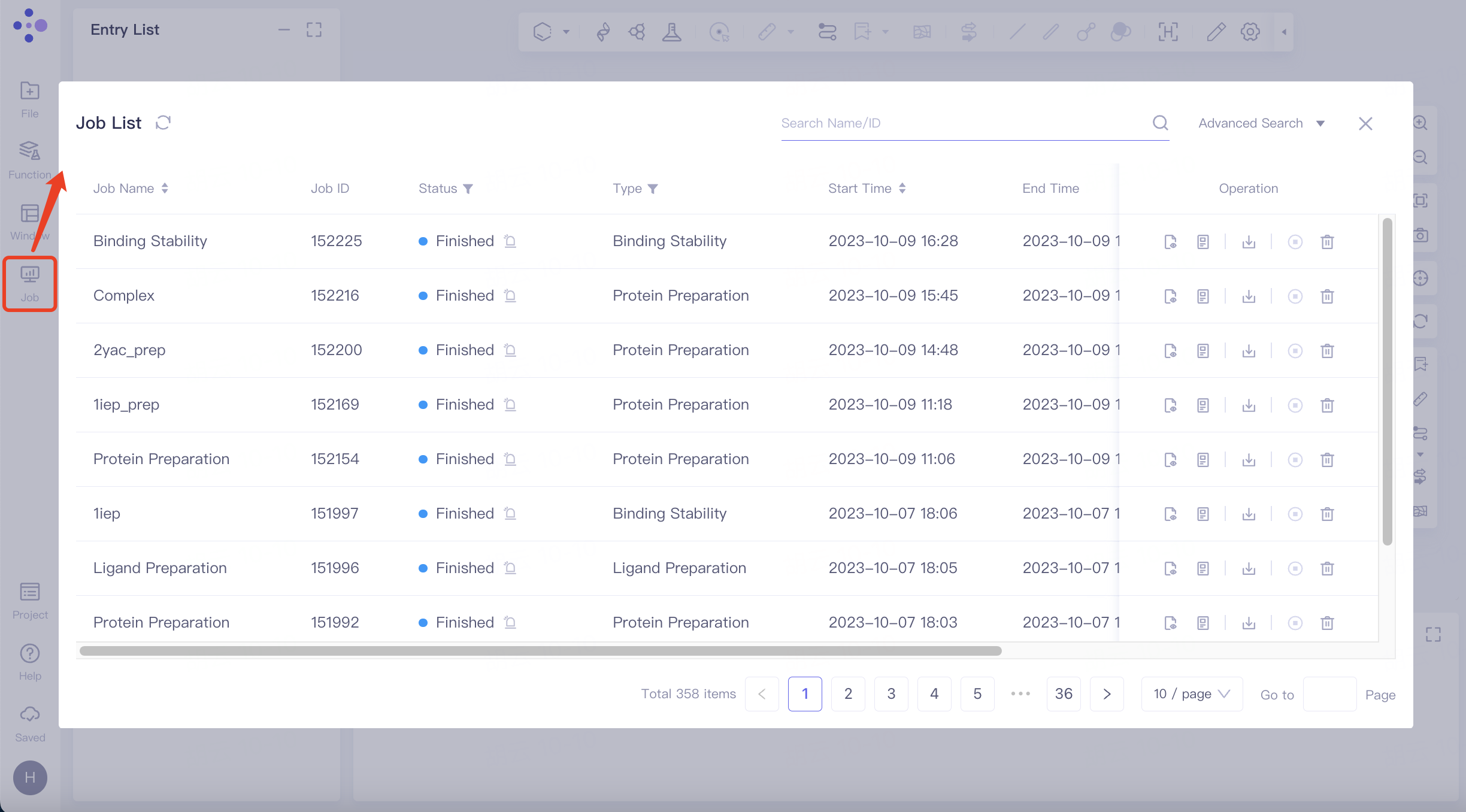
The left general menu bar Job → Find the required task.
The task can be found by searching for Job Name or by filtering through Job Type.
2.2 Results
2.2.1 Dynamic trajectory video
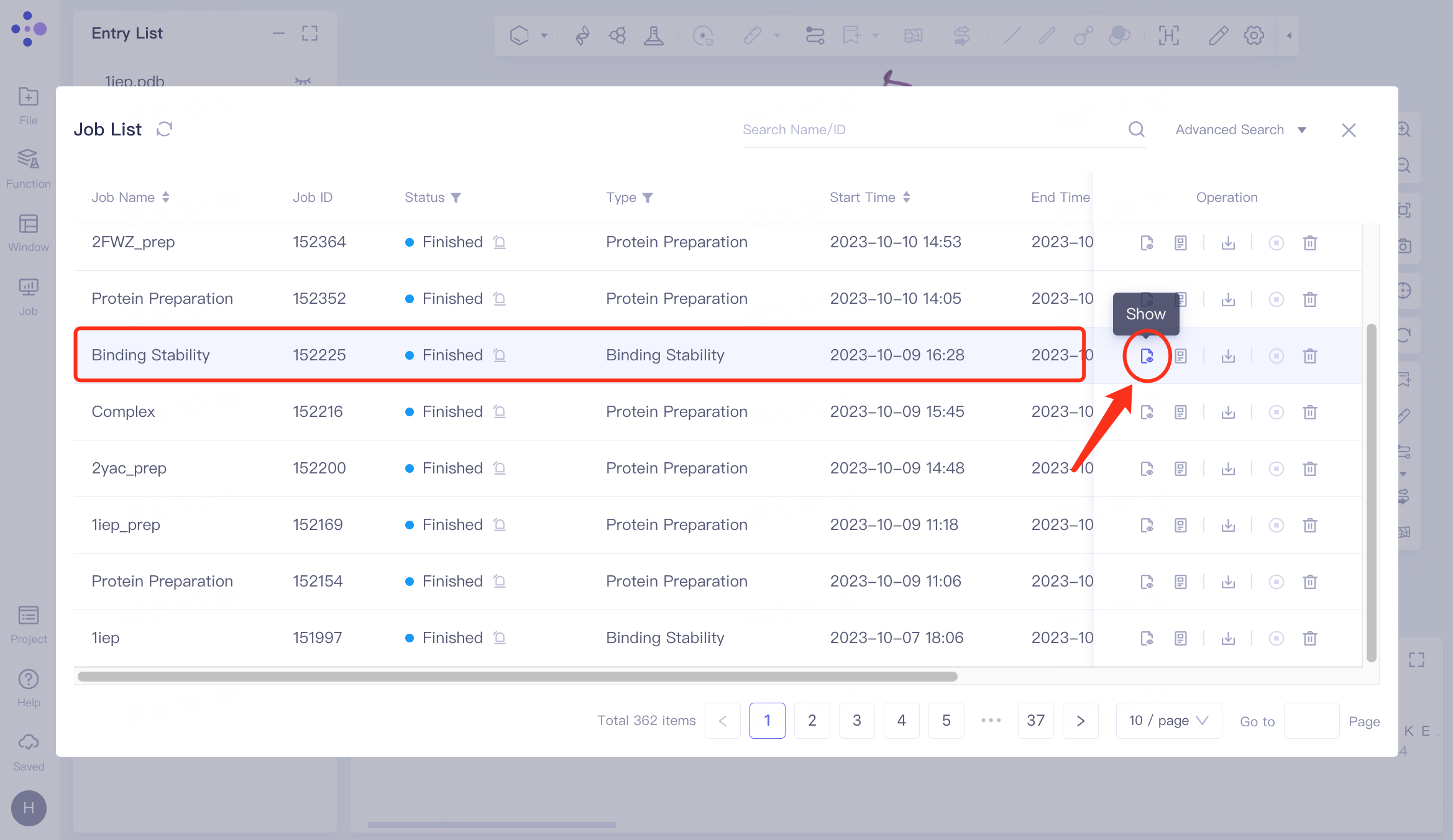
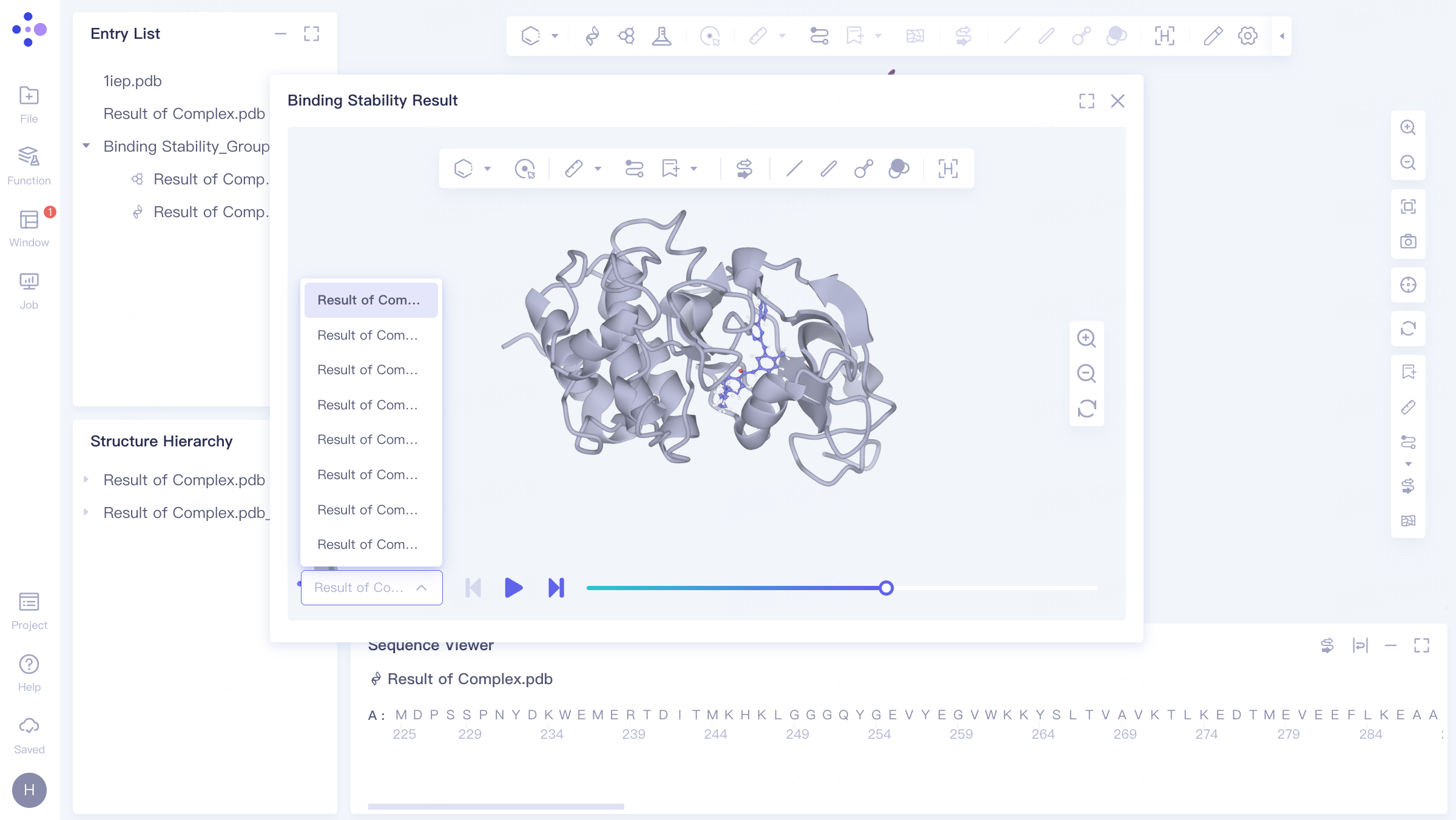
Select the task you want to view, and click Show in the Operation column to display the result of the task. The interface is shown in the left figure.
The video on the right shows the dynamic trajectory of Binding Stability , which can be switched to display 10 trajectory videos.
 |  |
2.2.2 Download result description
Download file
Select the task to be downloaded, and click Download in the Operation column to download the result of the task. The interface is shown in the figure.
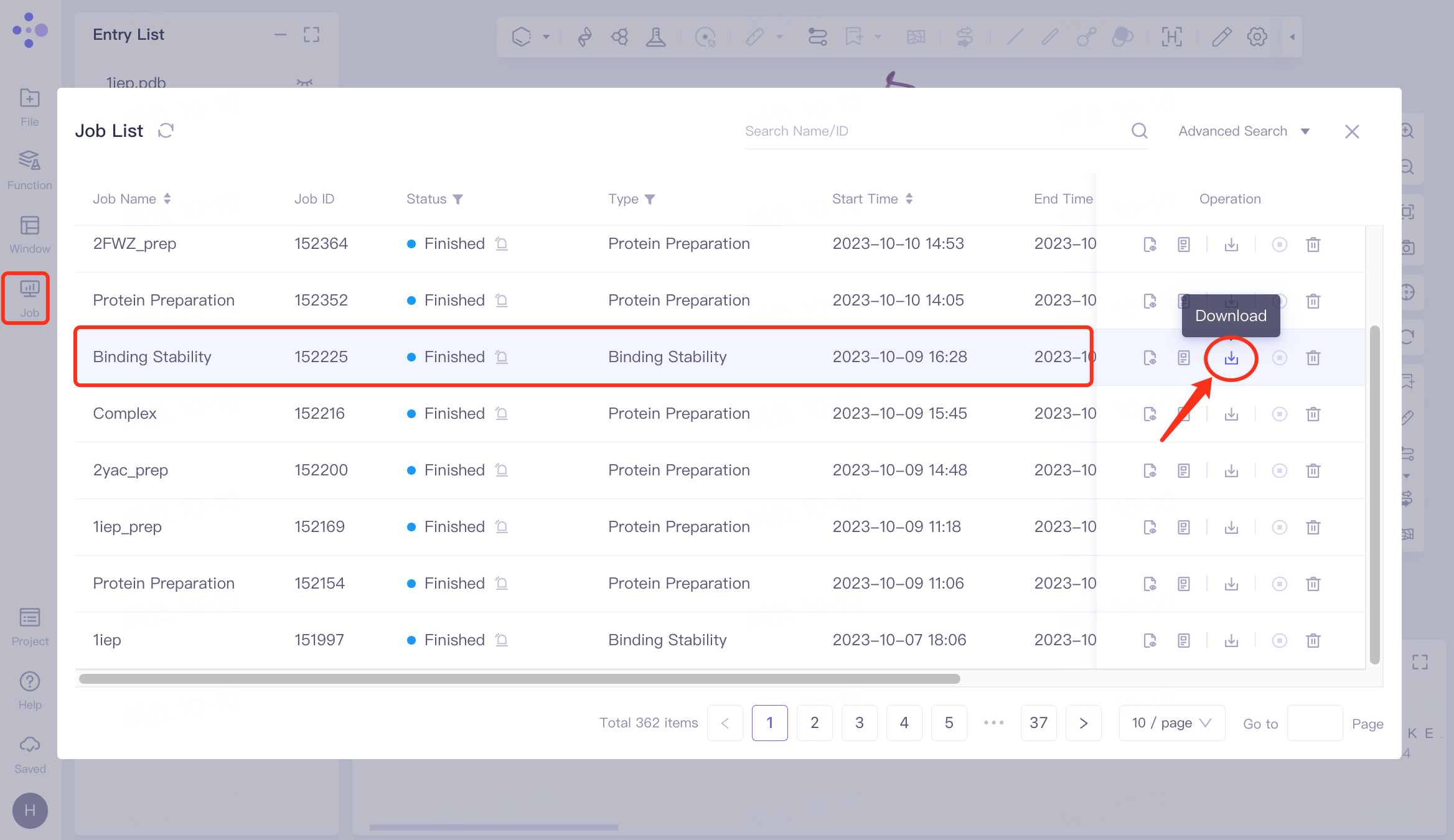
Results analysis :
- Combination stability :
results.csv: In this case, PoseScore is less than 2 - and CompScore is less than 0, indicating that the ligand binding to the protein is conformationally stable during kinetic processes.
fig_MOL_BPMD.png file: Change chart of PoseScore, ContactScore, and CompScore under 10 ns duration. In this example, all three evaluation indicators have good convergence in the dynamic process, indicating high reliability of the results.
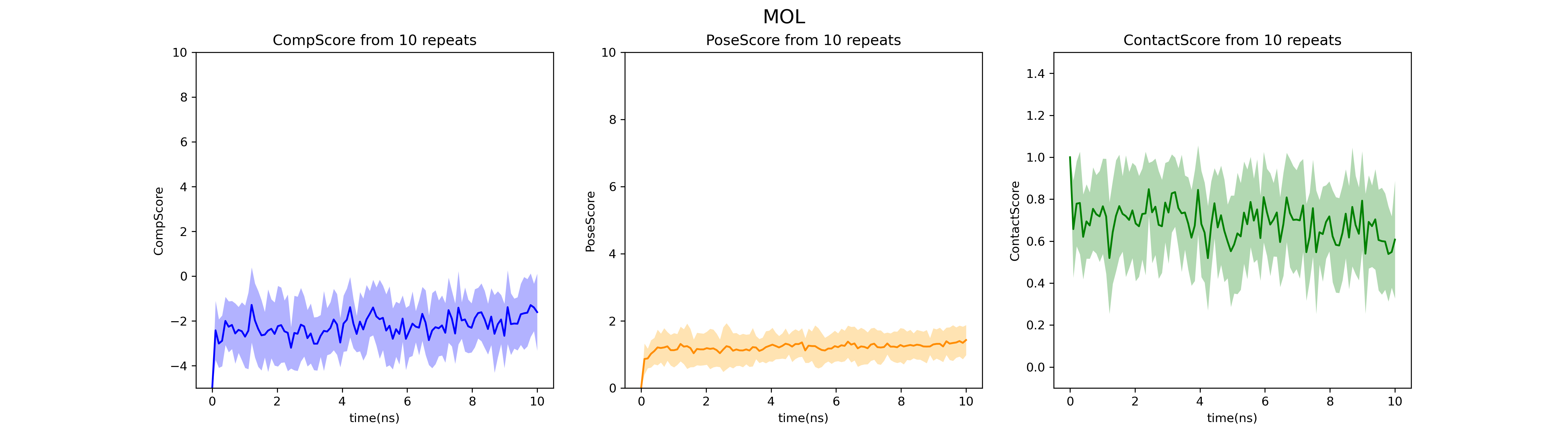
Notes:
PoseScore: The average value of the root mean square deviation (RMSD) between the atom and the initial atom in 10 simulations.
ContactScore: The ratio of the number of interactions between the conformation per 100 ps and the previous 1 ns conformation is recorded as the ContactScore 'at each time. The ContactScore value in this table is the average of the ContactScore' in the last 2 ns of 10 simulations.
In a single simulation, CompScore '= PoseScore - 5 * ContactScore. The CompScore in this table is recorded as the average of CompScore' in 10 simulations.
PoseScoreSD: Standard Deviation of PoseScore for 10 simulations.
ContactScoreSD: Standard Deviation of ContactScore for 10 simulations.
CompScoreSD: Standard Deviation of CompScoreSD for 10 simulations.
- Binding free energy
GBSA.csv: GBSA free energy values under different frames. The binding free energy of the ligand protein was calculated, and the contributions of different interactions to the binding free energy were split for subsequent analysis.
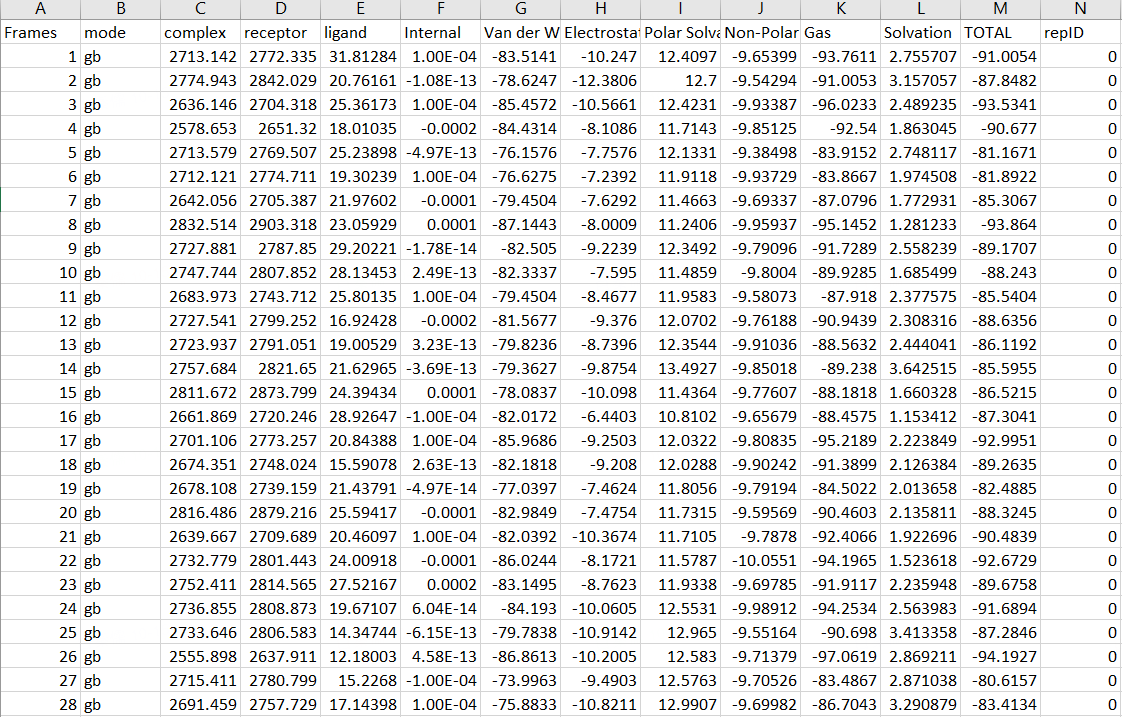
References
[1]:Nagar B, Bornmann WG, Pellicena P, et al. Crystal structures of the kinase domain of c-Abl in complex with the small molecule inhibitors PD173955 and imatinib (STI-571). Cancer Res. 2002;62(15):4236-4243.