EM Structure Fitting
1. Introduction
In recent years, cryo-electron microscopy (cryo Electron Microscopy, cryo-em) technology has become an important research tool in the field of structural biology, with a series of breakthroughs in hardware and algorithms, the applicability and resolution of cryo-em technology has been steadily improved. In the process of processing cryo-em data, it is very important to use the reconstructed three-dimensional density map to model biomolecules and further optimize the atomic coordinates. The EM Structure Fitting module (Hermite ® Uni-em | structure fitting) of the Hermite ® platform provides the flexible building function of the all-atom structure model based on the cryo-em density map. With the friendly operation interface, optimized algorithm and flexible cloud computing resources of Hermite platform, you can easily and quickly obtain high-quality cryo-em molecular structure model consistent with the density map.
2. How to use
2.1 Entrance
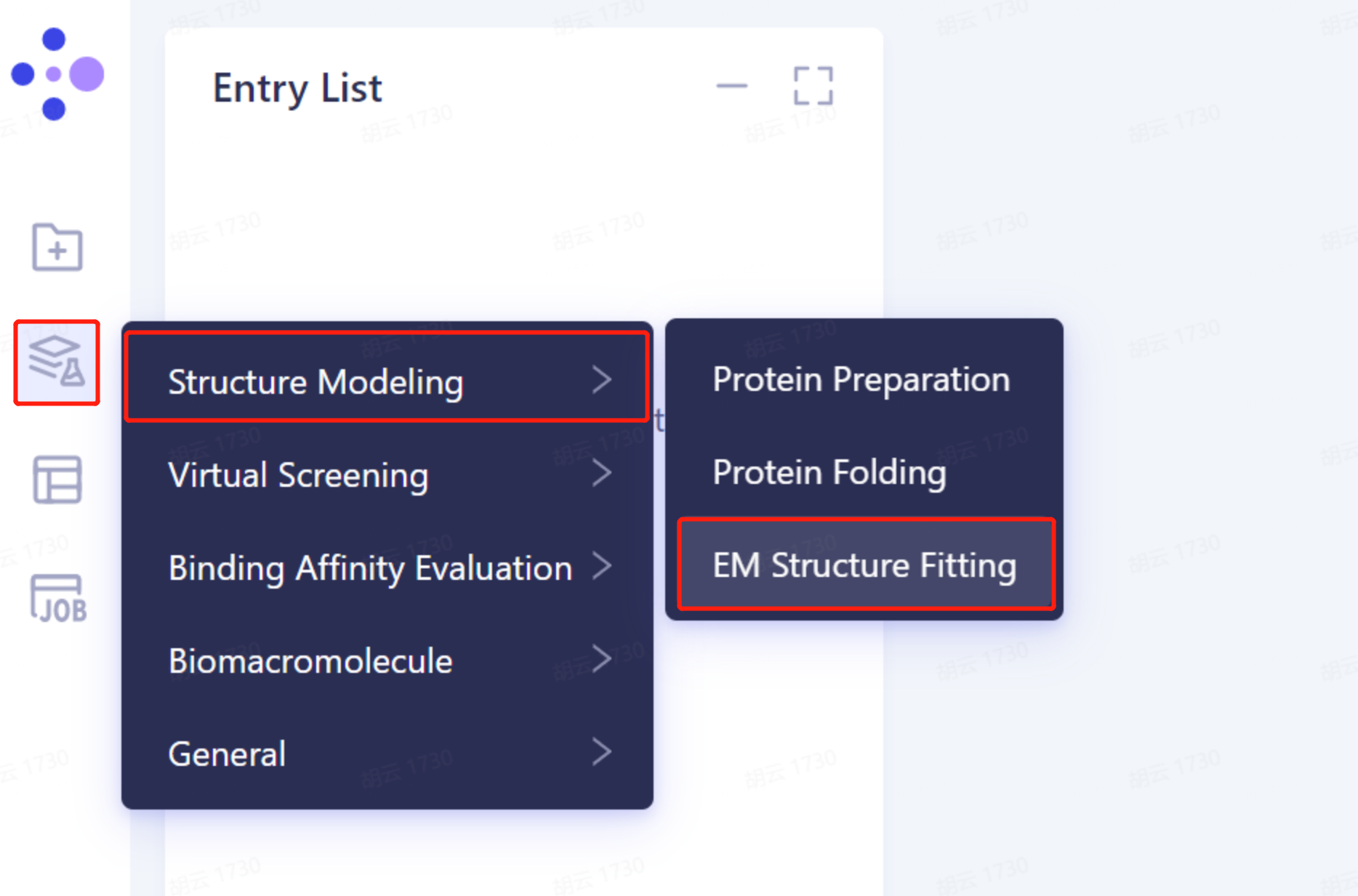
- The left general menu bar Function → Structure Modeling → EM Structure Fitting.
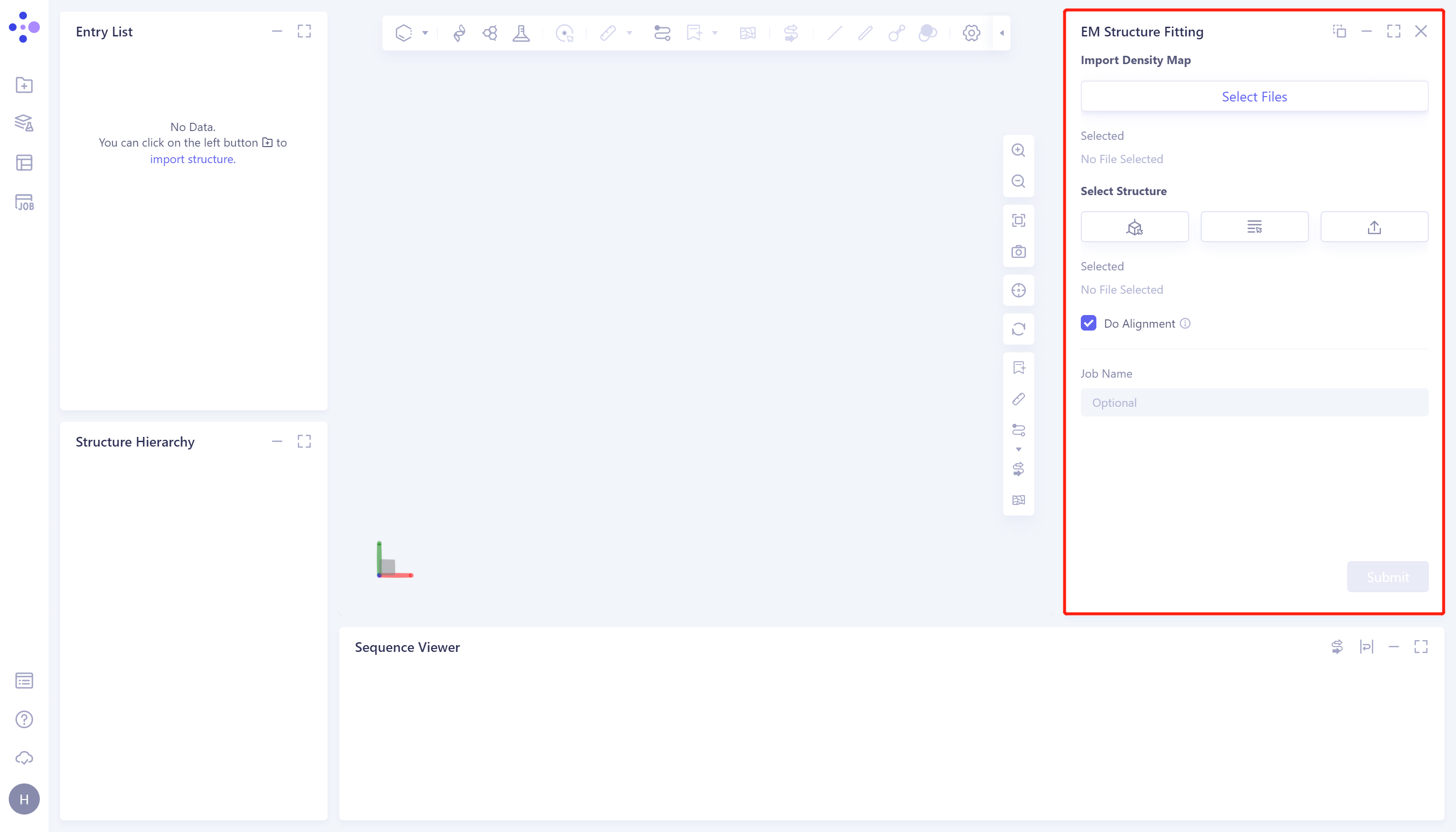
- The operation box of the EM Structure Fitting (shown in the red box) appears on the right side, and the overall interface is as follows:
2.2 Operation
-
Upload the electronic density map
- Support up to 200 M (upload large files will take a long time to load, please wait patiently); support.mrc and.ccp4 format files.
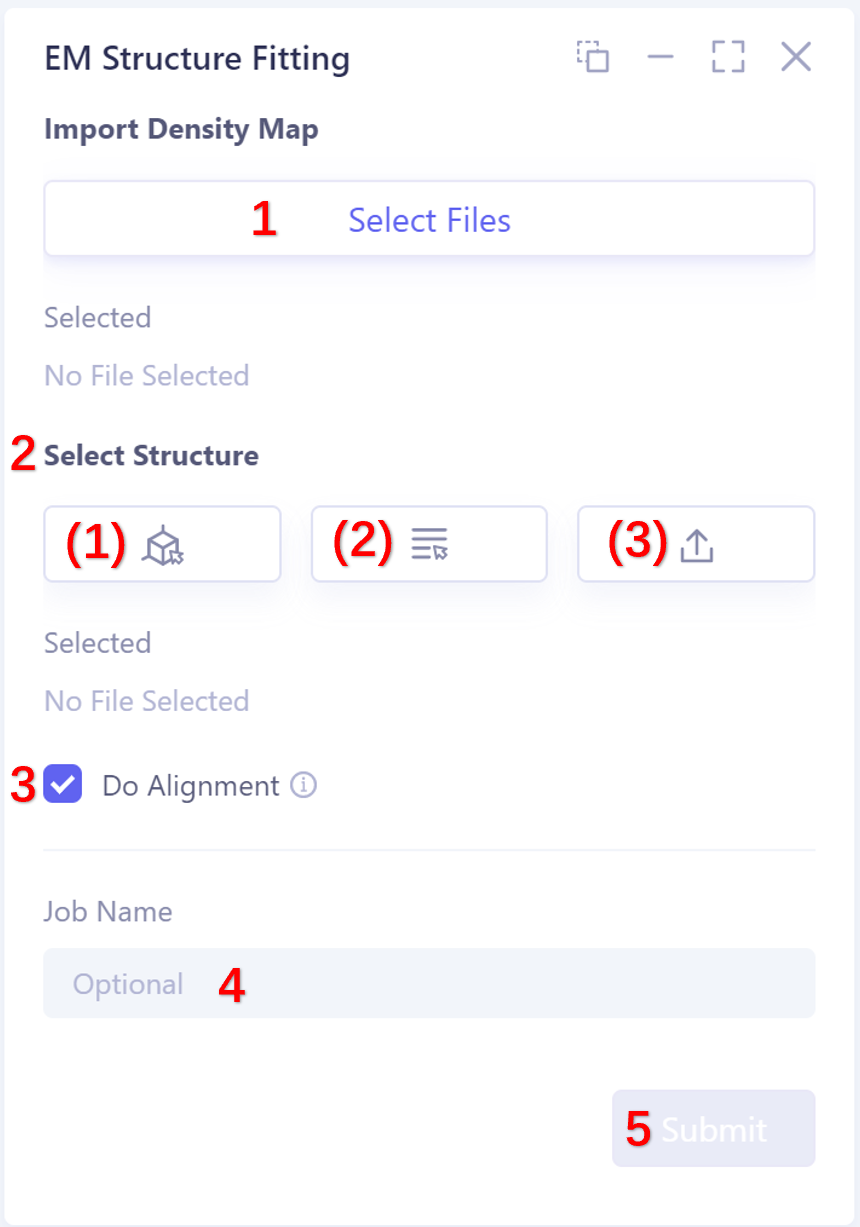
-
The selection protein structure supports three ways, as follows:
-
1)Select from 3D Workspace: Select the protein structure that has been imported into this task. The specific steps are as follows: click the Select from 3D Workspace icon to display the Select Structure interface → select protein structure from Structure Herarchy on the left side//select protein structure from 3D Workspace window → select on the right side Click OK in the Structure interface.
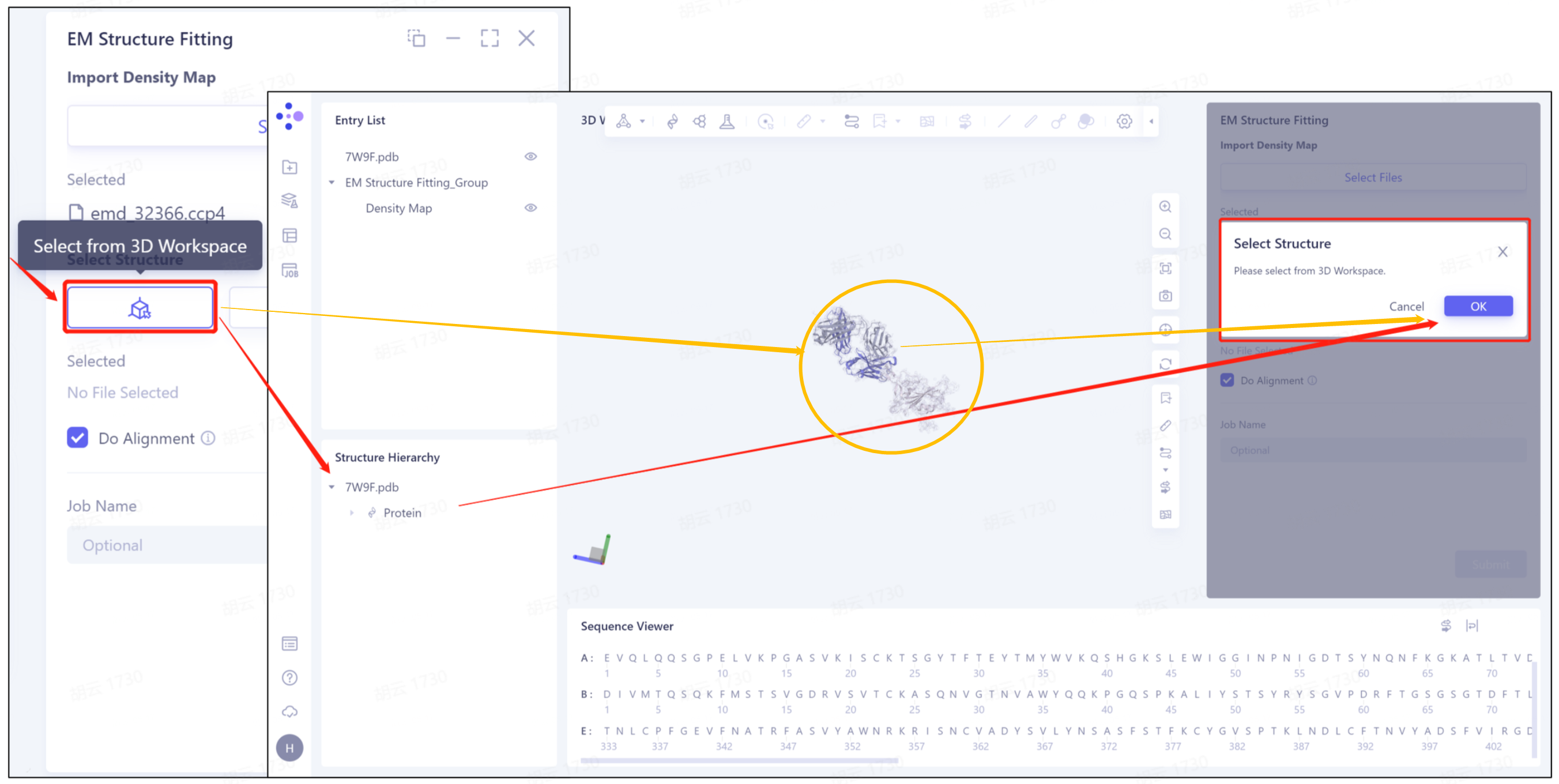
- 2)Select from Project: select the protein structure in the historical project. The steps are as follows: click the Select from Project icon to display the Select from Project interface → select the required protein structure in the Select from project interface → click OK.
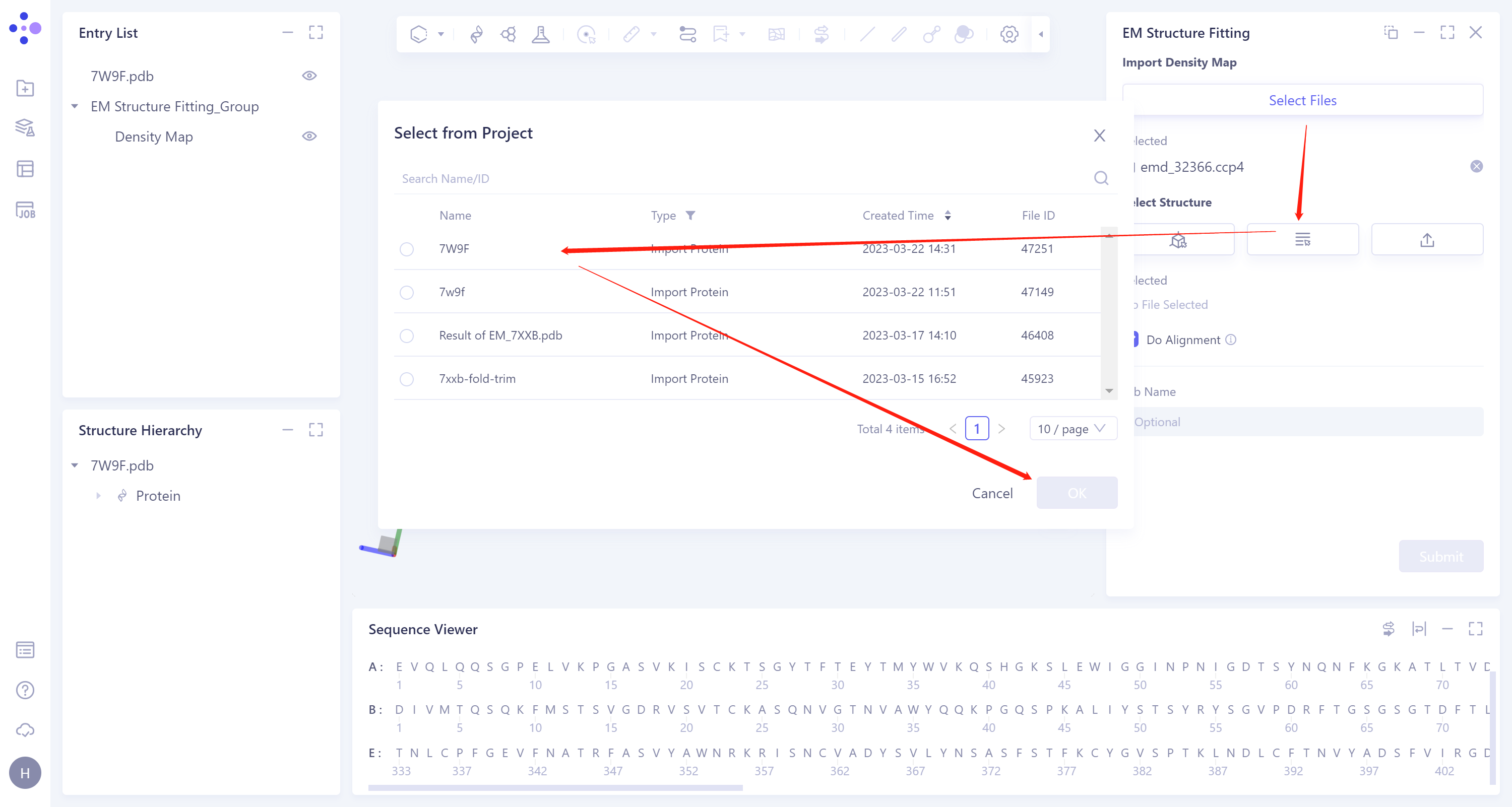
- 3)Select File: Click the Select File icon to import the local protein structure file, which supports the.pdb format file.
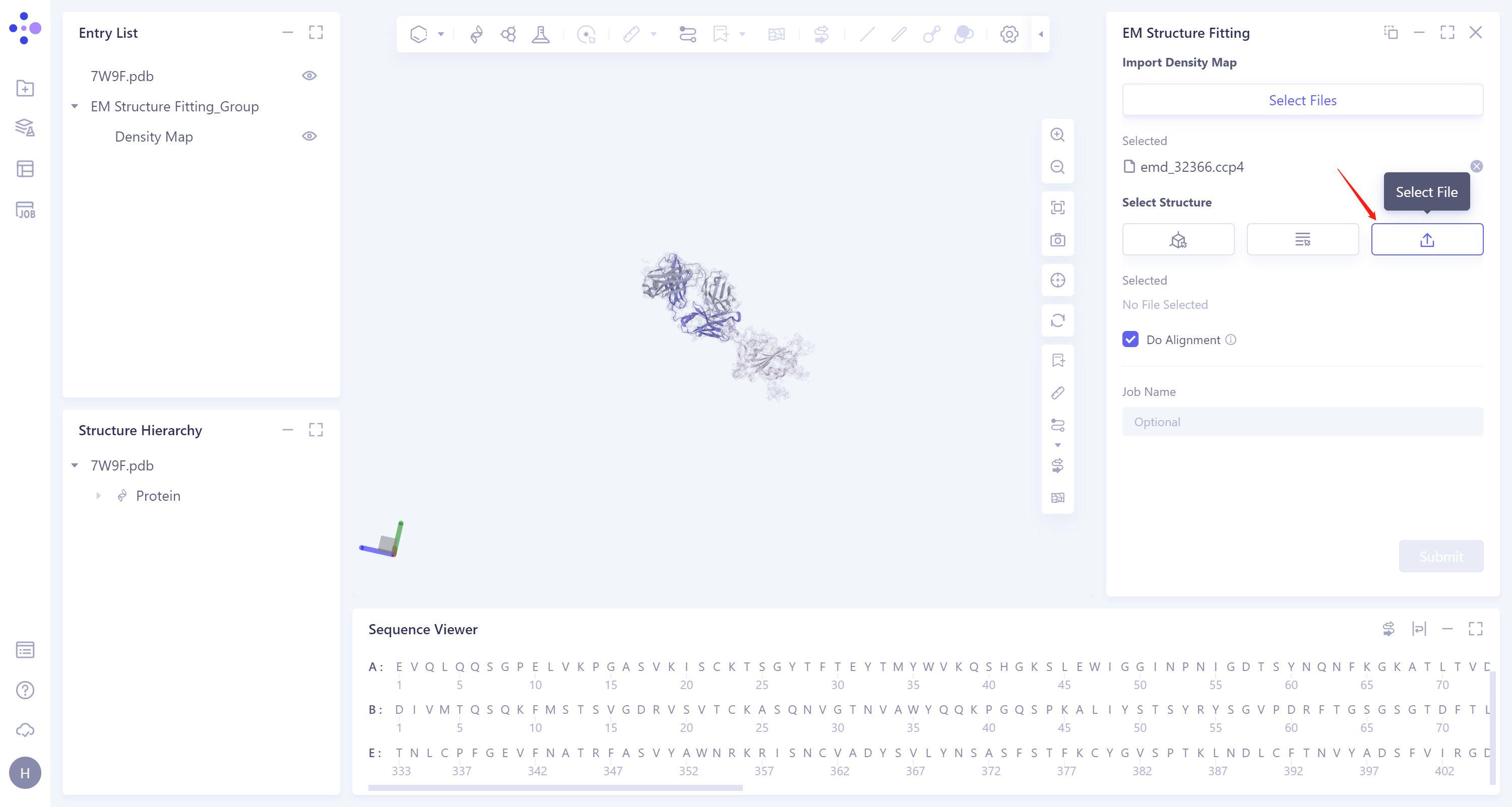
-
Do Alignment: This option compares the input structure with the electron density map. It is recommended to check Do Alignment.
-
Name the task at the Job Name ”.
-
Click “Submit” to submit the task.
3. Analysis of results
3.1 Entrance
-
General menu bar on the left: Menu Job → Find the required task.
-
The task can be found by searching for the Job Name, or by filtering the Job Type.
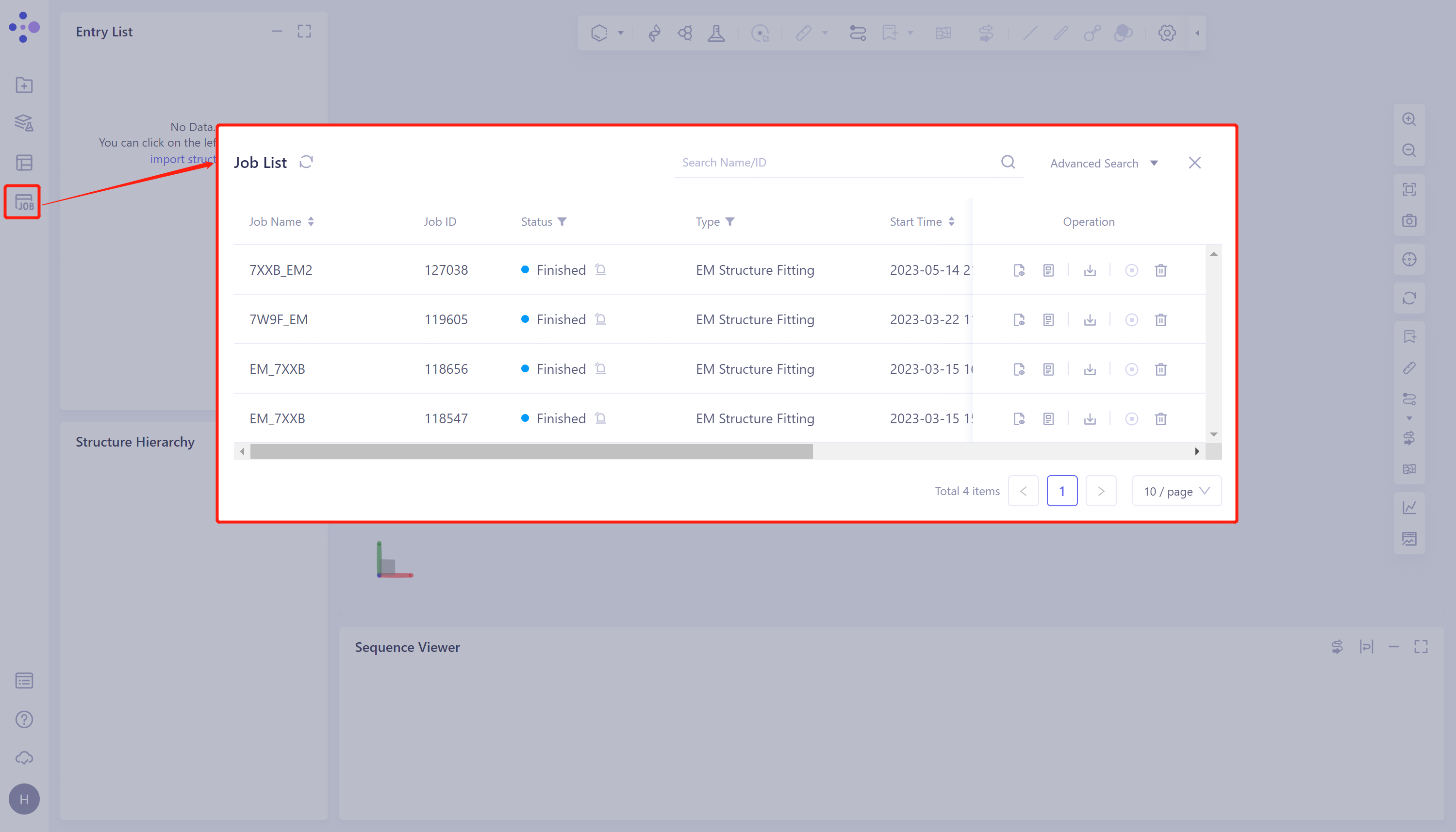
3.2 Results presentation
-
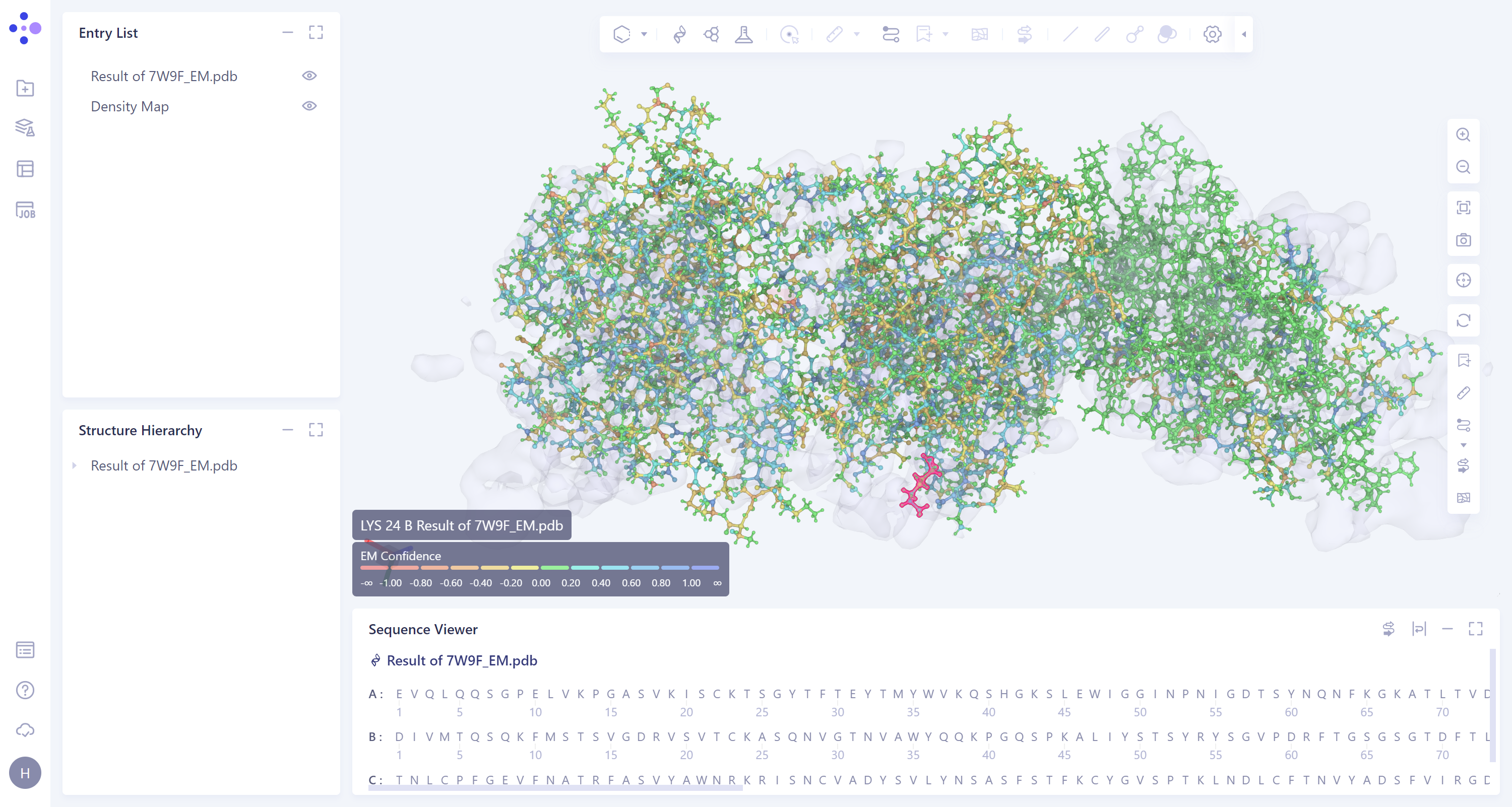
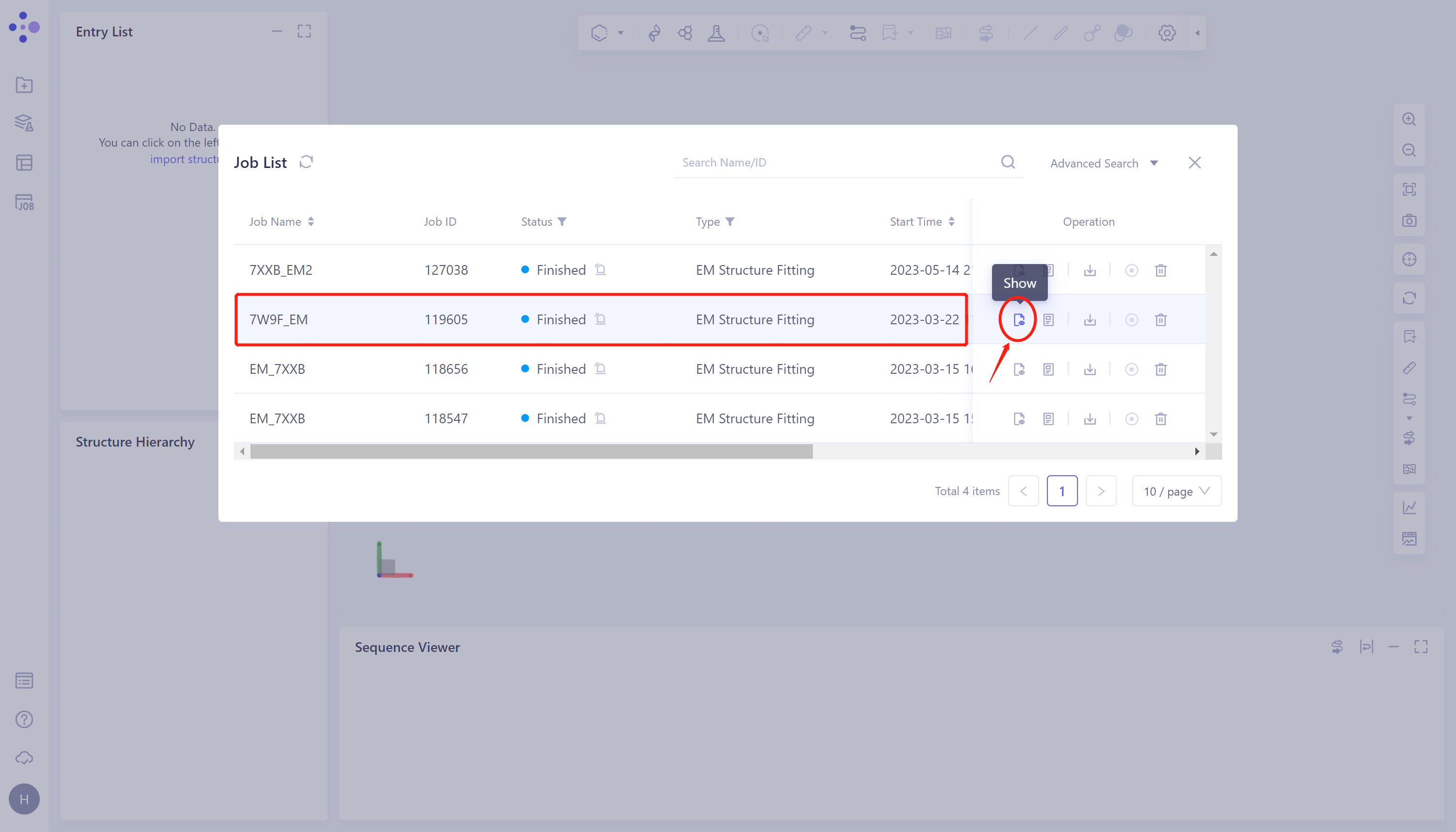
Select the task to be viewed and click Show in the Operation column to display the result of the task, as shown in the figure below.
-
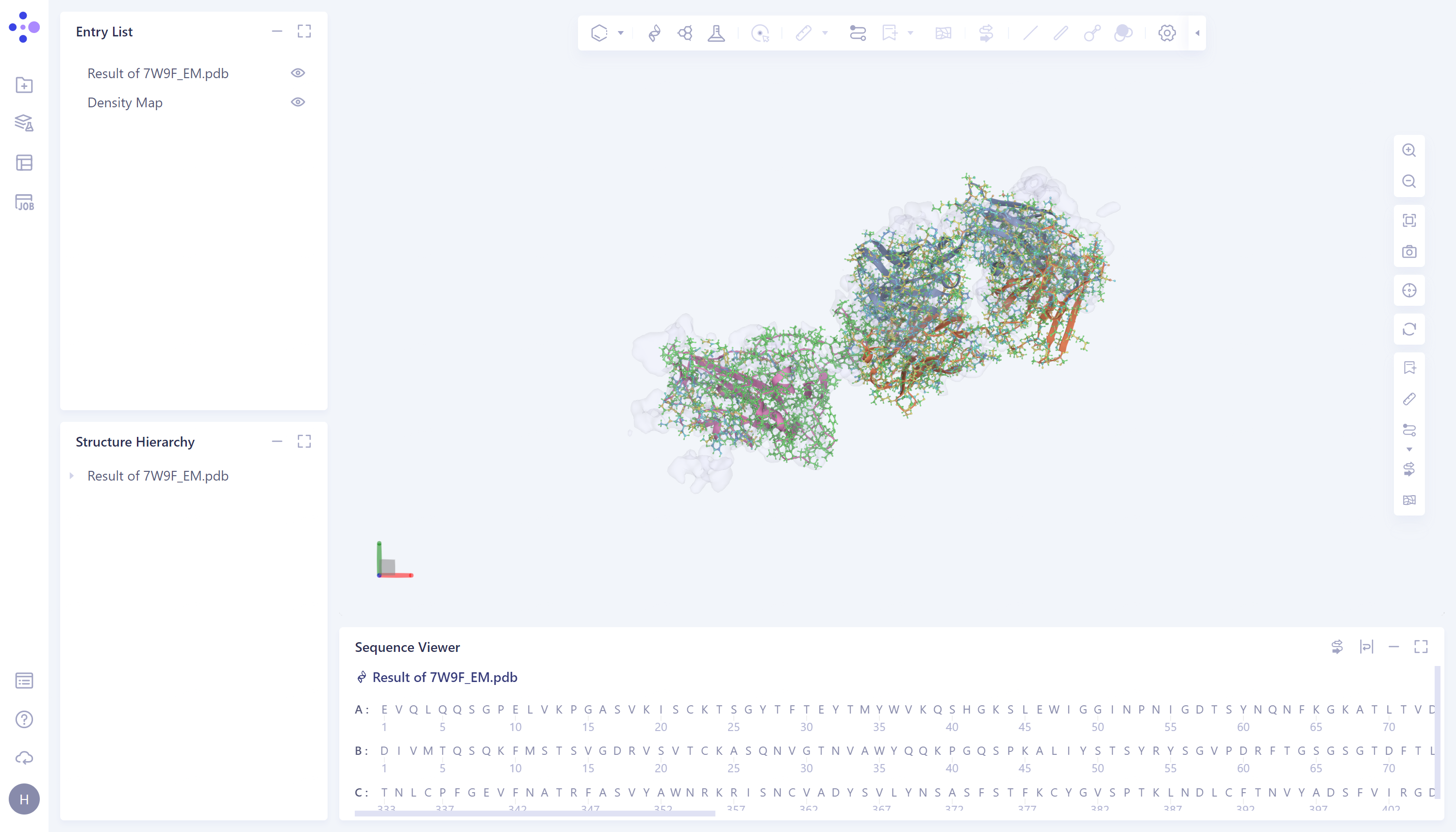
The imported task results are in the Entry List on the left. The results can be displayed or not displayed by adjusting the opening and closing of the small eyes. When the small eyes are opened, the results are displayed.
-
It is recommended to turn off Secondary Structure by clicking on the Ribbon icon on the right.
- The mouse hovers over an amino acid residue and the confidence information appears.