Aquasite
1. Introduction
Water molecules are widely distributed in drug target proteins and play an important role in the interaction between drugs and target proteins. In drug design, it is an effective way to optimize the structure of compounds by replacing water molecules and forming interactions with water molecules. In the process of using water molecules to guide the modification of compounds, because the free energy of water molecules can not be known simply from the observation of crystal structure, theoretical calculation has become an important means to predict the sites and properties of water molecules. Uni-Aquasite calculates the free energy of each water molecule by molecular dynamics simulation, and finds out the position of stable and unstable water molecules at the binding site of the target, which provides structural basis and guiding ideas for drug molecular design.
2. How to use
2.1 Entrance
- Left general menu bar Menu → Function → General → Aquasite
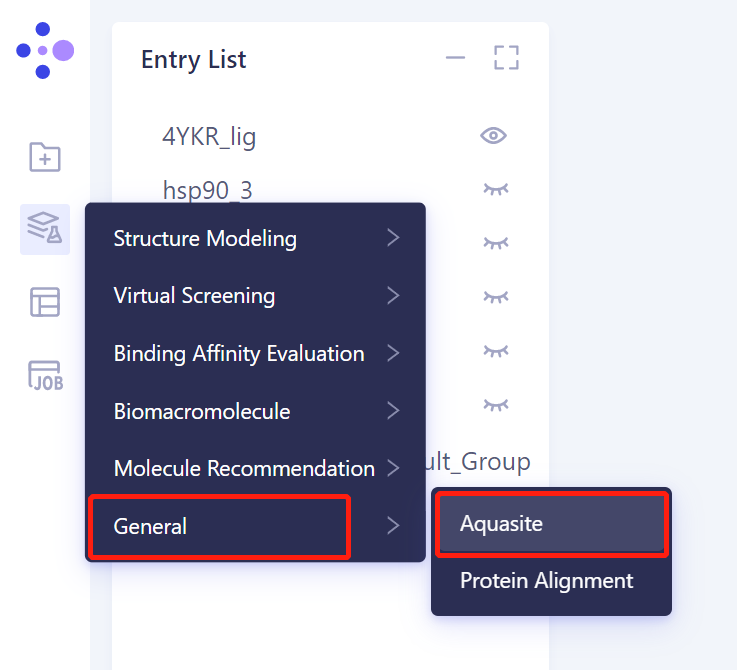
- The operation box of Aquasite (shown in the red box) appears on the right side, and the overall interface is as follows:
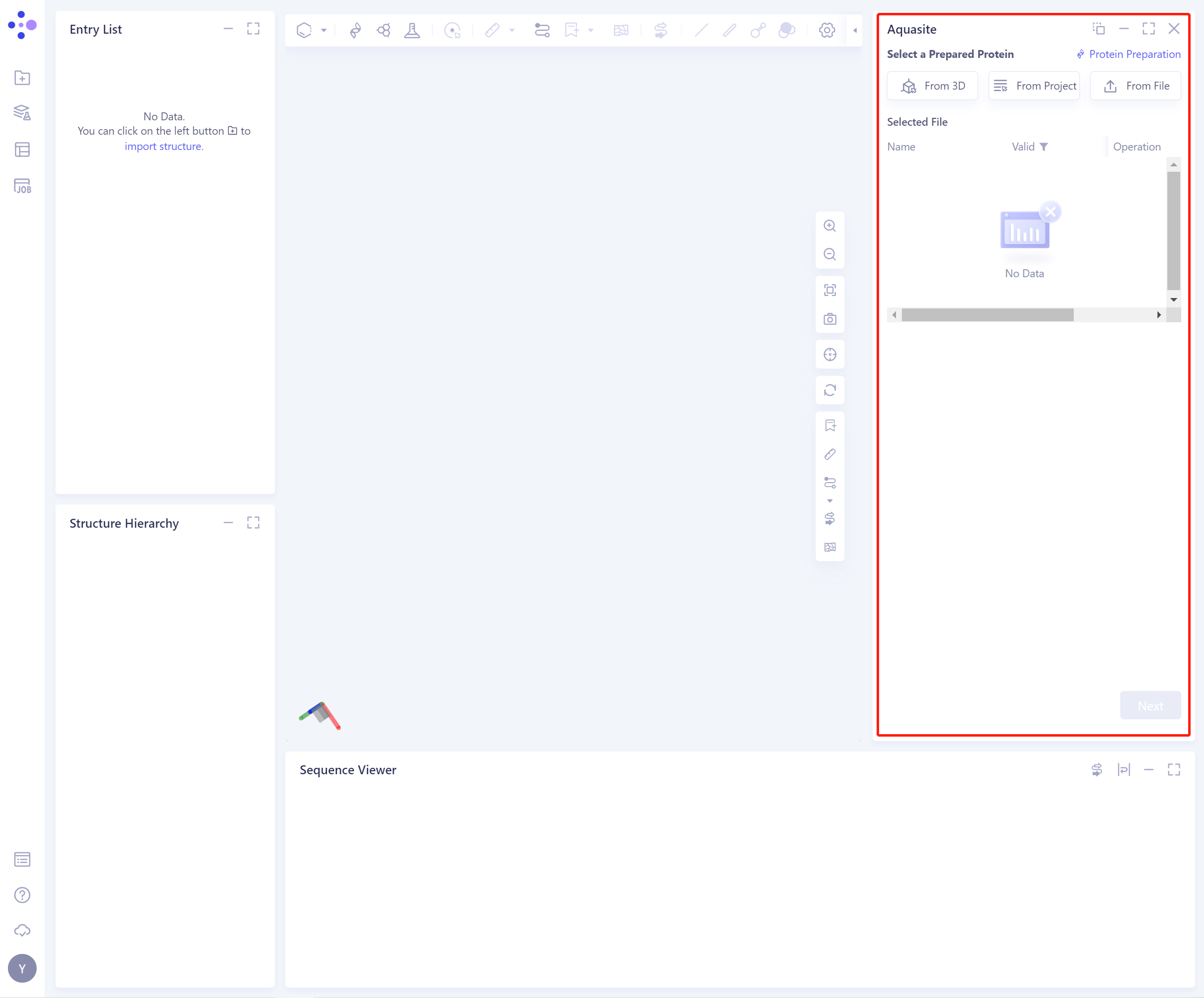
2.2 Operation
-
There are three ways to introduce proteins:
-
1)From 3D Works pace: Select proteins from graphical components.
-
3D Workspace only supports the selection of protein level, which can be selected in Hierarchy (linkage 3D).
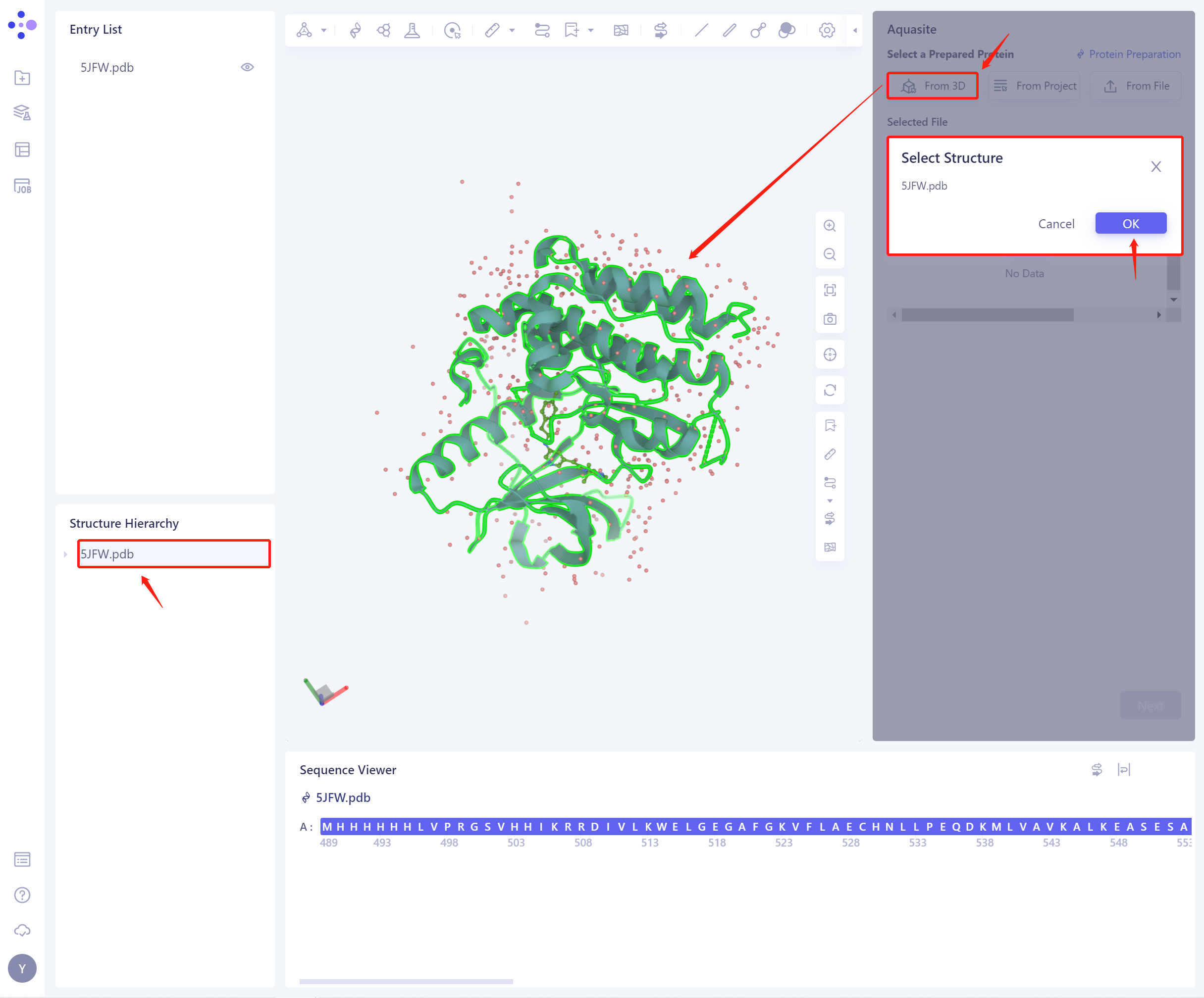
-
2)From Project: Select proteins from historical projects.
-
In the pop-up Select from Project operation interface, you can search for files by file name and ID through Search Name/ID, and adjust the Type to select files.
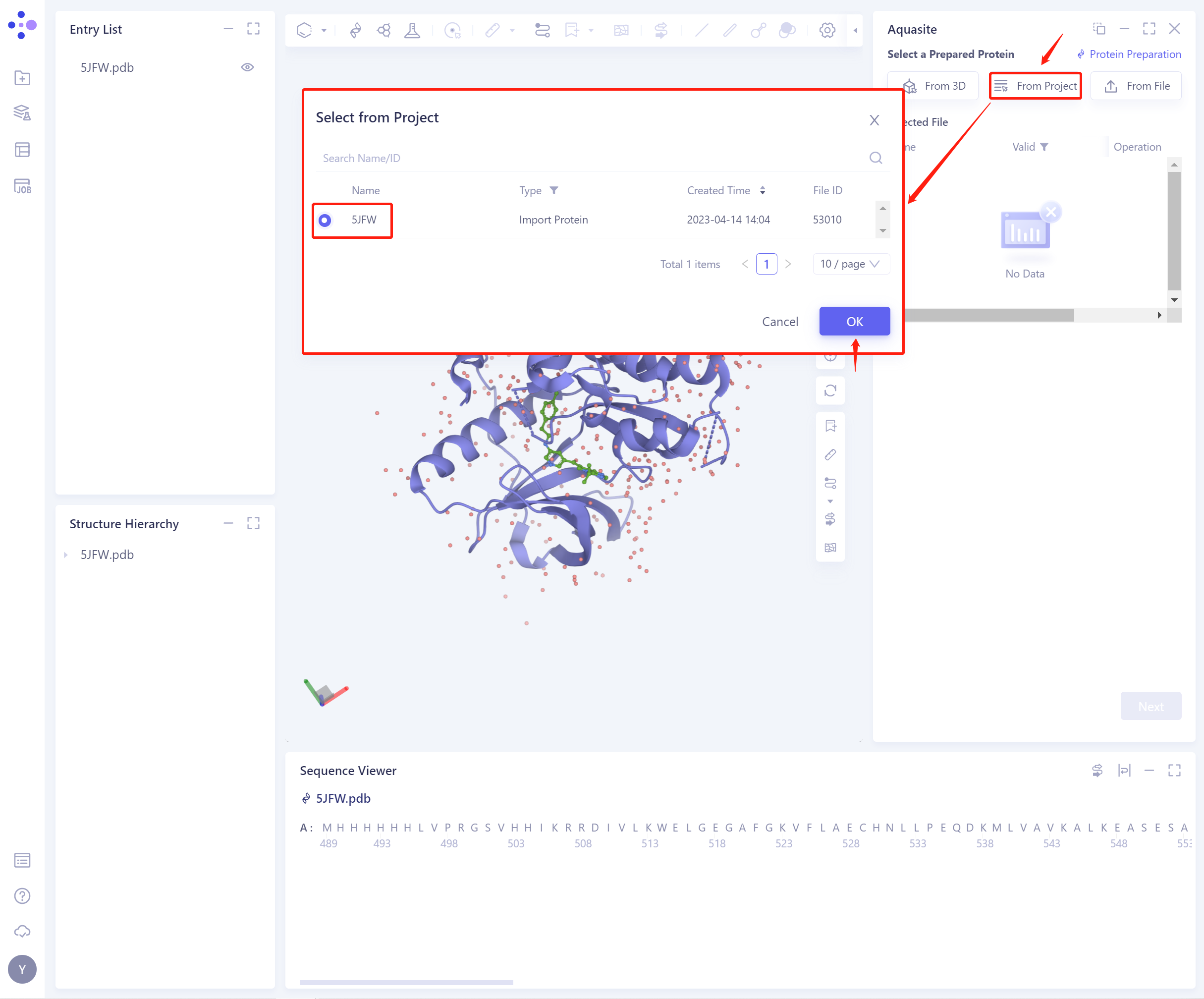
- 3)From File: Upload a local protein file in.pdb format.
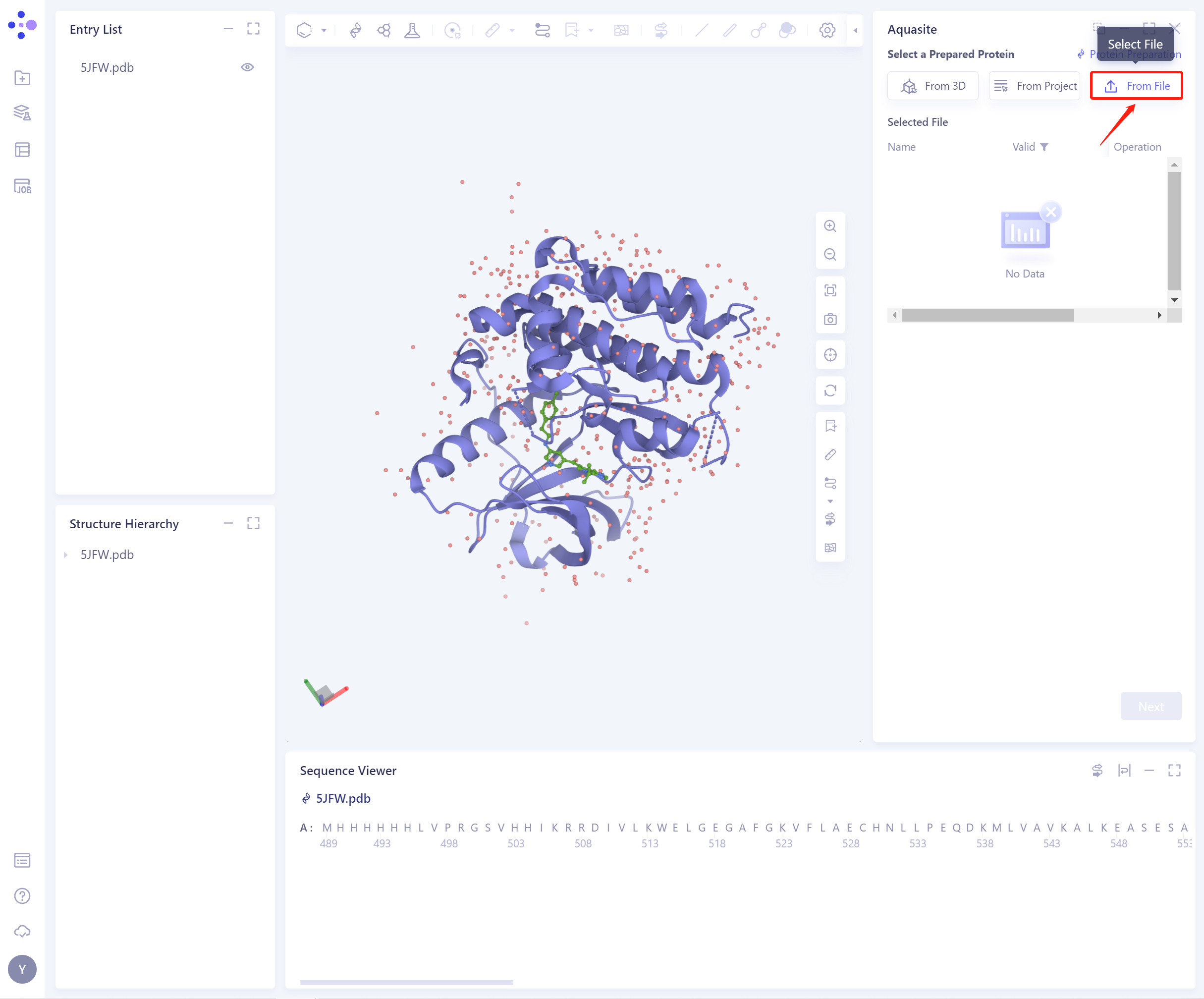
- The selected protein will be checked by the force field, and the following figure shows the checking.
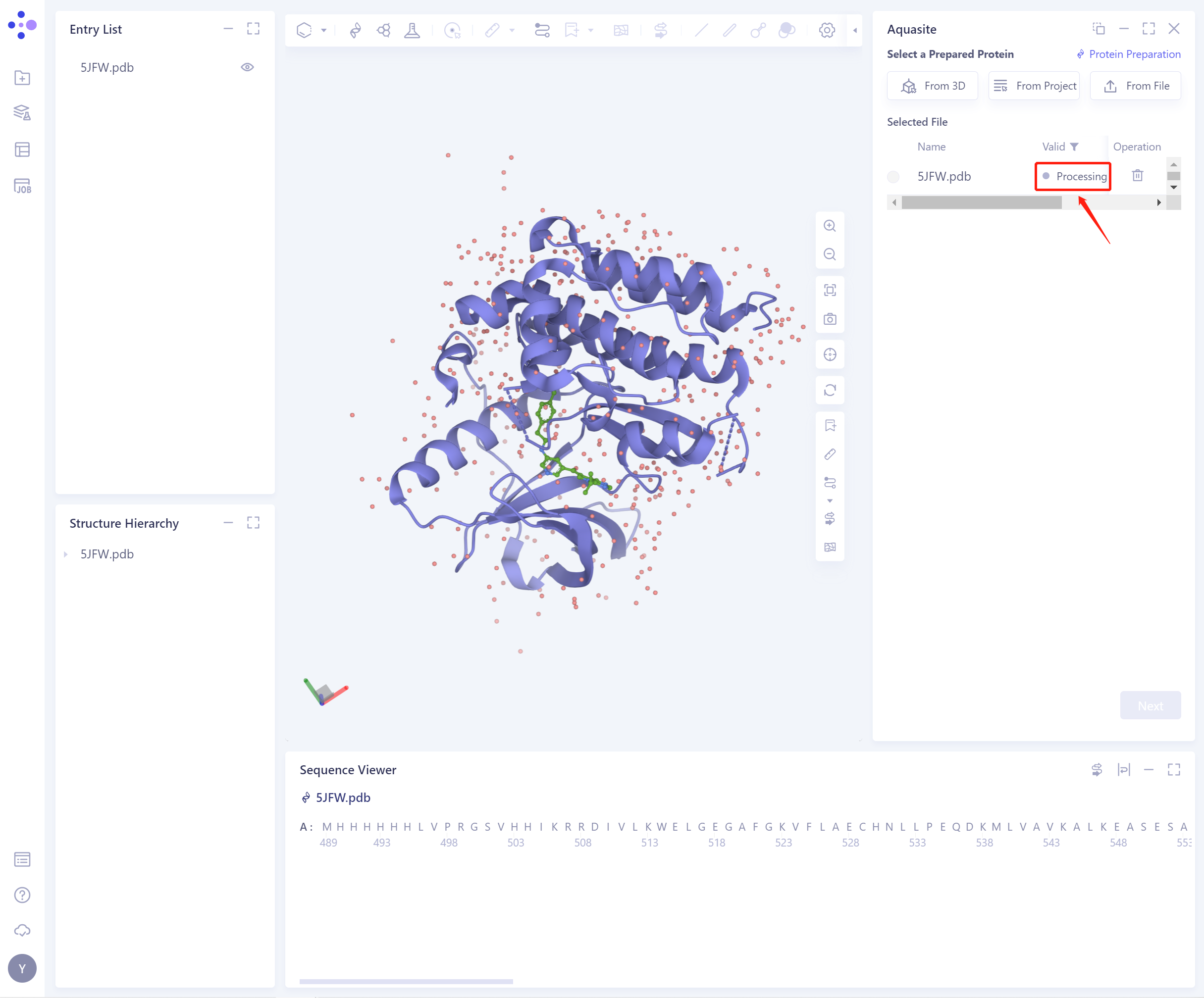
- If the protein does not pass the force field check (here is Valid, that is, the force field check passes), protein preparation is required (see the red box on the upper right of the figure for the shortcut).
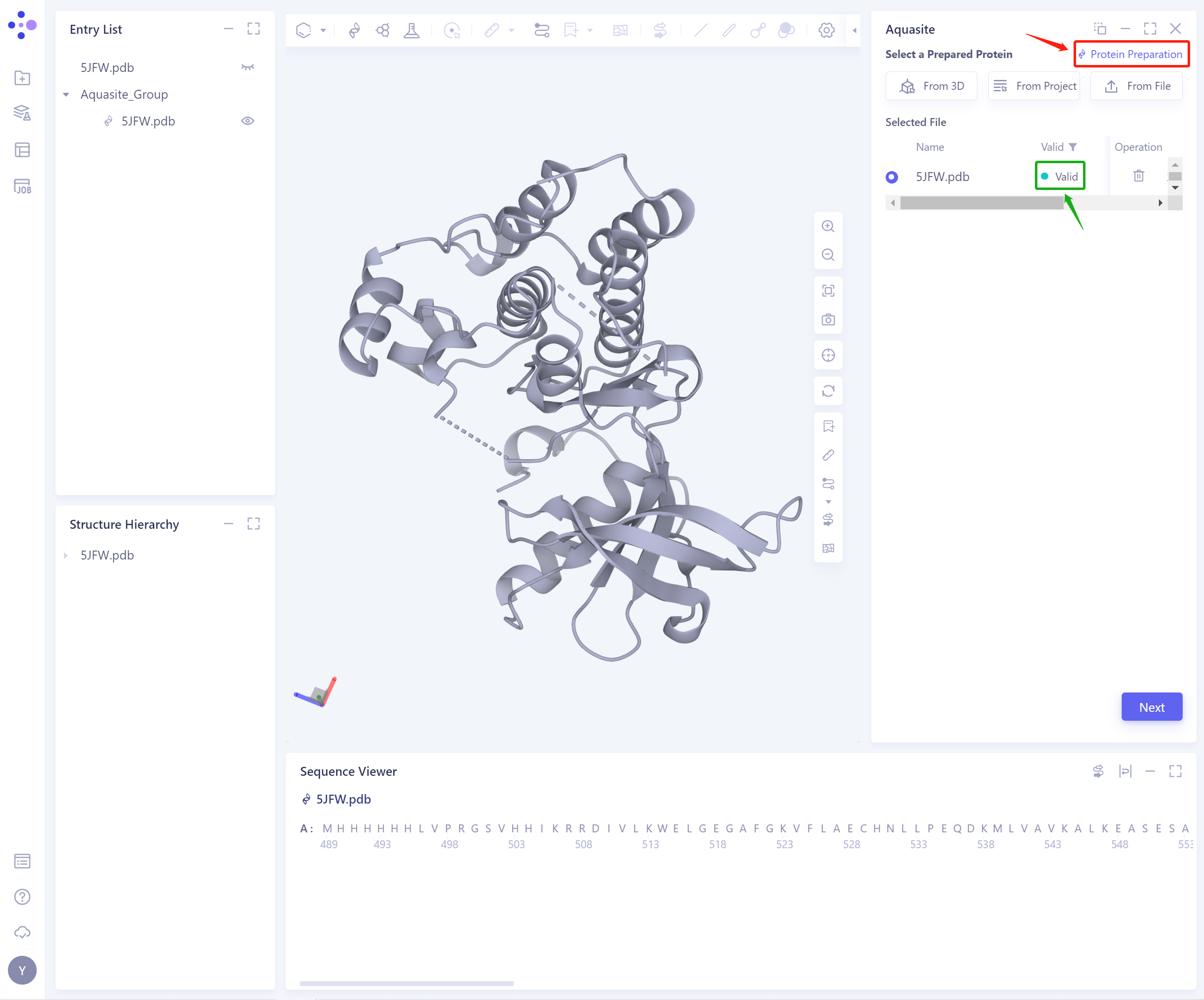
- After the ligand force field check is passed, the function can continue to be used.
-
There are three ways to import ligand:
-
1)From 3D Works pace: Select the ligand from the graphical component.
-
3D Workspace only supports the selection of Residue level, which can be selected in Hierarchy (linkage 3D).
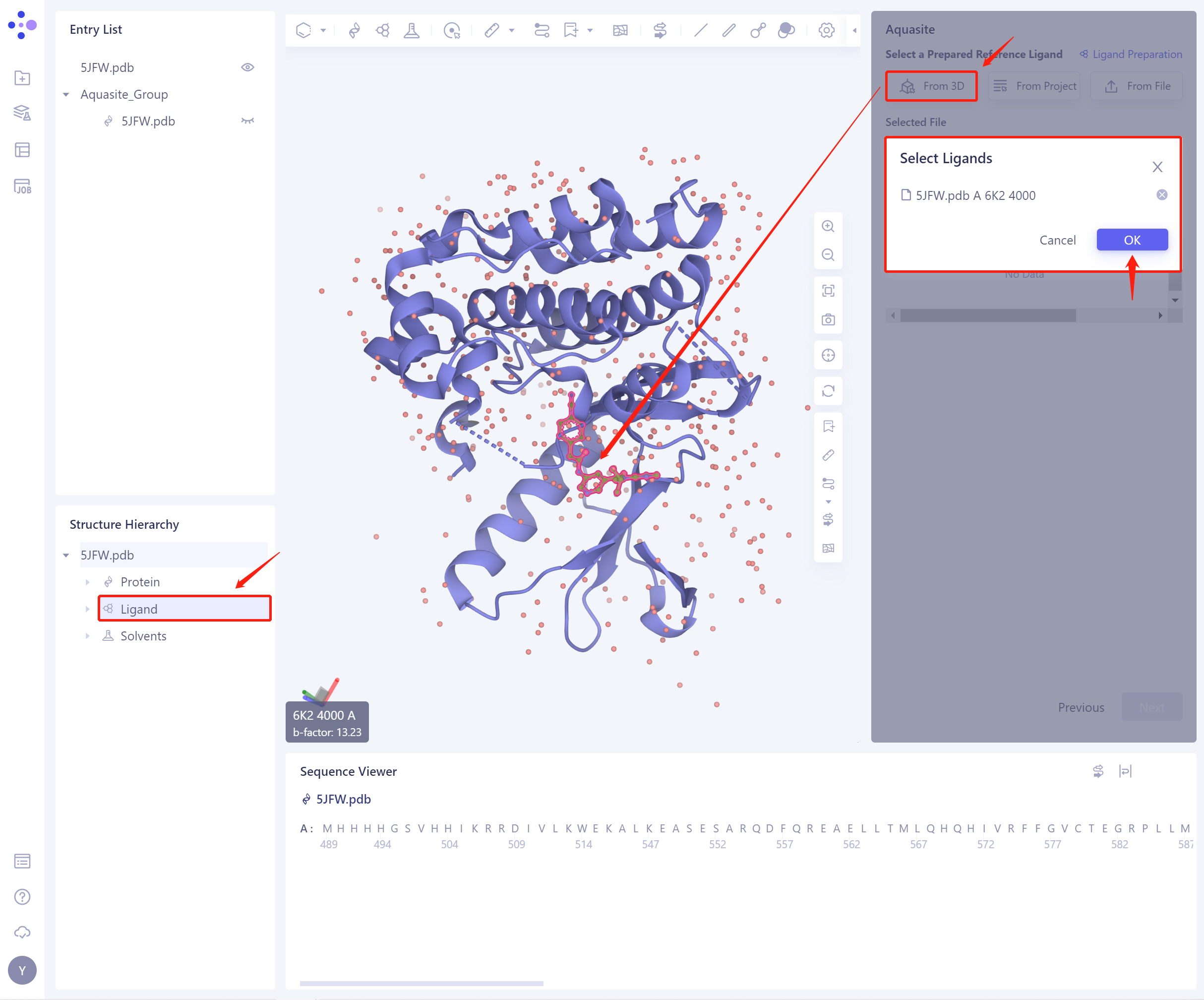
-
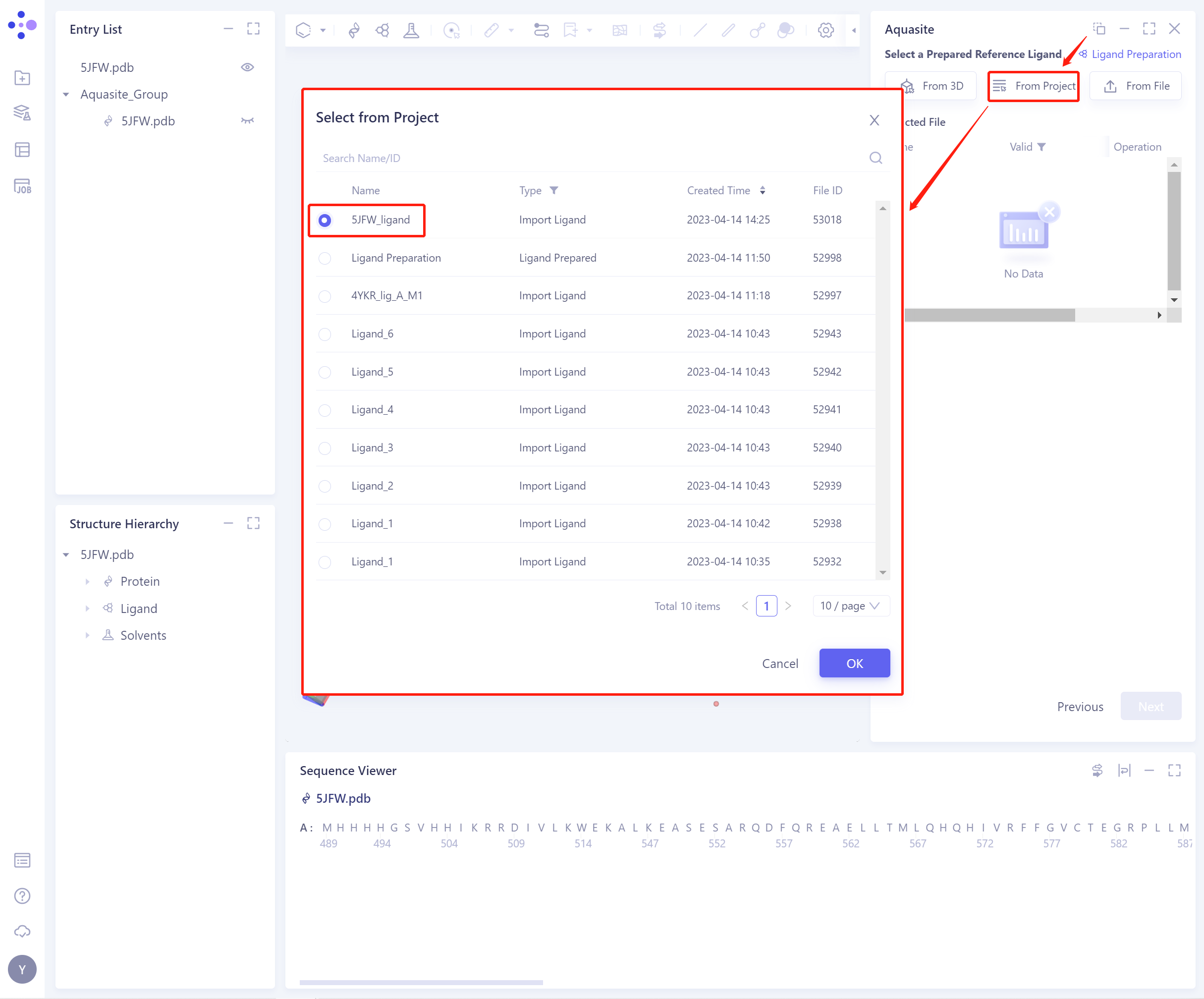
2)From Project: select the ligand from the historical project.
-
In the pop-up Select from Project operation interface, you can search for files by file name and ID through Search Name/ID, and select the ligand file by adjusting the type (Type).
- 3)From File: Upload the local ligand file, which supports the.sdf/.mol format.
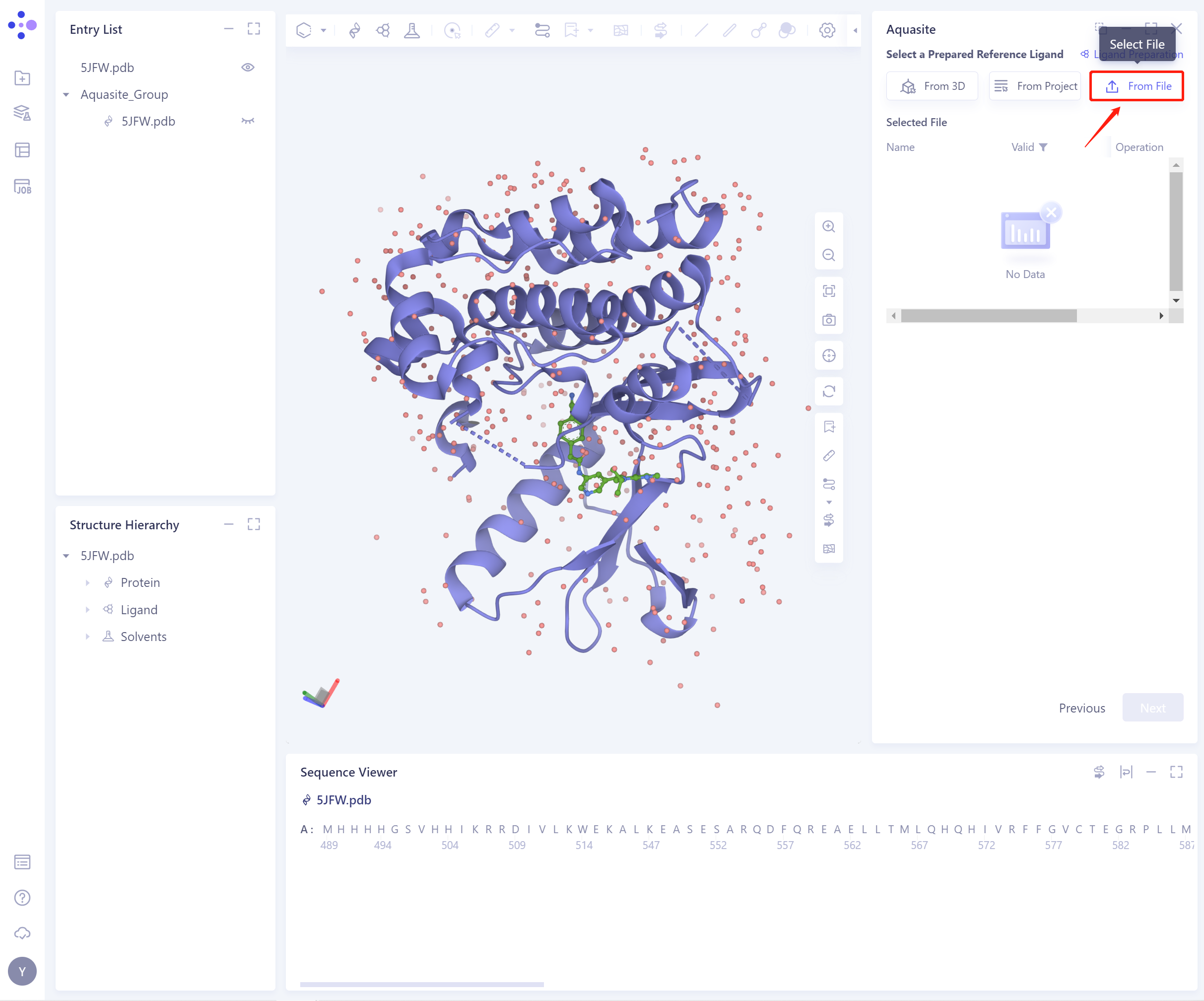
- Before importing the ligand, you need to select whether to repair the bond information that may be lost by the ligand. The default is yes.
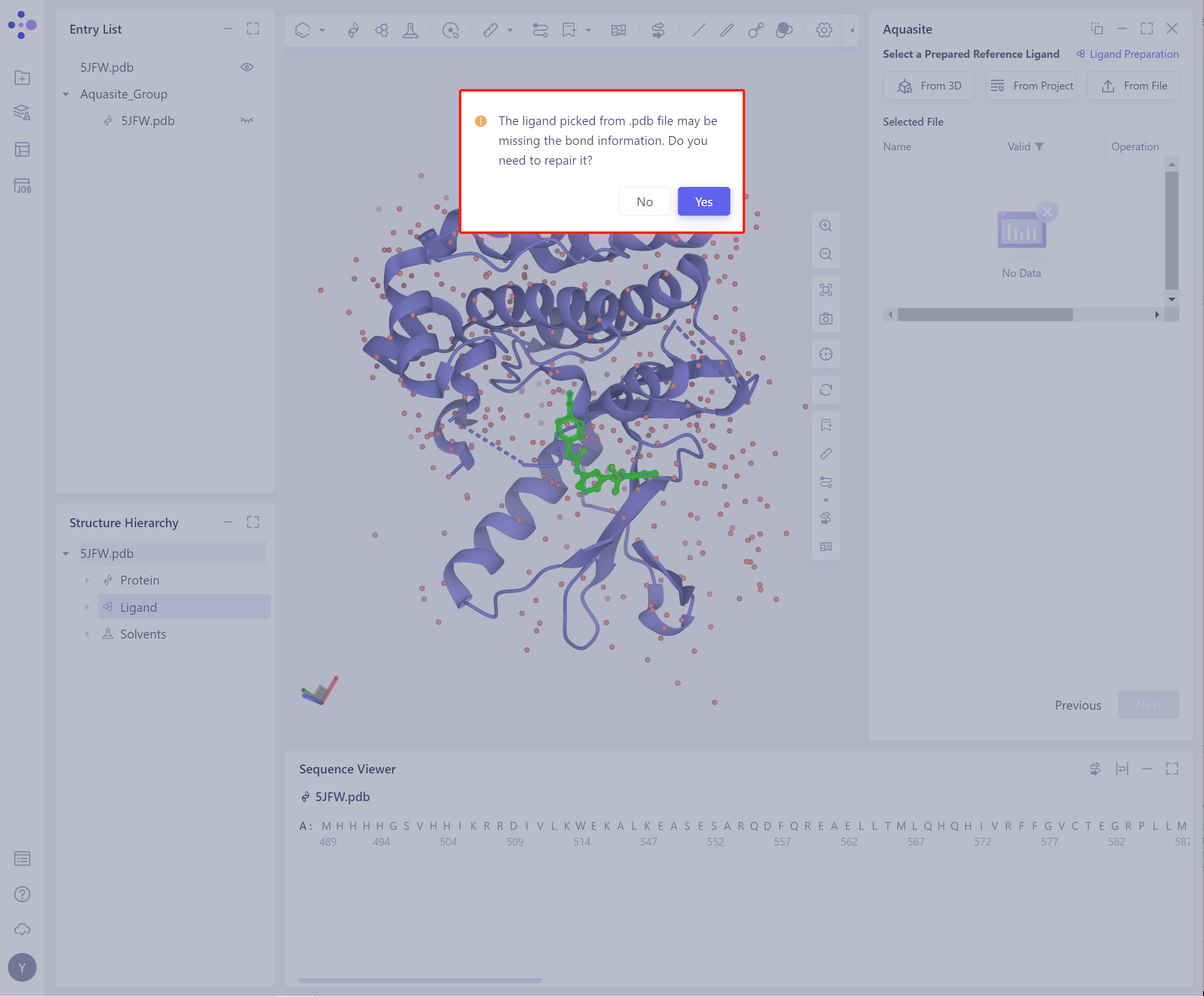
- The selected ligand will be checked by the force field, and the following figure shows the checking.
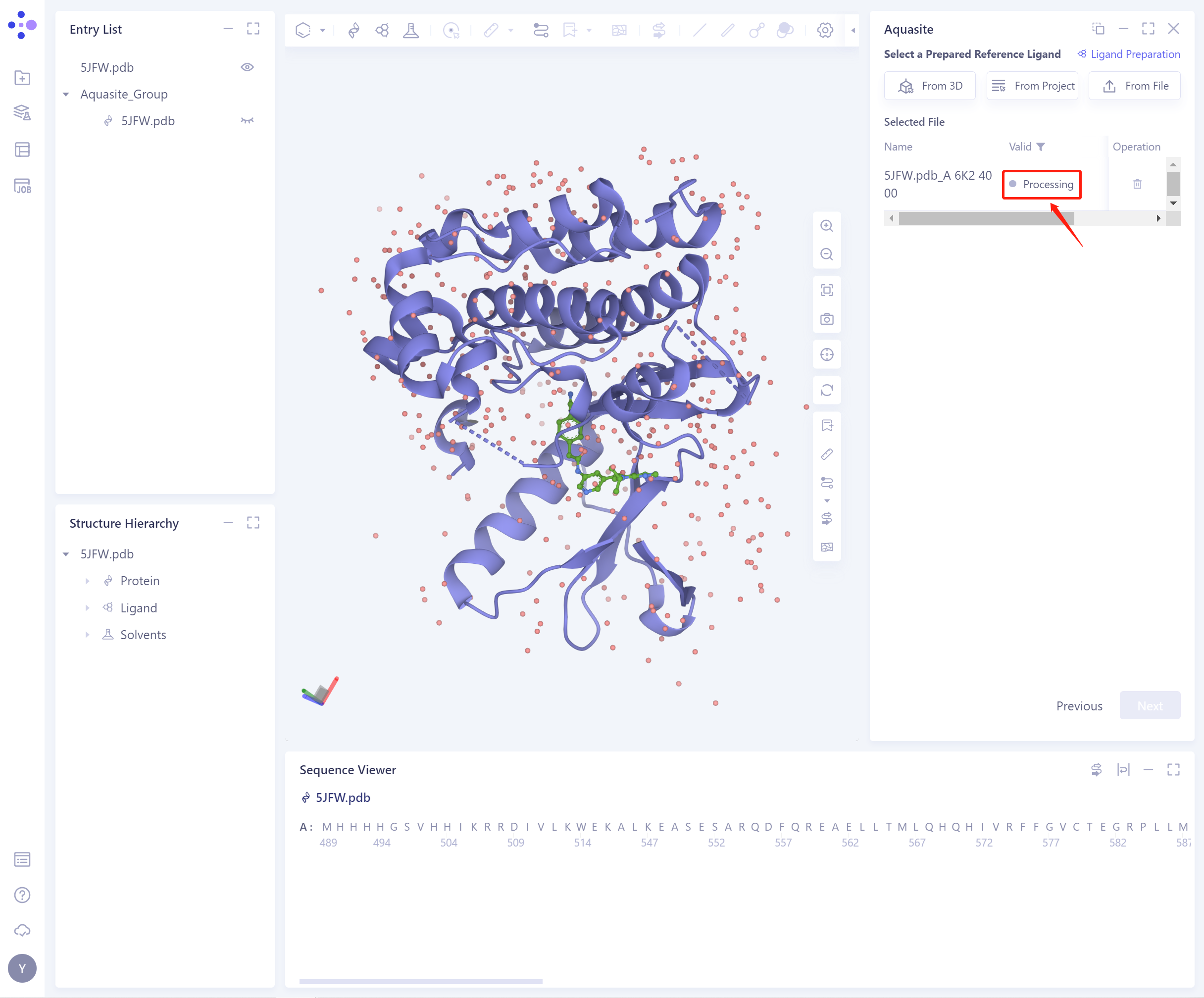
- If the ligand force field inspection is not passed (here is Valid, that is, the force field inspection is passed), the ligand preparation (Ligand Preparation, see the red box on the upper right of the figure for the shortcut) is required.
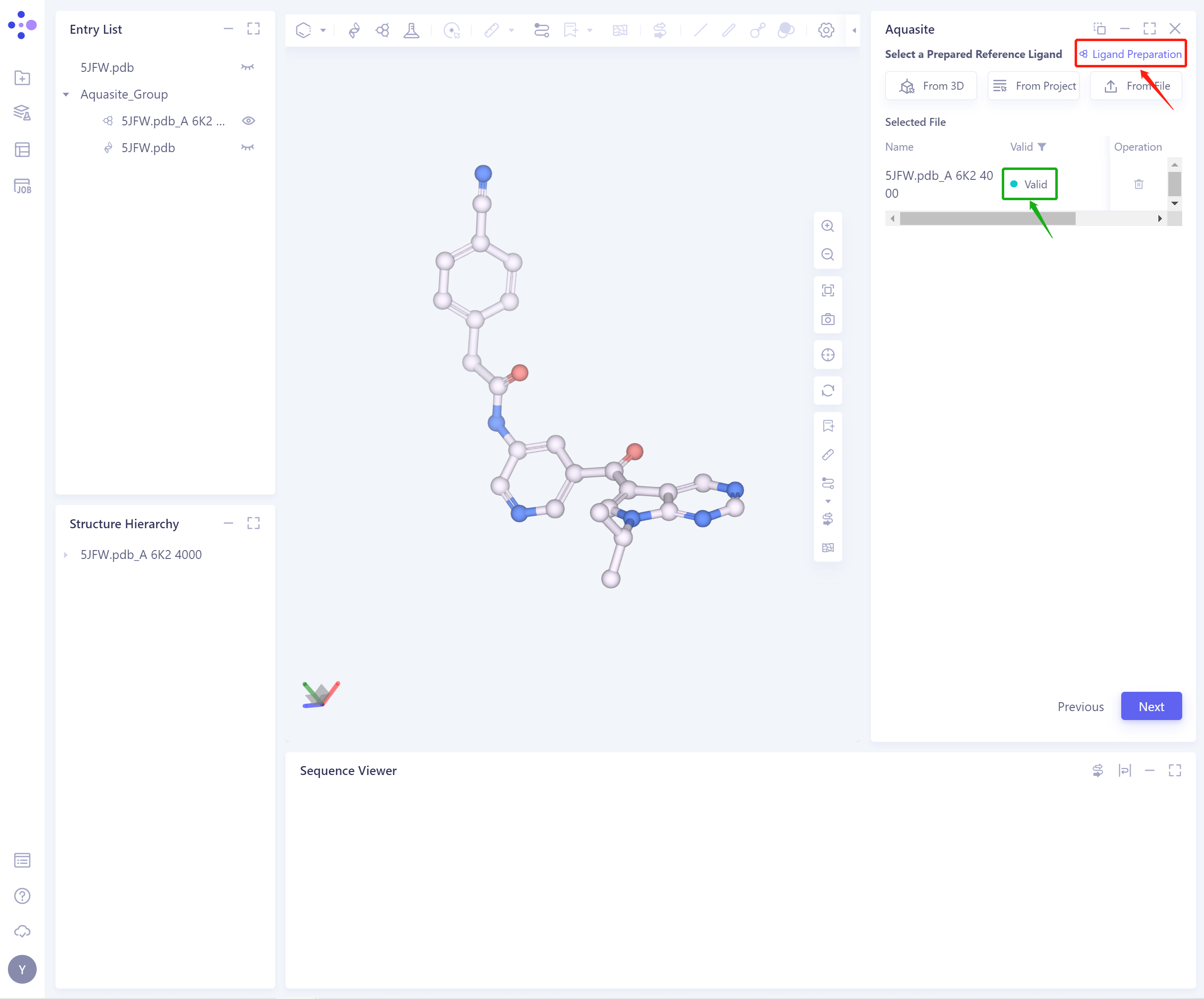
- After the ligand force field check is passed, the function can continue to be used.
-
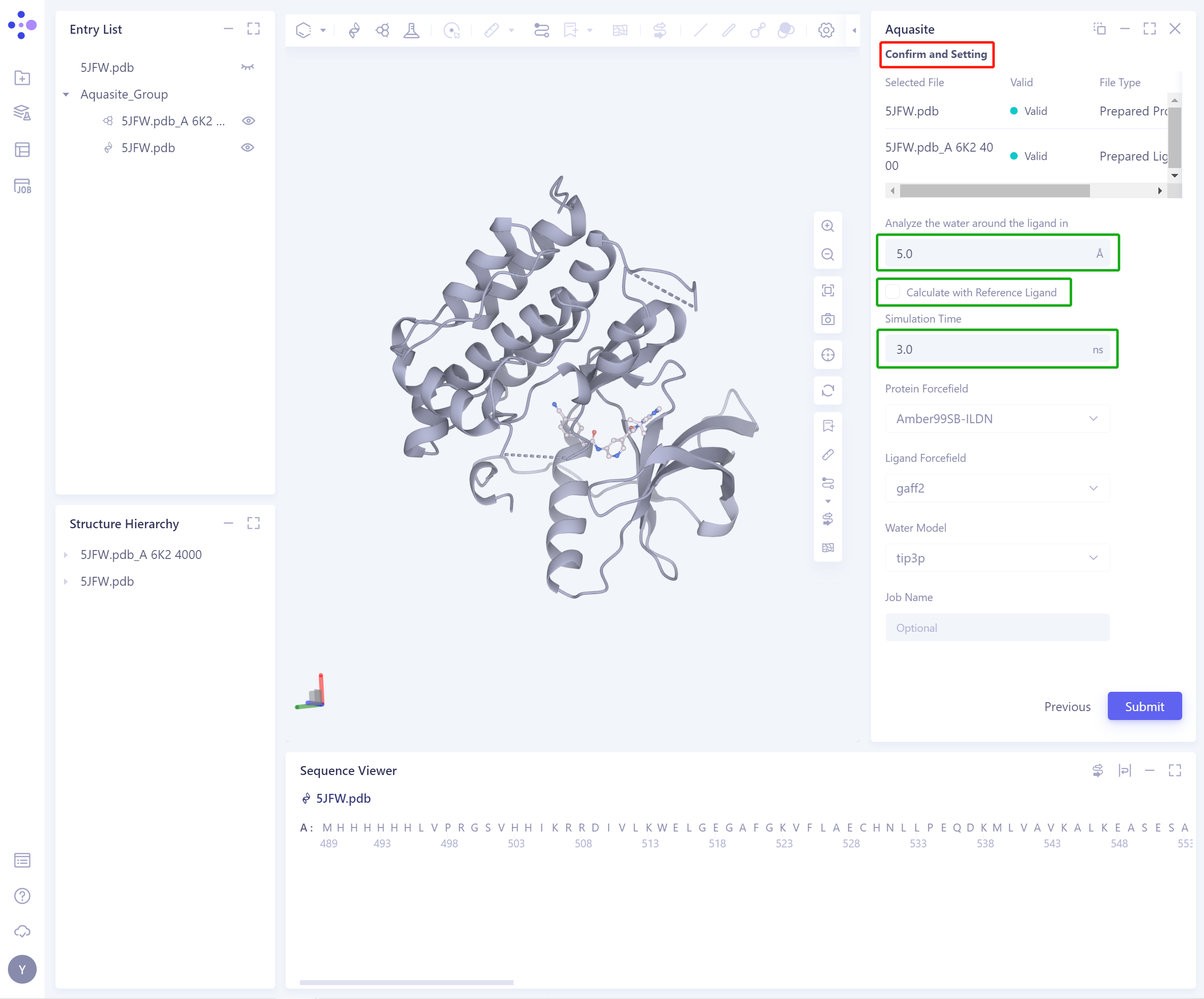
Confirm and Setting:
-
Analyze the water around the ligand in: determine the space where water molecules are generated around the ligand;
-
Calculate with Reference Ligand: If you want to keep the ligand for simulation, check this option and remove the original system water molecules that may collide with the ligand in 3D Workspace. If the ligand is not reserved for simulation, there is no need to check this option;
-
Simulation Time: determine the simulation duration;
-
Protein Forcefield, Ligand Forcefield and Water model are not supported to be changed, and the default parameters are maintained.
-
-
Job Name: Name the task.
-
Submit: Submit the task.
3. Analysis of results
3.1 Entrance
-
The generic menu bar on the left Job → Find the Aquasite Job.
-
The task can be found by searching for the Job Name, or by filtering the Job Type.
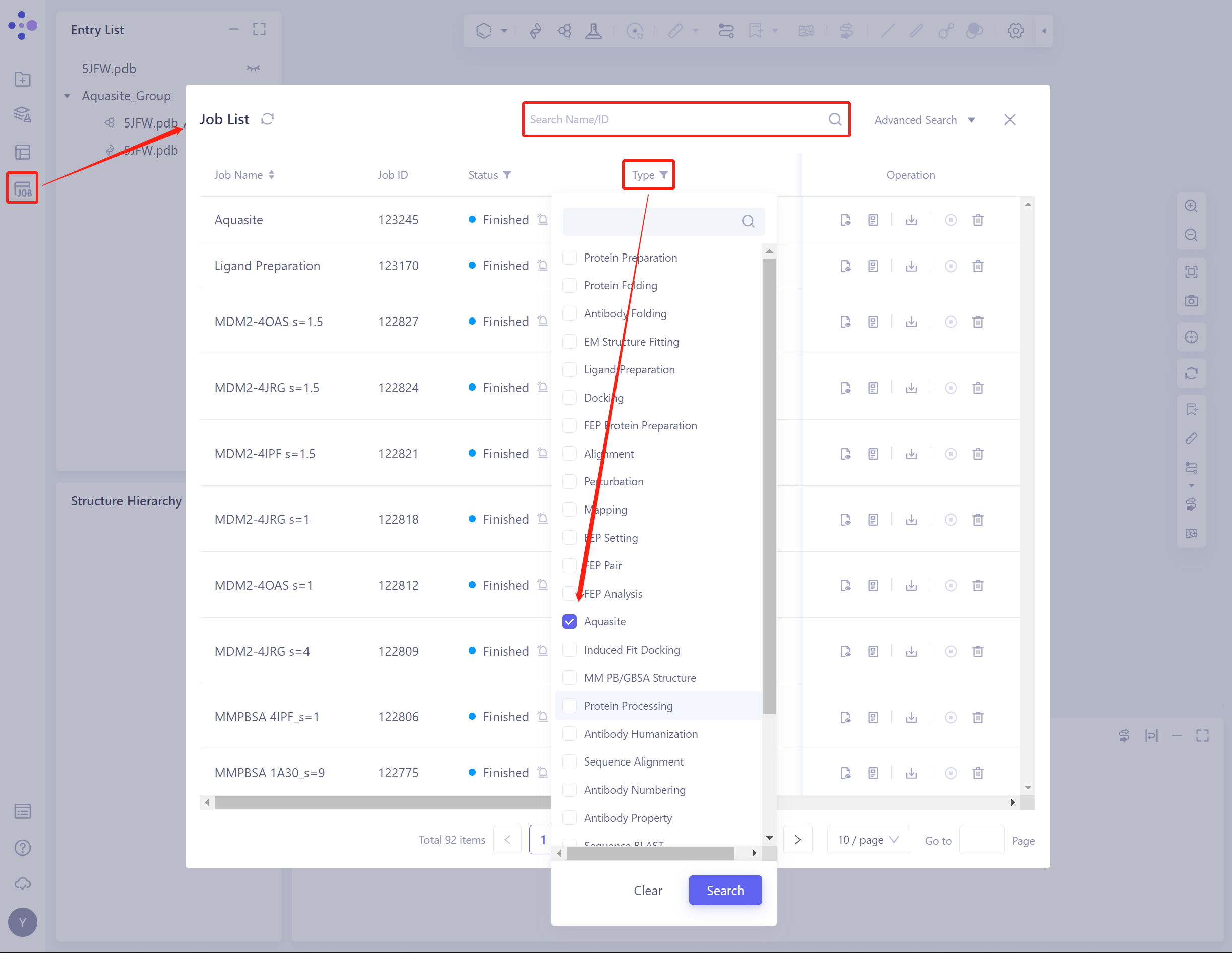
- Select the task to be viewed, and click Show in the Operation column to display the result of the task.
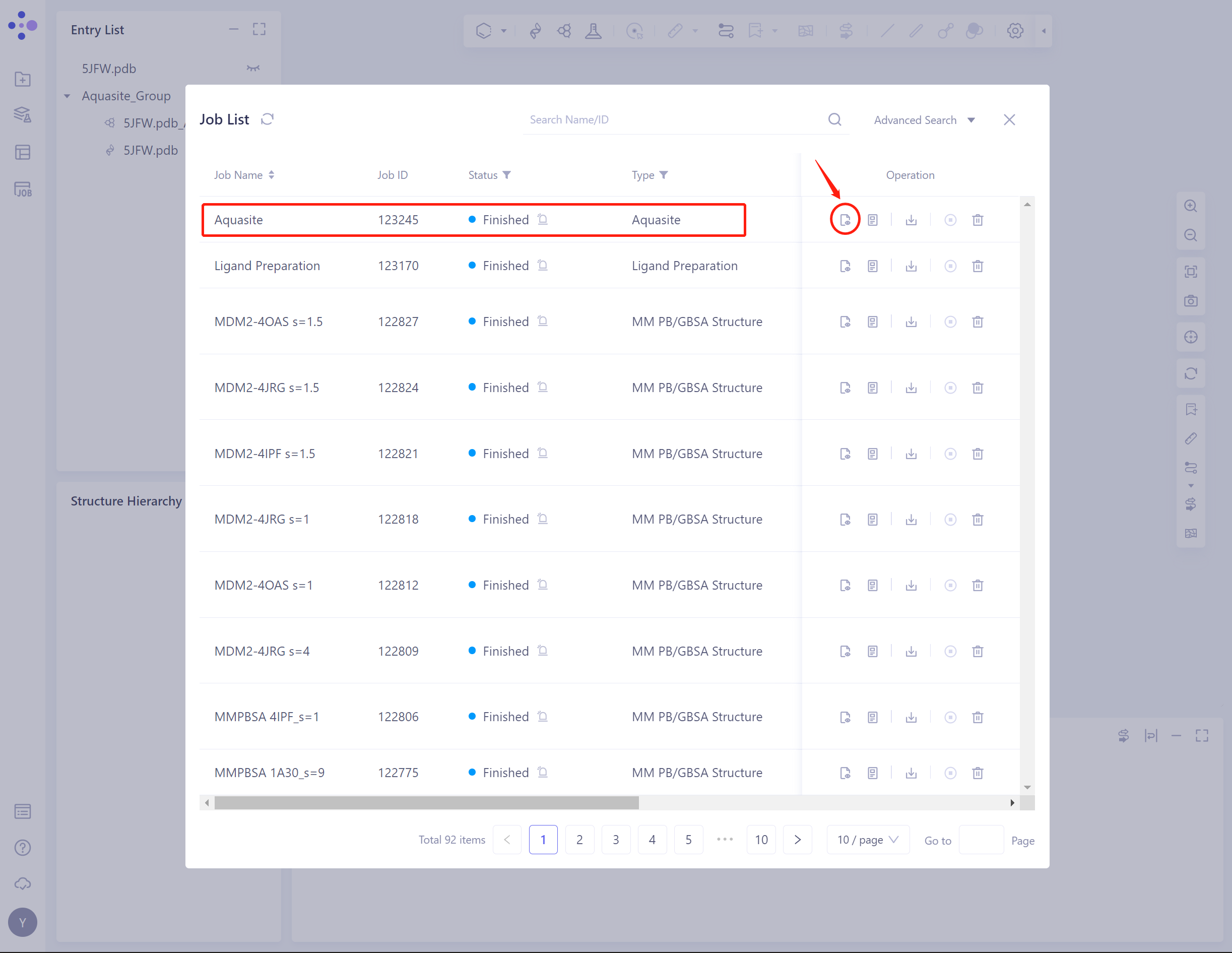
3.2 Results presentation
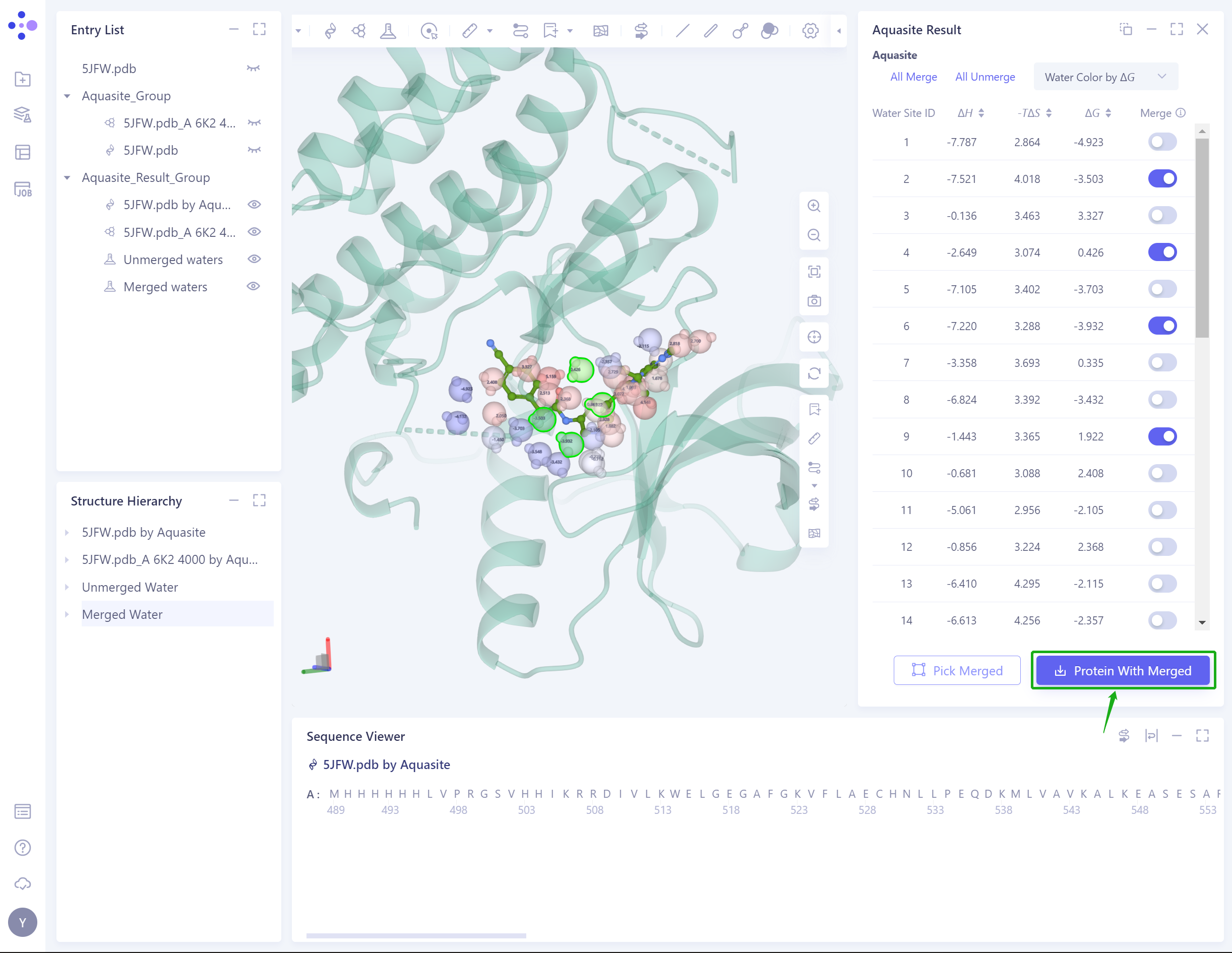
- The Result: list shows the predicted binding free energy of water molecules and the contribution of enthalpy change and entropy change, and the contribution of each energy is identified in 3D Workspace.
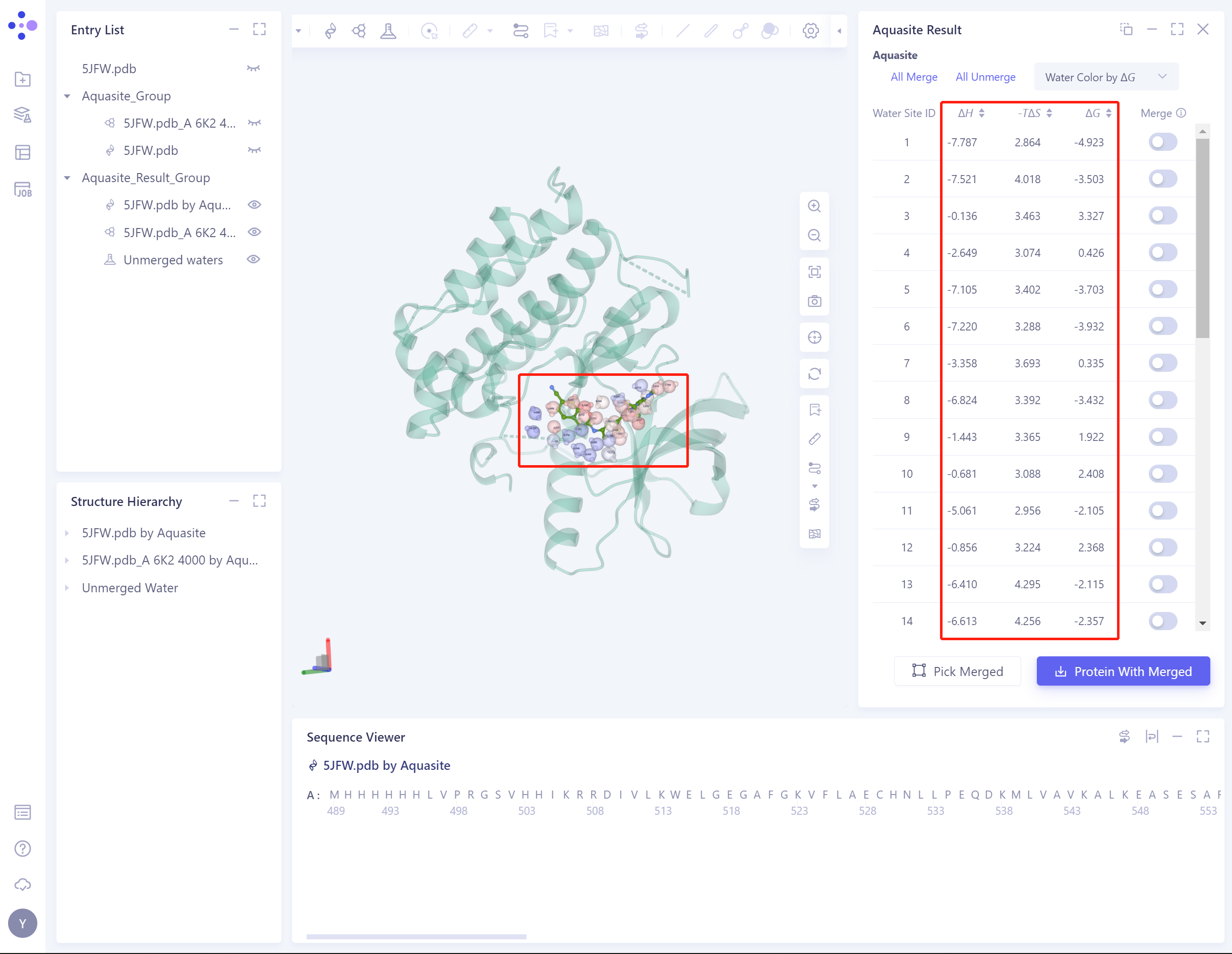
- Supports coloring water molecules with binding free energy, enthalpy change, or entropy change:
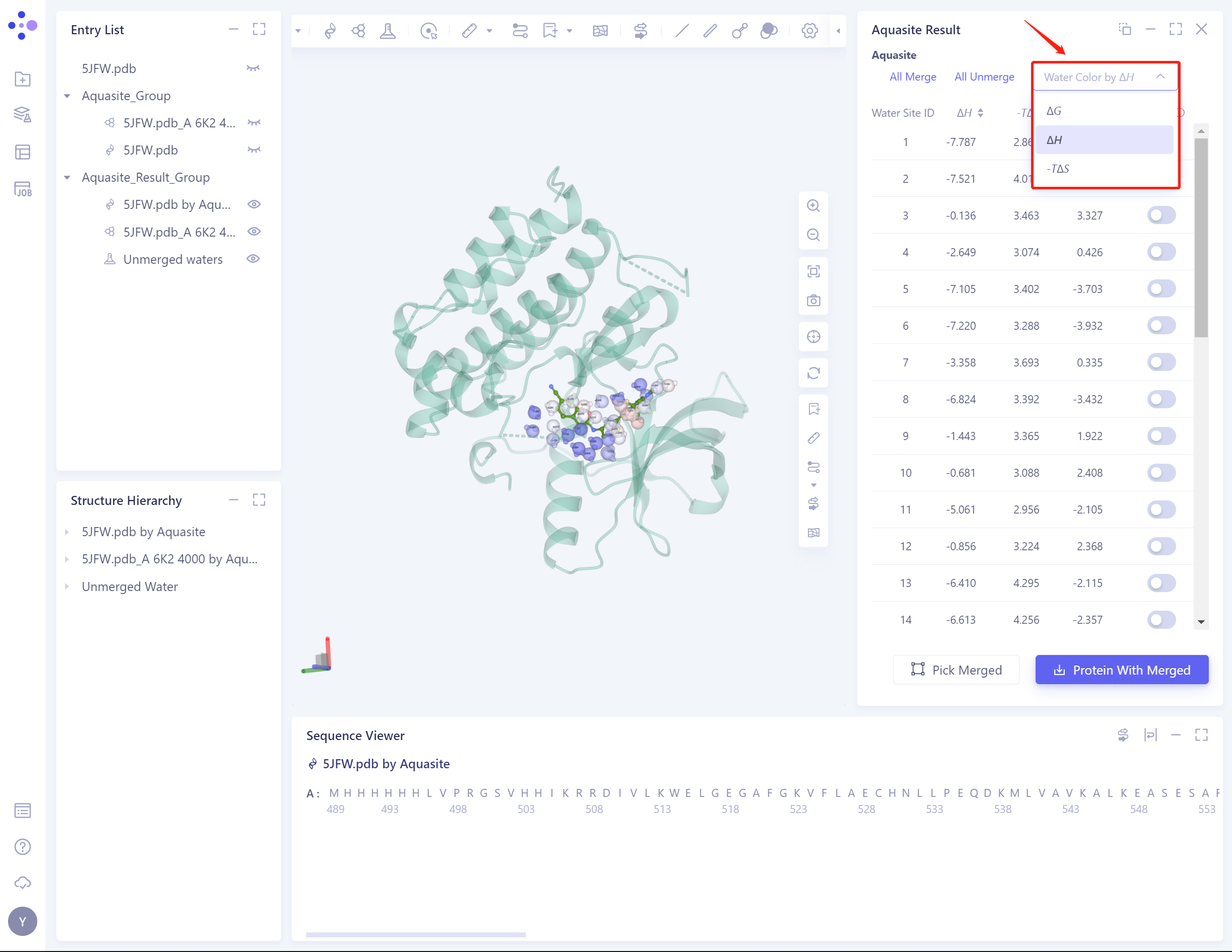
-
Merge:
-
Red box: from the predicted water molecules, select the water molecules to be retained in the system and continue to be used subsequently;
-
Yellow box: keep the batch shortcut of water molecules.
-
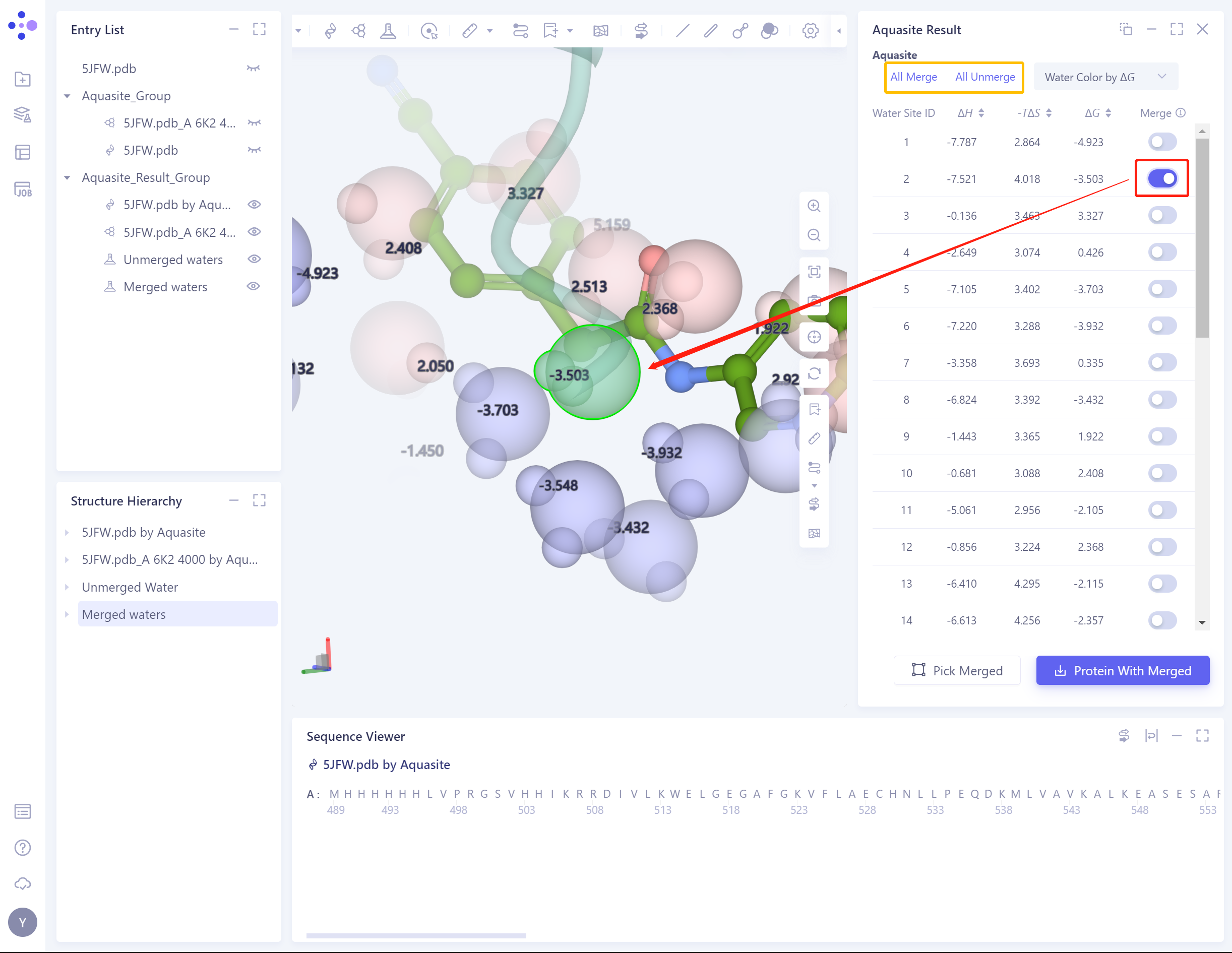
- Pick Merged: Click the green box to highlight all Merged water molecules, so that users can check whether all the water molecules they want to keep have been selected before downloading.
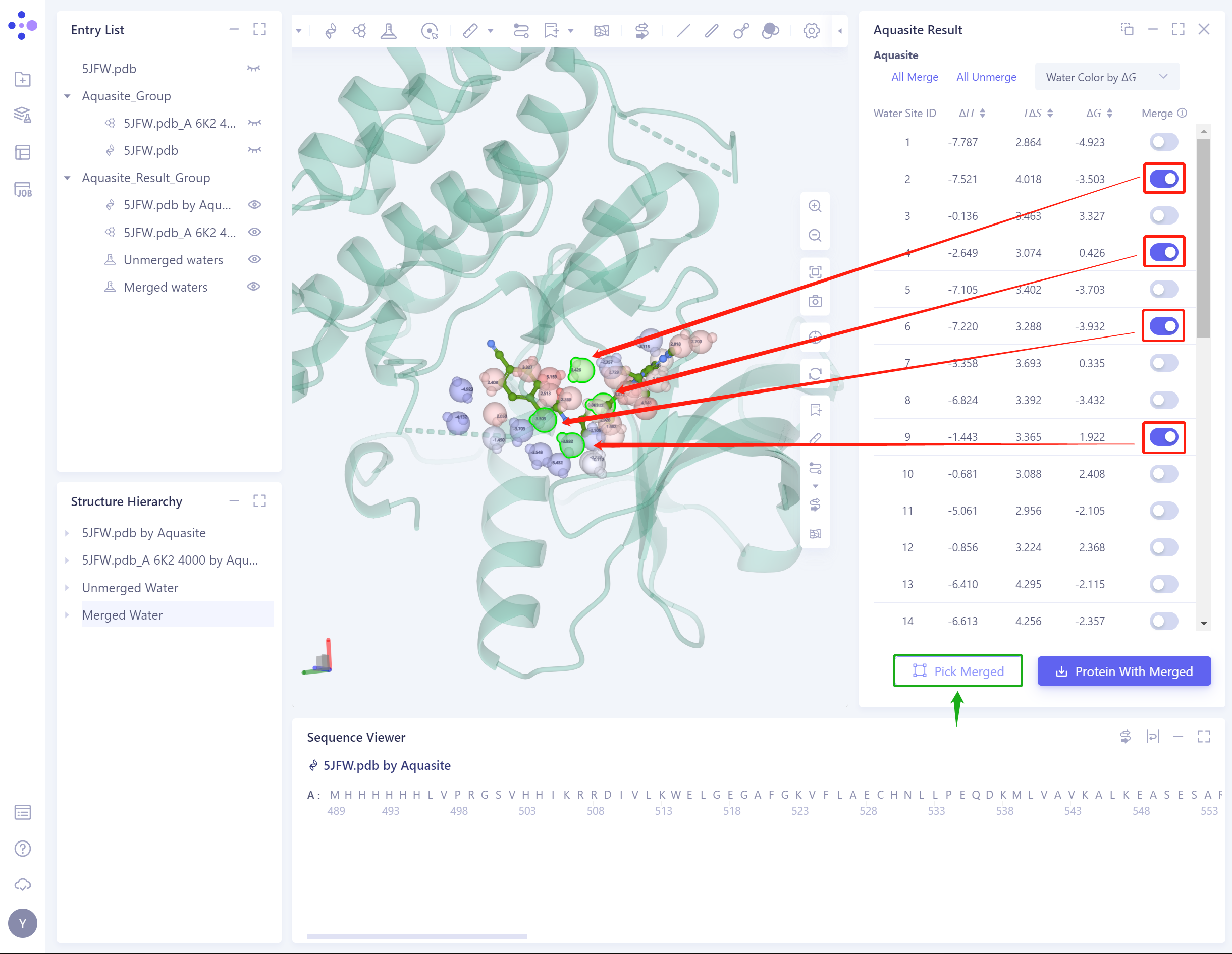
- Protein with Merged: The water molecules and proteins of Merged are downloaded as a whole system in the format of.pdb.
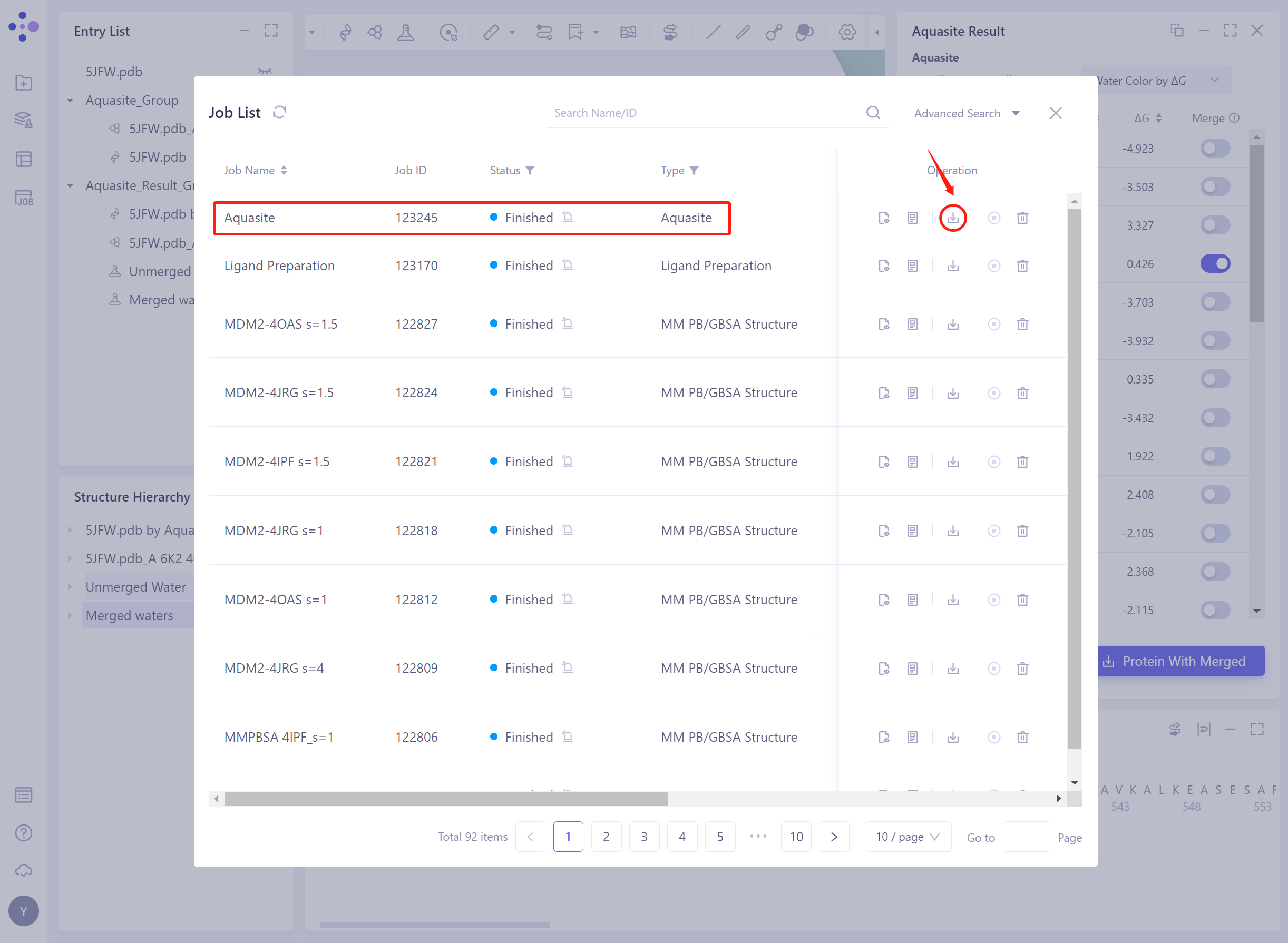
- You can also download the result file from the Job List, which contains all the predicted water molecules, if you want to fine-tune, see above.
- Water molecules with different binding free energies, stained as follows:
